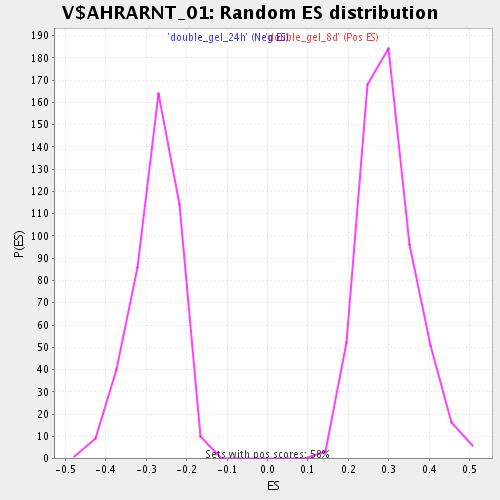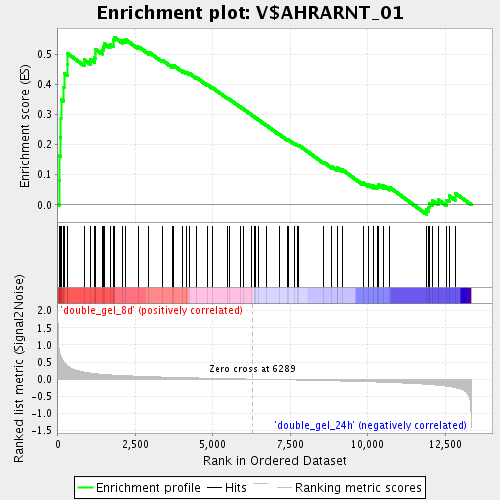
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$AHRARNT_01 |
| Enrichment Score (ES) | 0.5566408 |
| Normalized Enrichment Score (NES) | 1.8609873 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.01728834 |
| FWER p-Value | 0.719 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 37 | 0.888 | 0.0818 | Yes |
| 2 | WNT4 | WNT4 Entrez, Source | wingless-type MMTV integration site family, member 4 | 42 | 0.852 | 0.1626 | Yes |
| 3 | TACC1 | TACC1 Entrez, Source | transforming, acidic coiled-coil containing protein 1 | 80 | 0.688 | 0.2254 | Yes |
| 4 | BHLHB3 | BHLHB3 Entrez, Source | basic helix-loop-helix domain containing, class B, 3 | 98 | 0.666 | 0.2876 | Yes |
| 5 | SOAT1 | SOAT1 Entrez, Source | sterol O-acyltransferase (acyl-Coenzyme A: cholesterol acyltransferase) 1 | 100 | 0.661 | 0.3505 | Yes |
| 6 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 191 | 0.498 | 0.3912 | Yes |
| 7 | KLF15 | KLF15 Entrez, Source | Kruppel-like factor 15 | 202 | 0.486 | 0.4367 | Yes |
| 8 | IHPK2 | IHPK2 Entrez, Source | inositol hexaphosphate kinase 2 | 313 | 0.388 | 0.4655 | Yes |
| 9 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 318 | 0.386 | 0.5020 | Yes |
| 10 | GRM7 | GRM7 Entrez, Source | glutamate receptor, metabotropic 7 | 845 | 0.206 | 0.4819 | Yes |
| 11 | GHR | GHR Entrez, Source | growth hormone receptor | 1057 | 0.176 | 0.4828 | Yes |
| 12 | MAGED2 | MAGED2 Entrez, Source | melanoma antigen family D, 2 | 1191 | 0.162 | 0.4882 | Yes |
| 13 | BDNF | BDNF Entrez, Source | brain-derived neurotrophic factor | 1218 | 0.159 | 0.5014 | Yes |
| 14 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 1221 | 0.159 | 0.5164 | Yes |
| 15 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 1427 | 0.140 | 0.5143 | Yes |
| 16 | JUB | JUB Entrez, Source | jub, ajuba homolog (Xenopus laevis) | 1462 | 0.138 | 0.5249 | Yes |
| 17 | HPCAL1 | HPCAL1 Entrez, Source | hippocalcin-like 1 | 1490 | 0.136 | 0.5358 | Yes |
| 18 | CNTNAP1 | CNTNAP1 Entrez, Source | contactin associated protein 1 | 1684 | 0.124 | 0.5330 | Yes |
| 19 | SEZ6 | SEZ6 Entrez, Source | seizure related 6 homolog (mouse) | 1784 | 0.118 | 0.5368 | Yes |
| 20 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 1785 | 0.118 | 0.5480 | Yes |
| 21 | ID4 | ID4 Entrez, Source | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein | 1818 | 0.116 | 0.5566 | Yes |
| 22 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 2076 | 0.105 | 0.5472 | No |
| 23 | CRH | CRH Entrez, Source | corticotropin releasing hormone | 2185 | 0.100 | 0.5486 | No |
| 24 | BCL11A | BCL11A Entrez, Source | B-cell CLL/lymphoma 11A (zinc finger protein) | 2584 | 0.084 | 0.5266 | No |
| 25 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 2930 | 0.073 | 0.5076 | No |
| 26 | RGS6 | RGS6 Entrez, Source | regulator of G-protein signalling 6 | 3368 | 0.061 | 0.4805 | No |
| 27 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 3704 | 0.052 | 0.4602 | No |
| 28 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 3725 | 0.052 | 0.4637 | No |
| 29 | REPIN1 | REPIN1 Entrez, Source | replication initiator 1 | 4032 | 0.044 | 0.4448 | No |
| 30 | PTF1A | PTF1A Entrez, Source | pancreas specific transcription factor, 1a | 4139 | 0.041 | 0.4407 | No |
| 31 | HES1 | HES1 Entrez, Source | hairy and enhancer of split 1, (Drosophila) | 4248 | 0.038 | 0.4363 | No |
| 32 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 4456 | 0.035 | 0.4240 | No |
| 33 | CYP26B1 | CYP26B1 Entrez, Source | cytochrome P450, family 26, subfamily B, polypeptide 1 | 4829 | 0.027 | 0.3986 | No |
| 34 | RGS8 | RGS8 Entrez, Source | regulator of G-protein signalling 8 | 4985 | 0.024 | 0.3892 | No |
| 35 | NLGN3 | NLGN3 Entrez, Source | neuroligin 3 | 5467 | 0.016 | 0.3544 | No |
| 36 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 5523 | 0.014 | 0.3517 | No |
| 37 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 5876 | 0.008 | 0.3259 | No |
| 38 | CAMK2D | CAMK2D Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II delta | 5975 | 0.006 | 0.3190 | No |
| 39 | LHX1 | LHX1 Entrez, Source | LIM homeobox 1 | 6258 | 0.000 | 0.2978 | No |
| 40 | HK2 | HK2 Entrez, Source | hexokinase 2 | 6340 | -0.001 | 0.2918 | No |
| 41 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 6380 | -0.001 | 0.2890 | No |
| 42 | GNAO1 | GNAO1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O | 6475 | -0.003 | 0.2822 | No |
| 43 | PHF1 | PHF1 Entrez, Source | PHD finger protein 1 | 6733 | -0.008 | 0.2636 | No |
| 44 | MAEA | MAEA Entrez, Source | macrophage erythroblast attacher | 7146 | -0.015 | 0.2339 | No |
| 45 | FBXL20 | FBXL20 Entrez, Source | F-box and leucine-rich repeat protein 20 | 7404 | -0.020 | 0.2165 | No |
| 46 | TAF11 | TAF11 Entrez, Source | TAF11 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 28kDa | 7434 | -0.020 | 0.2162 | No |
| 47 | GABRA1 | GABRA1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 1 | 7649 | -0.025 | 0.2024 | No |
| 48 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 7732 | -0.026 | 0.1987 | No |
| 49 | SHC1 | SHC1 Entrez, Source | SHC (Src homology 2 domain containing) transforming protein 1 | 7776 | -0.027 | 0.1980 | No |
| 50 | KAZALD1 | KAZALD1 Entrez, Source | Kazal-type serine peptidase inhibitor domain 1 | 8576 | -0.043 | 0.1419 | No |
| 51 | VDP | VDP Entrez, Source | - | 8845 | -0.049 | 0.1264 | No |
| 52 | NEUROD2 | NEUROD2 Entrez, Source | neurogenic differentiation 2 | 9017 | -0.052 | 0.1185 | No |
| 53 | SYT12 | SYT12 Entrez, Source | synaptotagmin XII | 9021 | -0.052 | 0.1232 | No |
| 54 | IL7 | IL7 Entrez, Source | interleukin 7 | 9178 | -0.055 | 0.1168 | No |
| 55 | NR3C1 | NR3C1 Entrez, Source | nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) | 9848 | -0.072 | 0.0733 | No |
| 56 | TNPO3 | TNPO3 Entrez, Source | transportin 3 | 10016 | -0.077 | 0.0680 | No |
| 57 | RBBP7 | RBBP7 Entrez, Source | retinoblastoma binding protein 7 | 10181 | -0.081 | 0.0634 | No |
| 58 | WSB1 | WSB1 Entrez, Source | WD repeat and SOCS box-containing 1 | 10301 | -0.084 | 0.0625 | No |
| 59 | PABPC1 | PABPC1 Entrez, Source | poly(A) binding protein, cytoplasmic 1 | 10349 | -0.086 | 0.0671 | No |
| 60 | UGCGL1 | UGCGL1 Entrez, Source | UDP-glucose ceramide glucosyltransferase-like 1 | 10515 | -0.091 | 0.0633 | No |
| 61 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 10702 | -0.097 | 0.0585 | No |
| 62 | ABTB2 | ABTB2 Entrez, Source | ankyrin repeat and BTB (POZ) domain containing 2 | 11886 | -0.149 | -0.0163 | No |
| 63 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 11961 | -0.154 | -0.0073 | No |
| 64 | EIF4A2 | EIF4A2 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 2 | 11996 | -0.157 | 0.0051 | No |
| 65 | XPO6 | XPO6 Entrez, Source | exportin 6 | 12092 | -0.164 | 0.0136 | No |
| 66 | TRIM23 | TRIM23 Entrez, Source | tripartite motif-containing 23 | 12278 | -0.177 | 0.0166 | No |
| 67 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 12547 | -0.204 | 0.0158 | No |
| 68 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 12629 | -0.215 | 0.0302 | No |
| 69 | GTF2A1 | GTF2A1 Entrez, Source | general transcription factor IIA, 1, 19/37kDa | 12822 | -0.246 | 0.0392 | No |