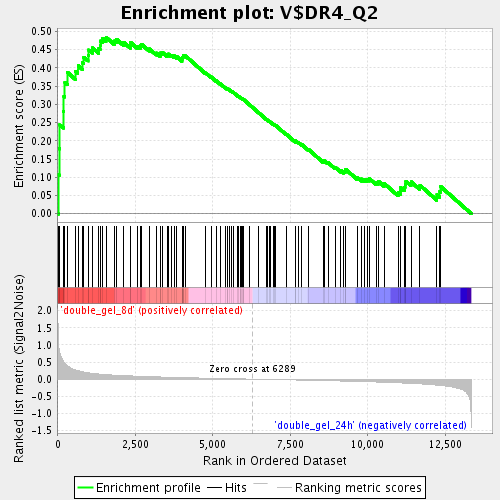
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$DR4_Q2 |
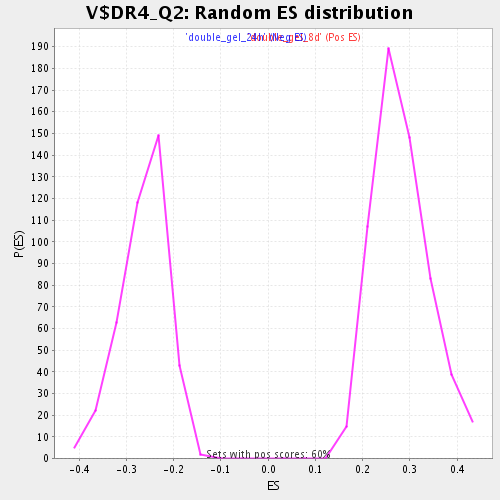
| Enrichment Score (ES) | 0.48413777 |
| Normalized Enrichment Score (NES) | 1.7189989 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.049597975 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 12 | 1.254 | 0.1081 | Yes |
| 2 | ARHGAP24 | ARHGAP24 Entrez, Source | Rho GTPase activating protein 24 | 44 | 0.838 | 0.1787 | Yes |
| 3 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 60 | 0.762 | 0.2438 | Yes |
| 4 | ABCG1 | ABCG1 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 1 | 179 | 0.519 | 0.2800 | Yes |
| 5 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 191 | 0.498 | 0.3224 | Yes |
| 6 | PTK2B | PTK2B Entrez, Source | PTK2B protein tyrosine kinase 2 beta | 224 | 0.463 | 0.3603 | Yes |
| 7 | IHPK2 | IHPK2 Entrez, Source | inositol hexaphosphate kinase 2 | 313 | 0.388 | 0.3874 | Yes |
| 8 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 565 | 0.265 | 0.3914 | Yes |
| 9 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 648 | 0.245 | 0.4066 | Yes |
| 10 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 796 | 0.215 | 0.4141 | Yes |
| 11 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 833 | 0.207 | 0.4294 | Yes |
| 12 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 973 | 0.186 | 0.4351 | Yes |
| 13 | OPTN | OPTN Entrez, Source | optineurin | 997 | 0.183 | 0.4493 | Yes |
| 14 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 1115 | 0.169 | 0.4551 | Yes |
| 15 | ABCA1 | ABCA1 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 1 | 1319 | 0.149 | 0.4527 | Yes |
| 16 | RASL11B | RASL11B Entrez, Source | RAS-like, family 11, member B | 1364 | 0.146 | 0.4621 | Yes |
| 17 | TNFRSF12A | TNFRSF12A Entrez, Source | tumor necrosis factor receptor superfamily, member 12A | 1374 | 0.145 | 0.4740 | Yes |
| 18 | TGOLN2 | TGOLN2 Entrez, Source | trans-golgi network protein 2 | 1434 | 0.140 | 0.4817 | Yes |
| 19 | NDUFS2 | NDUFS2 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 2, 49kDa (NADH-coenzyme Q reductase) | 1554 | 0.131 | 0.4841 | Yes |
| 20 | LRP3 | LRP3 Entrez, Source | low density lipoprotein receptor-related protein 3 | 1822 | 0.116 | 0.4741 | No |
| 21 | ENO2 | ENO2 Entrez, Source | enolase 2 (gamma, neuronal) | 1889 | 0.112 | 0.4788 | No |
| 22 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 2116 | 0.103 | 0.4707 | No |
| 23 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 2337 | 0.094 | 0.4623 | No |
| 24 | NRG1 | NRG1 Entrez, Source | neuregulin 1 | 2347 | 0.094 | 0.4697 | No |
| 25 | OBSCN | OBSCN Entrez, Source | obscurin, cytoskeletal calmodulin and titin-interacting RhoGEF | 2576 | 0.084 | 0.4598 | No |
| 26 | APLP2 | APLP2 Entrez, Source | amyloid beta (A4) precursor-like protein 2 | 2657 | 0.082 | 0.4609 | No |
| 27 | CX36 | CX36 Entrez, Source | - | 2695 | 0.080 | 0.4651 | No |
| 28 | ACACA | ACACA Entrez, Source | acetyl-Coenzyme A carboxylase alpha | 2948 | 0.073 | 0.4524 | No |
| 29 | ACCN3 | ACCN3 Entrez, Source | amiloride-sensitive cation channel 3 | 3175 | 0.066 | 0.4411 | No |
| 30 | SNIP | SNIP Entrez, Source | - | 3301 | 0.063 | 0.4371 | No |
| 31 | COL4A4 | COL4A4 Entrez, Source | collagen, type IV, alpha 4 | 3320 | 0.062 | 0.4411 | No |
| 32 | MYH13 | MYH13 Entrez, Source | myosin, heavy chain 13, skeletal muscle | 3383 | 0.061 | 0.4417 | No |
| 33 | NR5A2 | NR5A2 Entrez, Source | nuclear receptor subfamily 5, group A, member 2 | 3528 | 0.057 | 0.4358 | No |
| 34 | SLC7A9 | SLC7A9 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 9 | 3559 | 0.056 | 0.4383 | No |
| 35 | NOVA1 | NOVA1 Entrez, Source | neuro-oncological ventral antigen 1 | 3664 | 0.053 | 0.4351 | No |
| 36 | GREM1 | GREM1 Entrez, Source | gremlin 1, cysteine knot superfamily, homolog (Xenopus laevis) | 3749 | 0.051 | 0.4332 | No |
| 37 | TEAD3 | TEAD3 Entrez, Source | TEA domain family member 3 | 3839 | 0.049 | 0.4308 | No |
| 38 | ERBB3 | ERBB3 Entrez, Source | v-erb-b2 erythroblastic leukemia viral oncogene homolog 3 (avian) | 4005 | 0.044 | 0.4222 | No |
| 39 | ATP5G3 | ATP5G3 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C3 (subunit 9) | 4011 | 0.044 | 0.4256 | No |
| 40 | APOC1 | APOC1 Entrez, Source | apolipoprotein C-I | 4022 | 0.044 | 0.4287 | No |
| 41 | MAP1LC3A | MAP1LC3A Entrez, Source | microtubule-associated protein 1 light chain 3 alpha | 4039 | 0.044 | 0.4313 | No |
| 42 | NPEPPS | NPEPPS Entrez, Source | aminopeptidase puromycin sensitive | 4043 | 0.044 | 0.4348 | No |
| 43 | TGFB3 | TGFB3 Entrez, Source | transforming growth factor, beta 3 | 4113 | 0.042 | 0.4333 | No |
| 44 | NDUFS4 | NDUFS4 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 4, 18kDa (NADH-coenzyme Q reductase) | 4767 | 0.029 | 0.3864 | No |
| 45 | ATP1A2 | ATP1A2 Entrez, Source | ATPase, Na+/K+ transporting, alpha 2 (+) polypeptide | 4943 | 0.025 | 0.3754 | No |
| 46 | GGPS1 | GGPS1 Entrez, Source | geranylgeranyl diphosphate synthase 1 | 5108 | 0.022 | 0.3649 | No |
| 47 | DLX1 | DLX1 Entrez, Source | distal-less homeobox 1 | 5243 | 0.020 | 0.3565 | No |
| 48 | AMHR2 | AMHR2 Entrez, Source | anti-Mullerian hormone receptor, type II | 5408 | 0.017 | 0.3456 | No |
| 49 | NLGN3 | NLGN3 Entrez, Source | neuroligin 3 | 5467 | 0.016 | 0.3425 | No |
| 50 | NEUROG1 | NEUROG1 Entrez, Source | neurogenin 1 | 5488 | 0.015 | 0.3423 | No |
| 51 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 5532 | 0.014 | 0.3403 | No |
| 52 | ATP5G1 | ATP5G1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C1 (subunit 9) | 5604 | 0.013 | 0.3361 | No |
| 53 | SNF1LK | SNF1LK Entrez, Source | SNF1-like kinase | 5650 | 0.012 | 0.3337 | No |
| 54 | GNB2 | GNB2 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 2 | 5801 | 0.009 | 0.3231 | No |
| 55 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 5820 | 0.009 | 0.3225 | No |
| 56 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 5876 | 0.008 | 0.3190 | No |
| 57 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 5892 | 0.007 | 0.3185 | No |
| 58 | UQCRC1 | UQCRC1 Entrez, Source | ubiquinol-cytochrome c reductase core protein I | 5927 | 0.007 | 0.3165 | No |
| 59 | FGF11 | FGF11 Entrez, Source | fibroblast growth factor 11 | 5965 | 0.006 | 0.3143 | No |
| 60 | PEX11B | PEX11B Entrez, Source | peroxisomal biogenesis factor 11B | 5988 | 0.005 | 0.3131 | No |
| 61 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 5997 | 0.005 | 0.3129 | No |
| 62 | DCX | DCX Entrez, Source | doublecortex; lissencephaly, X-linked (doublecortin) | 6186 | 0.001 | 0.2988 | No |
| 63 | NELL2 | NELL2 Entrez, Source | NEL-like 2 (chicken) | 6480 | -0.003 | 0.2770 | No |
| 64 | MYH6 | MYH6 Entrez, Source | myosin, heavy chain 6, cardiac muscle, alpha (cardiomyopathy, hypertrophic 1) | 6716 | -0.007 | 0.2599 | No |
| 65 | NNAT | NNAT Entrez, Source | neuronatin | 6767 | -0.008 | 0.2568 | No |
| 66 | HRH3 | HRH3 Entrez, Source | histamine receptor H3 | 6821 | -0.009 | 0.2536 | No |
| 67 | ACY1 | ACY1 Entrez, Source | aminoacylase 1 | 6868 | -0.010 | 0.2510 | No |
| 68 | SEMA6C | SEMA6C Entrez, Source | sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6C | 6955 | -0.011 | 0.2455 | No |
| 69 | NXPH4 | NXPH4 Entrez, Source | neurexophilin 4 | 7004 | -0.012 | 0.2430 | No |
| 70 | GART | GART Entrez, Source | phosphoribosylglycinamide formyltransferase, phosphoribosylglycinamide synthetase, phosphoribosylaminoimidazole synthetase | 7019 | -0.013 | 0.2430 | No |
| 71 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 7366 | -0.019 | 0.2185 | No |
| 72 | GDNF | GDNF Entrez, Source | glial cell derived neurotrophic factor | 7663 | -0.025 | 0.1983 | No |
| 73 | SCAMP3 | SCAMP3 Entrez, Source | secretory carrier membrane protein 3 | 7665 | -0.025 | 0.2004 | No |
| 74 | PCSK1N | PCSK1N Entrez, Source | proprotein convertase subtilisin/kexin type 1 inhibitor | 7755 | -0.026 | 0.1960 | No |
| 75 | KCND3 | KCND3 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 3 | 7872 | -0.029 | 0.1897 | No |
| 76 | ESRRA | ESRRA Entrez, Source | estrogen-related receptor alpha | 8092 | -0.033 | 0.1761 | No |
| 77 | CAMKK2 | CAMKK2 Entrez, Source | calcium/calmodulin-dependent protein kinase kinase 2, beta | 8568 | -0.043 | 0.1439 | No |
| 78 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 8595 | -0.043 | 0.1457 | No |
| 79 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 8718 | -0.046 | 0.1405 | No |
| 80 | TFEB | TFEB Entrez, Source | transcription factor EB | 8952 | -0.051 | 0.1273 | No |
| 81 | RAB10 | RAB10 Entrez, Source | RAB10, member RAS oncogene family | 9126 | -0.055 | 0.1190 | No |
| 82 | SON | SON Entrez, Source | SON DNA binding protein | 9229 | -0.057 | 0.1162 | No |
| 83 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 9270 | -0.057 | 0.1182 | No |
| 84 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 9277 | -0.058 | 0.1227 | No |
| 85 | BTBD14B | BTBD14B Entrez, Source | BTB (POZ) domain containing 14B | 9670 | -0.067 | 0.0989 | No |
| 86 | UBE2D3 | UBE2D3 Entrez, Source | ubiquitin-conjugating enzyme E2D 3 (UBC4/5 homolog, yeast) | 9797 | -0.071 | 0.0956 | No |
| 87 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 9896 | -0.074 | 0.0946 | No |
| 88 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 9978 | -0.076 | 0.0950 | No |
| 89 | TIMM10 | TIMM10 Entrez, Source | translocase of inner mitochondrial membrane 10 homolog (yeast) | 10050 | -0.078 | 0.0965 | No |
| 90 | PUM2 | PUM2 Entrez, Source | pumilio homolog 2 (Drosophila) | 10272 | -0.084 | 0.0870 | No |
| 91 | IL11 | IL11 Entrez, Source | interleukin 11 | 10348 | -0.086 | 0.0888 | No |
| 92 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 10528 | -0.091 | 0.0832 | No |
| 93 | EIF4G1 | EIF4G1 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 1 | 10984 | -0.108 | 0.0582 | No |
| 94 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 11050 | -0.110 | 0.0629 | No |
| 95 | ADM | ADM Entrez, Source | adrenomedullin | 11063 | -0.110 | 0.0716 | No |
| 96 | PDGFRL | PDGFRL Entrez, Source | platelet-derived growth factor receptor-like | 11182 | -0.116 | 0.0727 | No |
| 97 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 11205 | -0.116 | 0.0811 | No |
| 98 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 11229 | -0.117 | 0.0896 | No |
| 99 | POLR3C | POLR3C Entrez, Source | polymerase (RNA) III (DNA directed) polypeptide C (62kD) | 11398 | -0.125 | 0.0877 | No |
| 100 | BZW1 | BZW1 Entrez, Source | basic leucine zipper and W2 domains 1 | 11682 | -0.137 | 0.0783 | No |
| 101 | ARID4B | ARID4B Entrez, Source | AT rich interactive domain 4B (RBP1- like) | 12232 | -0.174 | 0.0519 | No |
| 102 | ADNP | ADNP Entrez, Source | activity-dependent neuroprotector | 12323 | -0.181 | 0.0609 | No |
| 103 | NR1D1 | NR1D1 Entrez, Source | nuclear receptor subfamily 1, group D, member 1 | 12353 | -0.184 | 0.0747 | No |