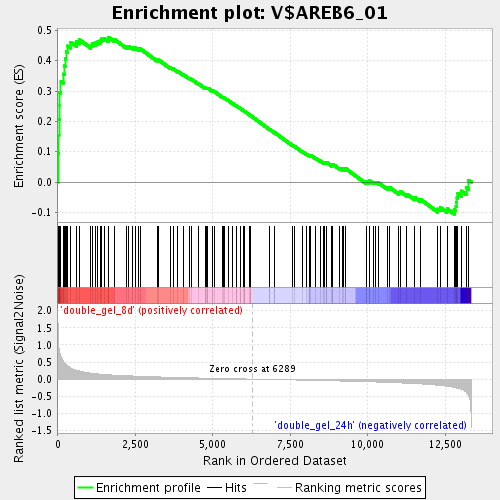
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$AREB6_01 |
| Enrichment Score (ES) | 0.47752327 |
| Normalized Enrichment Score (NES) | 1.6954372 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.057442974 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 3 | 1.622 | 0.0957 | Yes |
| 2 | LAMC2 | LAMC2 Entrez, Source | laminin, gamma 2 | 23 | 1.021 | 0.1547 | Yes |
| 3 | CLDN7 | CLDN7 Entrez, Source | claudin 7 | 36 | 0.888 | 0.2063 | Yes |
| 4 | ANKRD1 | ANKRD1 Entrez, Source | ankyrin repeat domain 1 (cardiac muscle) | 56 | 0.784 | 0.2513 | Yes |
| 5 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 60 | 0.762 | 0.2961 | Yes |
| 6 | BHLHB3 | BHLHB3 Entrez, Source | basic helix-loop-helix domain containing, class B, 3 | 98 | 0.666 | 0.3328 | Yes |
| 7 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 185 | 0.510 | 0.3564 | Yes |
| 8 | USP2 | USP2 Entrez, Source | ubiquitin specific peptidase 2 | 207 | 0.480 | 0.3832 | Yes |
| 9 | KIF12 | KIF12 Entrez, Source | kinesin family member 12 | 249 | 0.442 | 0.4063 | Yes |
| 10 | DDR1 | DDR1 Entrez, Source | discoidin domain receptor family, member 1 | 262 | 0.429 | 0.4307 | Yes |
| 11 | PC | PC Entrez, Source | pyruvate carboxylase | 322 | 0.380 | 0.4488 | Yes |
| 12 | CDH13 | CDH13 Entrez, Source | cadherin 13, H-cadherin (heart) | 417 | 0.328 | 0.4611 | Yes |
| 13 | PXN | PXN Entrez, Source | paxillin | 594 | 0.258 | 0.4630 | Yes |
| 14 | C1QA | C1QA Entrez, Source | complement component 1, q subcomponent, A chain | 699 | 0.236 | 0.4691 | Yes |
| 15 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1066 | 0.174 | 0.4518 | Yes |
| 16 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 1115 | 0.169 | 0.4582 | Yes |
| 17 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 1221 | 0.159 | 0.4597 | Yes |
| 18 | ACTN3 | ACTN3 Entrez, Source | actinin, alpha 3 | 1290 | 0.152 | 0.4635 | Yes |
| 19 | LRP10 | LRP10 Entrez, Source | low density lipoprotein receptor-related protein 10 | 1369 | 0.145 | 0.4662 | Yes |
| 20 | TEF | TEF Entrez, Source | thyrotrophic embryonic factor | 1404 | 0.143 | 0.4720 | Yes |
| 21 | CTHRC1 | CTHRC1 Entrez, Source | collagen triple helix repeat containing 1 | 1491 | 0.136 | 0.4736 | Yes |
| 22 | FKRP | FKRP Entrez, Source | fukutin related protein | 1635 | 0.126 | 0.4702 | Yes |
| 23 | CRB3 | CRB3 Entrez, Source | crumbs homolog 3 (Drosophila) | 1638 | 0.126 | 0.4775 | Yes |
| 24 | ID4 | ID4 Entrez, Source | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein | 1818 | 0.116 | 0.4709 | No |
| 25 | SLC6A11 | SLC6A11 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, GABA), member 11 | 2212 | 0.099 | 0.4470 | No |
| 26 | IL2 | IL2 Entrez, Source | interleukin 2 | 2282 | 0.096 | 0.4475 | No |
| 27 | VWF | VWF Entrez, Source | von Willebrand factor | 2400 | 0.092 | 0.4441 | No |
| 28 | AGC1 | AGC1 Entrez, Source | aggrecan 1 (chondroitin sulfate proteoglycan 1, large aggregating proteoglycan, antigen identified by monoclonal antibody A0122) | 2489 | 0.088 | 0.4426 | No |
| 29 | IGFBP6 | IGFBP6 Entrez, Source | insulin-like growth factor binding protein 6 | 2598 | 0.084 | 0.4394 | No |
| 30 | CRELD1 | CRELD1 Entrez, Source | cysteine-rich with EGF-like domains 1 | 2660 | 0.082 | 0.4396 | No |
| 31 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 3202 | 0.066 | 0.4026 | No |
| 32 | SFRP2 | SFRP2 Entrez, Source | secreted frizzled-related protein 2 | 3247 | 0.064 | 0.4031 | No |
| 33 | EGR4 | EGR4 Entrez, Source | early growth response 4 | 3630 | 0.054 | 0.3774 | No |
| 34 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 3715 | 0.052 | 0.3741 | No |
| 35 | HCLS1 | HCLS1 Entrez, Source | hematopoietic cell-specific Lyn substrate 1 | 3872 | 0.048 | 0.3652 | No |
| 36 | CUGBP1 | CUGBP1 Entrez, Source | CUG triplet repeat, RNA binding protein 1 | 4049 | 0.043 | 0.3545 | No |
| 37 | HPCAL4 | HPCAL4 Entrez, Source | hippocalcin like 4 | 4243 | 0.039 | 0.3422 | No |
| 38 | NKX6-1 | NKX6-1 Entrez, Source | NK6 transcription factor related, locus 1 (Drosophila) | 4306 | 0.037 | 0.3397 | No |
| 39 | C6ORF47 | C6ORF47 Entrez, Source | chromosome 6 open reading frame 47 | 4532 | 0.033 | 0.3246 | No |
| 40 | HCRTR1 | HCRTR1 Entrez, Source | hypocretin (orexin) receptor 1 | 4747 | 0.029 | 0.3102 | No |
| 41 | STX1A | STX1A Entrez, Source | syntaxin 1A (brain) | 4763 | 0.029 | 0.3108 | No |
| 42 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 4797 | 0.028 | 0.3099 | No |
| 43 | CYP26B1 | CYP26B1 Entrez, Source | cytochrome P450, family 26, subfamily B, polypeptide 1 | 4829 | 0.027 | 0.3092 | No |
| 44 | HEPH | HEPH Entrez, Source | hephaestin | 4842 | 0.027 | 0.3099 | No |
| 45 | THRA | THRA Entrez, Source | thyroid hormone receptor, alpha (erythroblastic leukemia viral (v-erb-a) oncogene homolog, avian) | 4984 | 0.024 | 0.3007 | No |
| 46 | SYT3 | SYT3 Entrez, Source | synaptotagmin III | 4986 | 0.024 | 0.3020 | No |
| 47 | CEL | CEL Entrez, Source | carboxyl ester lipase (bile salt-stimulated lipase) | 5047 | 0.023 | 0.2989 | No |
| 48 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 5323 | 0.018 | 0.2792 | No |
| 49 | NRXN2 | NRXN2 Entrez, Source | neurexin 2 | 5341 | 0.018 | 0.2790 | No |
| 50 | SLC16A8 | SLC16A8 Entrez, Source | solute carrier family 16, member 8 (monocarboxylic acid transporter 3) | 5380 | 0.017 | 0.2771 | No |
| 51 | RRAD | RRAD Entrez, Source | Ras-related associated with diabetes | 5502 | 0.015 | 0.2688 | No |
| 52 | BTG4 | BTG4 Entrez, Source | B-cell translocation gene 4 | 5638 | 0.012 | 0.2593 | No |
| 53 | FXYD3 | FXYD3 Entrez, Source | FXYD domain containing ion transport regulator 3 | 5757 | 0.010 | 0.2510 | No |
| 54 | NRXN3 | NRXN3 Entrez, Source | neurexin 3 | 5770 | 0.010 | 0.2507 | No |
| 55 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 5876 | 0.008 | 0.2432 | No |
| 56 | KCNH3 | KCNH3 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 3 | 5891 | 0.007 | 0.2425 | No |
| 57 | CXXC5 | CXXC5 Entrez, Source | CXXC finger 5 | 5992 | 0.005 | 0.2353 | No |
| 58 | GRK6 | GRK6 Entrez, Source | G protein-coupled receptor kinase 6 | 6012 | 0.005 | 0.2341 | No |
| 59 | SLC16A6 | SLC16A6 Entrez, Source | solute carrier family 16, member 6 (monocarboxylic acid transporter 7) | 6019 | 0.005 | 0.2340 | No |
| 60 | PCP4 | PCP4 Entrez, Source | Purkinje cell protein 4 | 6190 | 0.001 | 0.2212 | No |
| 61 | RASGRF2 | RASGRF2 Entrez, Source | Ras protein-specific guanine nucleotide-releasing factor 2 | 6203 | 0.001 | 0.2204 | No |
| 62 | MCAM | MCAM Entrez, Source | melanoma cell adhesion molecule | 6217 | 0.001 | 0.2195 | No |
| 63 | PAX3 | PAX3 Entrez, Source | paired box gene 3 (Waardenburg syndrome 1) | 6824 | -0.009 | 0.1742 | No |
| 64 | ZIC1 | ZIC1 Entrez, Source | Zic family member 1 (odd-paired homolog, Drosophila) | 6837 | -0.009 | 0.1738 | No |
| 65 | FLNB | FLNB Entrez, Source | filamin B, beta (actin binding protein 278) | 7002 | -0.012 | 0.1622 | No |
| 66 | NXPH4 | NXPH4 Entrez, Source | neurexophilin 4 | 7004 | -0.012 | 0.1628 | No |
| 67 | SECISBP2 | SECISBP2 Entrez, Source | SECIS binding protein 2 | 7576 | -0.023 | 0.1210 | No |
| 68 | TRIB1 | TRIB1 Entrez, Source | tribbles homolog 1 (Drosophila) | 7632 | -0.024 | 0.1183 | No |
| 69 | GABARAP | GABARAP Entrez, Source | GABA(A) receptor-associated protein | 7908 | -0.029 | 0.0993 | No |
| 70 | UMOD | UMOD Entrez, Source | uromodulin (uromucoid, Tamm-Horsfall glycoprotein) | 8036 | -0.032 | 0.0915 | No |
| 71 | SHH | SHH Entrez, Source | sonic hedgehog homolog (Drosophila) | 8114 | -0.034 | 0.0877 | No |
| 72 | CLDN5 | CLDN5 Entrez, Source | claudin 5 (transmembrane protein deleted in velocardiofacial syndrome) | 8120 | -0.034 | 0.0893 | No |
| 73 | IL12B | IL12B Entrez, Source | interleukin 12B (natural killer cell stimulatory factor 2, cytotoxic lymphocyte maturation factor 2, p40) | 8161 | -0.035 | 0.0884 | No |
| 74 | SOSTDC1 | SOSTDC1 Entrez, Source | sclerostin domain containing 1 | 8320 | -0.038 | 0.0787 | No |
| 75 | LSP1 | LSP1 Entrez, Source | lymphocyte-specific protein 1 | 8463 | -0.041 | 0.0703 | No |
| 76 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 8582 | -0.043 | 0.0640 | No |
| 77 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 8595 | -0.043 | 0.0656 | No |
| 78 | KCNH7 | KCNH7 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 7 | 8656 | -0.045 | 0.0637 | No |
| 79 | TITF1 | TITF1 Entrez, Source | thyroid transcription factor 1 | 8658 | -0.045 | 0.0663 | No |
| 80 | VDP | VDP Entrez, Source | - | 8845 | -0.049 | 0.0551 | No |
| 81 | TCF1 | TCF1 Entrez, Source | transcription factor 1, hepatic; LF-B1, hepatic nuclear factor (HNF1), albumin proximal factor | 8858 | -0.049 | 0.0571 | No |
| 82 | CAMKK1 | CAMKK1 Entrez, Source | calcium/calmodulin-dependent protein kinase kinase 1, alpha | 8876 | -0.049 | 0.0587 | No |
| 83 | FGF5 | FGF5 Entrez, Source | fibroblast growth factor 5 | 9095 | -0.054 | 0.0455 | No |
| 84 | IL7 | IL7 Entrez, Source | interleukin 7 | 9178 | -0.055 | 0.0425 | No |
| 85 | CHRM1 | CHRM1 Entrez, Source | cholinergic receptor, muscarinic 1 | 9204 | -0.056 | 0.0440 | No |
| 86 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 9270 | -0.057 | 0.0424 | No |
| 87 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 9277 | -0.058 | 0.0454 | No |
| 88 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 9947 | -0.075 | -0.0007 | No |
| 89 | JUP | JUP Entrez, Source | junction plakoglobin | 9963 | -0.075 | 0.0026 | No |
| 90 | RPS27A | RPS27A Entrez, Source | ribosomal protein S27a | 10042 | -0.078 | 0.0013 | No |
| 91 | SOX10 | SOX10 Entrez, Source | SRY (sex determining region Y)-box 10 | 10051 | -0.078 | 0.0053 | No |
| 92 | TDRD3 | TDRD3 Entrez, Source | tudor domain containing 3 | 10193 | -0.081 | -0.0005 | No |
| 93 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 10248 | -0.083 | 0.0003 | No |
| 94 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 10344 | -0.086 | -0.0018 | No |
| 95 | FGF17 | FGF17 Entrez, Source | fibroblast growth factor 17 | 10631 | -0.095 | -0.0178 | No |
| 96 | LRRFIP2 | LRRFIP2 Entrez, Source | leucine rich repeat (in FLII) interacting protein 2 | 10707 | -0.097 | -0.0177 | No |
| 97 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 11004 | -0.108 | -0.0337 | No |
| 98 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 11050 | -0.110 | -0.0306 | No |
| 99 | TRPM7 | TRPM7 Entrez, Source | transient receptor potential cation channel, subfamily M, member 7 | 11259 | -0.119 | -0.0393 | No |
| 100 | EIF5 | EIF5 Entrez, Source | eukaryotic translation initiation factor 5 | 11521 | -0.130 | -0.0513 | No |
| 101 | BMF | BMF Entrez, Source | Bcl2 modifying factor | 11710 | -0.139 | -0.0573 | No |
| 102 | DNMT3A | DNMT3A Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 alpha | 12263 | -0.177 | -0.0886 | No |
| 103 | TUBA4 | TUBA4 Entrez, Source | tubulin, alpha 4 | 12345 | -0.183 | -0.0839 | No |
| 104 | CASK | CASK Entrez, Source | calcium/calmodulin-dependent serine protein kinase (MAGUK family) | 12559 | -0.206 | -0.0878 | No |
| 105 | ITGB3BP | ITGB3BP Entrez, Source | integrin beta 3 binding protein (beta3-endonexin) | 12793 | -0.240 | -0.0913 | No |
| 106 | REV3L | REV3L Entrez, Source | REV3-like, catalytic subunit of DNA polymerase zeta (yeast) | 12847 | -0.253 | -0.0803 | No |
| 107 | RNF38 | RNF38 Entrez, Source | ring finger protein 38 | 12858 | -0.256 | -0.0659 | No |
| 108 | VAT1 | VAT1 Entrez, Source | vesicle amine transport protein 1 homolog (T californica) | 12879 | -0.262 | -0.0520 | No |
| 109 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 12908 | -0.269 | -0.0382 | No |
| 110 | CSAD | CSAD Entrez, Source | cysteine sulfinic acid decarboxylase | 13026 | -0.301 | -0.0292 | No |
| 111 | EDG1 | EDG1 Entrez, Source | endothelial differentiation, sphingolipid G-protein-coupled receptor, 1 | 13191 | -0.391 | -0.0185 | No |
| 112 | LAMB3 | LAMB3 Entrez, Source | laminin, beta 3 | 13259 | -0.504 | 0.0063 | No |

