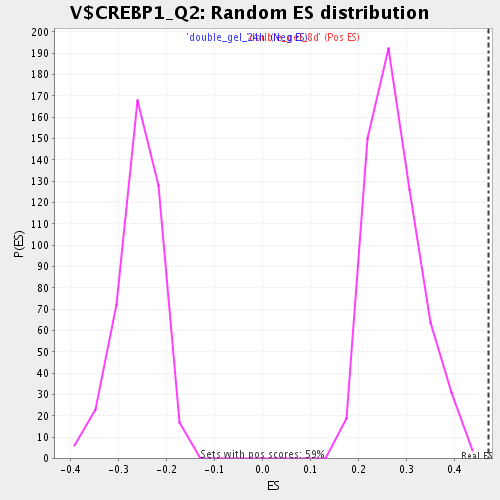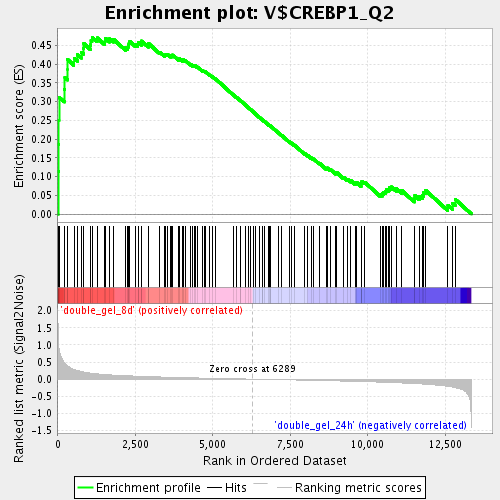
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$CREBP1_Q2 |
| Enrichment Score (ES) | 0.4719661 |
| Normalized Enrichment Score (NES) | 1.7076997 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.052497298 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 3 | 1.622 | 0.1135 | Yes |
| 2 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 21 | 1.039 | 0.1851 | Yes |
| 3 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 31 | 0.935 | 0.2500 | Yes |
| 4 | CLDN7 | CLDN7 Entrez, Source | claudin 7 | 36 | 0.888 | 0.3119 | Yes |
| 5 | USP2 | USP2 Entrez, Source | ubiquitin specific peptidase 2 | 207 | 0.480 | 0.3327 | Yes |
| 6 | NCALD | NCALD Entrez, Source | neurocalcin delta | 225 | 0.463 | 0.3639 | Yes |
| 7 | HS3ST2 | HS3ST2 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 2 | 308 | 0.394 | 0.3853 | Yes |
| 8 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 312 | 0.390 | 0.4124 | Yes |
| 9 | BAT5 | BAT5 Entrez, Source | HLA-B associated transcript 5 | 519 | 0.279 | 0.4164 | Yes |
| 10 | SLC38A2 | SLC38A2 Entrez, Source | solute carrier family 38, member 2 | 631 | 0.249 | 0.4255 | Yes |
| 11 | ICA1 | ICA1 Entrez, Source | islet cell autoantigen 1, 69kDa | 753 | 0.226 | 0.4321 | Yes |
| 12 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 824 | 0.209 | 0.4415 | Yes |
| 13 | FLT1 | FLT1 Entrez, Source | fms-related tyrosine kinase 1 (vascular endothelial growth factor/vascular permeability factor receptor) | 835 | 0.207 | 0.4552 | Yes |
| 14 | PITX2 | PITX2 Entrez, Source | paired-like homeodomain transcription factor 2 | 1047 | 0.177 | 0.4517 | Yes |
| 15 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1066 | 0.174 | 0.4626 | Yes |
| 16 | SLC25A25 | SLC25A25 Entrez, Source | solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 25 | 1101 | 0.171 | 0.4720 | Yes |
| 17 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 1266 | 0.154 | 0.4704 | No |
| 18 | CALML3 | CALML3 Entrez, Source | calmodulin-like 3 | 1518 | 0.133 | 0.4608 | No |
| 19 | GRM3 | GRM3 Entrez, Source | glutamate receptor, metabotropic 3 | 1523 | 0.133 | 0.4698 | No |
| 20 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 1666 | 0.124 | 0.4678 | No |
| 21 | CBX8 | CBX8 Entrez, Source | chromobox homolog 8 (Pc class homolog, Drosophila) | 1783 | 0.118 | 0.4673 | No |
| 22 | CRH | CRH Entrez, Source | corticotropin releasing hormone | 2185 | 0.100 | 0.4440 | No |
| 23 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 2261 | 0.097 | 0.4451 | No |
| 24 | NAP1L2 | NAP1L2 Entrez, Source | nucleosome assembly protein 1-like 2 | 2268 | 0.097 | 0.4514 | No |
| 25 | TPM4 | TPM4 Entrez, Source | tropomyosin 4 | 2293 | 0.096 | 0.4563 | No |
| 26 | PCSK1 | PCSK1 Entrez, Source | proprotein convertase subtilisin/kexin type 1 | 2319 | 0.095 | 0.4611 | No |
| 27 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 2510 | 0.088 | 0.4528 | No |
| 28 | BCL11A | BCL11A Entrez, Source | B-cell CLL/lymphoma 11A (zinc finger protein) | 2584 | 0.084 | 0.4532 | No |
| 29 | BRAF | BRAF Entrez, Source | v-raf murine sarcoma viral oncogene homolog B1 | 2605 | 0.083 | 0.4575 | No |
| 30 | CX36 | CX36 Entrez, Source | - | 2695 | 0.080 | 0.4565 | No |
| 31 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 2696 | 0.080 | 0.4621 | No |
| 32 | PNRC1 | PNRC1 Entrez, Source | proline-rich nuclear receptor coactivator 1 | 2915 | 0.074 | 0.4508 | No |
| 33 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 2930 | 0.073 | 0.4549 | No |
| 34 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 3286 | 0.063 | 0.4325 | No |
| 35 | AGPAT4 | AGPAT4 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 4 (lysophosphatidic acid acyltransferase, delta) | 3447 | 0.059 | 0.4245 | No |
| 36 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 3479 | 0.058 | 0.4263 | No |
| 37 | RBMS2 | RBMS2 Entrez, Source | RNA binding motif, single stranded interacting protein 2 | 3522 | 0.057 | 0.4271 | No |
| 38 | EGR4 | EGR4 Entrez, Source | early growth response 4 | 3630 | 0.054 | 0.4228 | No |
| 39 | PDLIM3 | PDLIM3 Entrez, Source | PDZ and LIM domain 3 | 3680 | 0.053 | 0.4228 | No |
| 40 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 3704 | 0.052 | 0.4247 | No |
| 41 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 3905 | 0.047 | 0.4129 | No |
| 42 | KCNK1 | KCNK1 Entrez, Source | potassium channel, subfamily K, member 1 | 3913 | 0.047 | 0.4156 | No |
| 43 | ELOVL5 | ELOVL5 Entrez, Source | ELOVL family member 5, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4036 | 0.044 | 0.4094 | No |
| 44 | MAP1LC3A | MAP1LC3A Entrez, Source | microtubule-associated protein 1 light chain 3 alpha | 4039 | 0.044 | 0.4123 | No |
| 45 | MESDC1 | MESDC1 Entrez, Source | mesoderm development candidate 1 | 4119 | 0.042 | 0.4093 | No |
| 46 | DIABLO | DIABLO Entrez, Source | diablo homolog (Drosophila) | 4276 | 0.038 | 0.4002 | No |
| 47 | ELL2 | ELL2 Entrez, Source | elongation factor, RNA polymerase II, 2 | 4354 | 0.037 | 0.3969 | No |
| 48 | SNAP25 | SNAP25 Entrez, Source | synaptosomal-associated protein, 25kDa | 4406 | 0.036 | 0.3956 | No |
| 49 | GNAS | GNAS Entrez, Source | GNAS complex locus | 4431 | 0.035 | 0.3962 | No |
| 50 | SLC6A4 | SLC6A4 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 | 4511 | 0.034 | 0.3926 | No |
| 51 | SPHK2 | SPHK2 Entrez, Source | sphingosine kinase 2 | 4673 | 0.031 | 0.3826 | No |
| 52 | SLC6A5 | SLC6A5 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, glycine), member 5 | 4715 | 0.030 | 0.3816 | No |
| 53 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 4753 | 0.029 | 0.3808 | No |
| 54 | CHGB | CHGB Entrez, Source | chromogranin B (secretogranin 1) | 4906 | 0.026 | 0.3711 | No |
| 55 | THRA | THRA Entrez, Source | thyroid hormone receptor, alpha (erythroblastic leukemia viral (v-erb-a) oncogene homolog, avian) | 4984 | 0.024 | 0.3670 | No |
| 56 | TACR1 | TACR1 Entrez, Source | tachykinin receptor 1 | 5085 | 0.023 | 0.3610 | No |
| 57 | SNF1LK | SNF1LK Entrez, Source | SNF1-like kinase | 5650 | 0.012 | 0.3192 | No |
| 58 | NAPA | NAPA Entrez, Source | N-ethylmaleimide-sensitive factor attachment protein, alpha | 5766 | 0.010 | 0.3112 | No |
| 59 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 5876 | 0.008 | 0.3035 | No |
| 60 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 5890 | 0.007 | 0.3030 | No |
| 61 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 6062 | 0.004 | 0.2904 | No |
| 62 | PER1 | PER1 Entrez, Source | period homolog 1 (Drosophila) | 6161 | 0.002 | 0.2831 | No |
| 63 | MCAM | MCAM Entrez, Source | melanoma cell adhesion molecule | 6217 | 0.001 | 0.2790 | No |
| 64 | RAB6A | RAB6A Entrez, Source | RAB6A, member RAS oncogene family | 6313 | -0.000 | 0.2719 | No |
| 65 | SCG2 | SCG2 Entrez, Source | secretogranin II (chromogranin C) | 6379 | -0.001 | 0.2670 | No |
| 66 | RPL41 | RPL41 Entrez, Source | ribosomal protein L41 | 6498 | -0.004 | 0.2584 | No |
| 67 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 6588 | -0.005 | 0.2520 | No |
| 68 | UCN | UCN Entrez, Source | urocortin | 6604 | -0.006 | 0.2513 | No |
| 69 | CBX3 | CBX3 Entrez, Source | chromobox homolog 3 (HP1 gamma homolog, Drosophila) | 6679 | -0.007 | 0.2462 | No |
| 70 | SCAMP5 | SCAMP5 Entrez, Source | secretory carrier membrane protein 5 | 6796 | -0.009 | 0.2380 | No |
| 71 | IKBKB | IKBKB Entrez, Source | inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase beta | 6828 | -0.009 | 0.2363 | No |
| 72 | CALM2 | CALM2 Entrez, Source | calmodulin 2 (phosphorylase kinase, delta) | 6851 | -0.010 | 0.2353 | No |
| 73 | KCNA5 | KCNA5 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 5 | 6863 | -0.010 | 0.2352 | No |
| 74 | TRAP1 | TRAP1 Entrez, Source | TNF receptor-associated protein 1 | 7108 | -0.014 | 0.2177 | No |
| 75 | FGF13 | FGF13 Entrez, Source | fibroblast growth factor 13 | 7222 | -0.016 | 0.2103 | No |
| 76 | TH | TH Entrez, Source | tyrosine hydroxylase | 7473 | -0.021 | 0.1929 | No |
| 77 | SMAD1 | SMAD1 Entrez, Source | SMAD, mothers against DPP homolog 1 (Drosophila) | 7536 | -0.022 | 0.1898 | No |
| 78 | TRIB1 | TRIB1 Entrez, Source | tribbles homolog 1 (Drosophila) | 7632 | -0.024 | 0.1843 | No |
| 79 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 7972 | -0.031 | 0.1608 | No |
| 80 | GPM6B | GPM6B Entrez, Source | glycoprotein M6B | 8060 | -0.032 | 0.1565 | No |
| 81 | PTPRU | PTPRU Entrez, Source | protein tyrosine phosphatase, receptor type, U | 8180 | -0.035 | 0.1500 | No |
| 82 | GNL1 | GNL1 Entrez, Source | guanine nucleotide binding protein-like 1 | 8250 | -0.036 | 0.1473 | No |
| 83 | FGF6 | FGF6 Entrez, Source | fibroblast growth factor 6 | 8436 | -0.040 | 0.1362 | No |
| 84 | SLC31A1 | SLC31A1 Entrez, Source | solute carrier family 31 (copper transporters), member 1 | 8681 | -0.045 | 0.1209 | No |
| 85 | HOXC10 | HOXC10 Entrez, Source | homeobox C10 | 8695 | -0.045 | 0.1231 | No |
| 86 | LCMT1 | LCMT1 Entrez, Source | leucine carboxyl methyltransferase 1 | 8783 | -0.047 | 0.1198 | No |
| 87 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 8961 | -0.051 | 0.1100 | No |
| 88 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 8985 | -0.051 | 0.1119 | No |
| 89 | AHI1 | AHI1 Entrez, Source | Abelson helper integration site 1 | 9218 | -0.056 | 0.0983 | No |
| 90 | THRAP3 | THRAP3 Entrez, Source | thyroid hormone receptor associated protein 3 | 9349 | -0.059 | 0.0926 | No |
| 91 | PPM1A | PPM1A Entrez, Source | protein phosphatase 1A (formerly 2C), magnesium-dependent, alpha isoform | 9447 | -0.062 | 0.0896 | No |
| 92 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 9594 | -0.065 | 0.0831 | No |
| 93 | UBAP1 | UBAP1 Entrez, Source | ubiquitin associated protein 1 | 9635 | -0.066 | 0.0847 | No |
| 94 | SLC18A2 | SLC18A2 Entrez, Source | solute carrier family 18 (vesicular monoamine), member 2 | 9786 | -0.070 | 0.0783 | No |
| 95 | EEF2 | EEF2 Entrez, Source | eukaryotic translation elongation factor 2 | 9787 | -0.070 | 0.0833 | No |
| 96 | RET | RET Entrez, Source | ret proto-oncogene (multiple endocrine neoplasia and medullary thyroid carcinoma 1, Hirschsprung disease) | 9808 | -0.071 | 0.0867 | No |
| 97 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 9896 | -0.074 | 0.0853 | No |
| 98 | RCE1 | RCE1 Entrez, Source | RCE1 homolog, prenyl protein peptidase (S. cerevisiae) | 10417 | -0.088 | 0.0522 | No |
| 99 | VGF | VGF Entrez, Source | VGF nerve growth factor inducible | 10474 | -0.089 | 0.0542 | No |
| 100 | RING1 | RING1 Entrez, Source | ring finger protein 1 | 10502 | -0.090 | 0.0585 | No |
| 101 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 10565 | -0.092 | 0.0603 | No |
| 102 | PRR3 | PRR3 Entrez, Source | proline rich 3 | 10589 | -0.093 | 0.0650 | No |
| 103 | STAT3 | STAT3 Entrez, Source | signal transducer and activator of transcription 3 (acute-phase response factor) | 10664 | -0.096 | 0.0662 | No |
| 104 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 10702 | -0.097 | 0.0702 | No |
| 105 | HHEX | HHEX Entrez, Source | homeobox, hematopoietically expressed | 10765 | -0.099 | 0.0725 | No |
| 106 | PPP1R10 | PPP1R10 Entrez, Source | protein phosphatase 1, regulatory subunit 10 | 10924 | -0.106 | 0.0679 | No |
| 107 | GAK | GAK Entrez, Source | cyclin G associated kinase | 11095 | -0.112 | 0.0629 | No |
| 108 | MRPS18B | MRPS18B Entrez, Source | mitochondrial ribosomal protein S18B | 11505 | -0.129 | 0.0411 | No |
| 109 | PPIG | PPIG Entrez, Source | peptidylprolyl isomerase G (cyclophilin G) | 11519 | -0.130 | 0.0492 | No |
| 110 | SENP2 | SENP2 Entrez, Source | SUMO1/sentrin/SMT3 specific peptidase 2 | 11664 | -0.136 | 0.0479 | No |
| 111 | YME1L1 | YME1L1 Entrez, Source | YME1-like 1 (S. cerevisiae) | 11766 | -0.142 | 0.0502 | No |
| 112 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 11805 | -0.145 | 0.0575 | No |
| 113 | MSX2 | MSX2 Entrez, Source | msh homeobox homolog 2 (Drosophila) | 11856 | -0.147 | 0.0641 | No |
| 114 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 12585 | -0.209 | 0.0237 | No |
| 115 | BIN1 | BIN1 Entrez, Source | bridging integrator 1 | 12736 | -0.231 | 0.0285 | No |
| 116 | GTF2A1 | GTF2A1 Entrez, Source | general transcription factor IIA, 1, 19/37kDa | 12822 | -0.246 | 0.0393 | No |