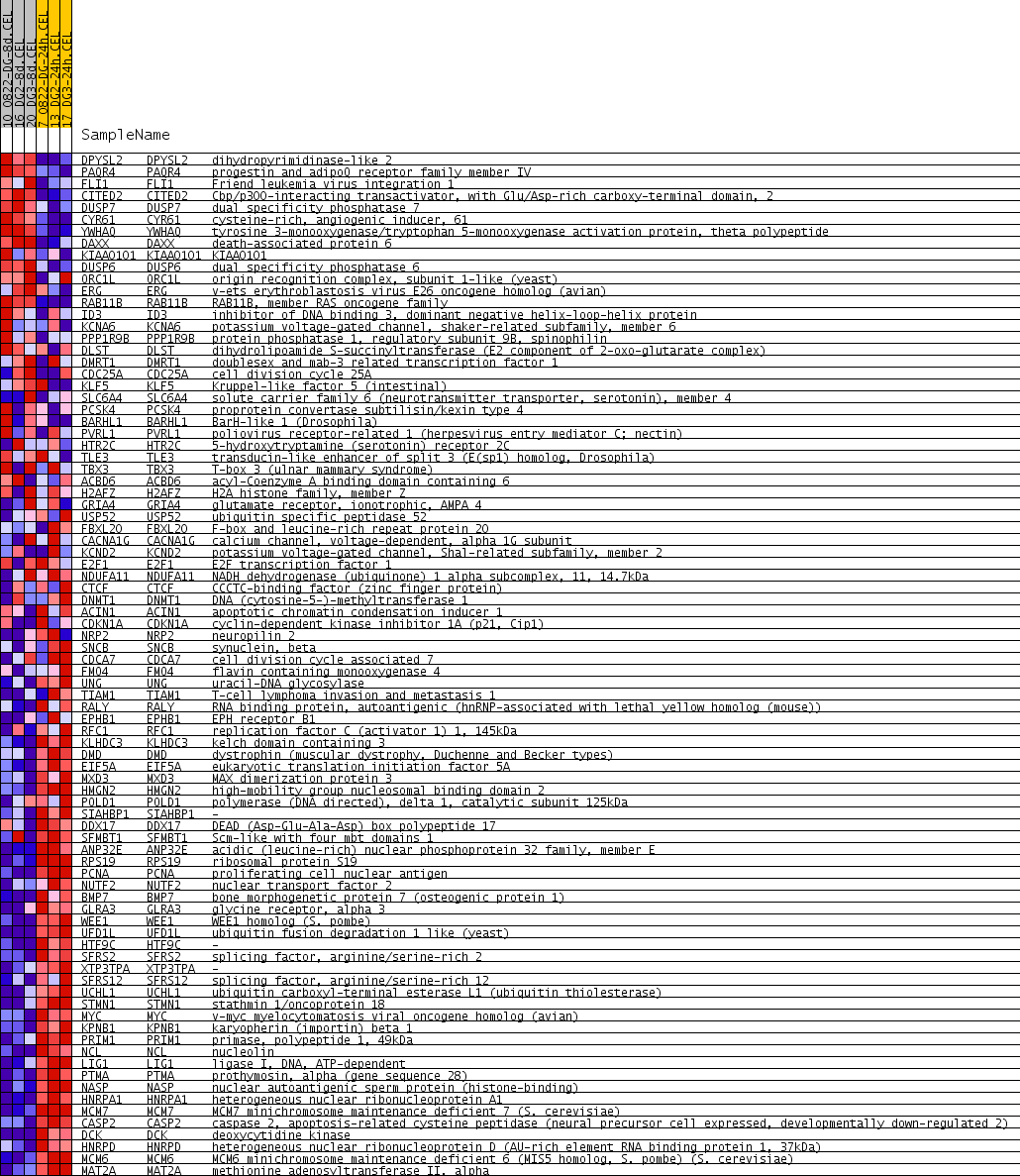
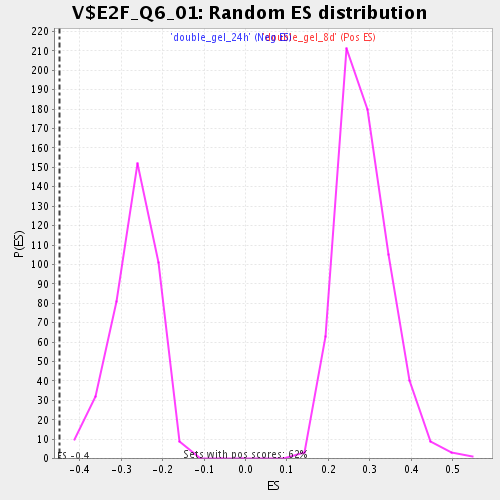
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | V$E2F_Q6_01 |
| Enrichment Score (ES) | -0.4490558 |
| Normalized Enrichment Score (NES) | -1.6777145 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.110000364 |
| FWER p-Value | 0.999 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 158 | 0.552 | 0.0367 | No |
| 2 | PAQR4 | PAQR4 Entrez, Source | progestin and adipoQ receptor family member IV | 293 | 0.407 | 0.0625 | No |
| 3 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 724 | 0.231 | 0.0505 | No |
| 4 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 793 | 0.215 | 0.0643 | No |
| 5 | DUSP7 | DUSP7 Entrez, Source | dual specificity phosphatase 7 | 970 | 0.186 | 0.0675 | No |
| 6 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 1062 | 0.175 | 0.0760 | No |
| 7 | YWHAQ | YWHAQ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide | 1447 | 0.139 | 0.0593 | No |
| 8 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 1796 | 0.117 | 0.0434 | No |
| 9 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 2376 | 0.093 | 0.0079 | No |
| 10 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 2490 | 0.088 | 0.0071 | No |
| 11 | ORC1L | ORC1L Entrez, Source | origin recognition complex, subunit 1-like (yeast) | 2867 | 0.075 | -0.0146 | No |
| 12 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 3040 | 0.070 | -0.0214 | No |
| 13 | RAB11B | RAB11B Entrez, Source | RAB11B, member RAS oncogene family | 3071 | 0.069 | -0.0176 | No |
| 14 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 3479 | 0.058 | -0.0431 | No |
| 15 | KCNA6 | KCNA6 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 6 | 3580 | 0.055 | -0.0458 | No |
| 16 | PPP1R9B | PPP1R9B Entrez, Source | protein phosphatase 1, regulatory subunit 9B, spinophilin | 3862 | 0.048 | -0.0627 | No |
| 17 | DLST | DLST Entrez, Source | dihydrolipoamide S-succinyltransferase (E2 component of 2-oxo-glutarate complex) | 3905 | 0.047 | -0.0617 | No |
| 18 | DMRT1 | DMRT1 Entrez, Source | doublesex and mab-3 related transcription factor 1 | 3981 | 0.045 | -0.0634 | No |
| 19 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 4102 | 0.042 | -0.0688 | No |
| 20 | KLF5 | KLF5 Entrez, Source | Kruppel-like factor 5 (intestinal) | 4503 | 0.034 | -0.0959 | No |
| 21 | SLC6A4 | SLC6A4 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 | 4511 | 0.034 | -0.0935 | No |
| 22 | PCSK4 | PCSK4 Entrez, Source | proprotein convertase subtilisin/kexin type 4 | 4552 | 0.033 | -0.0936 | No |
| 23 | BARHL1 | BARHL1 Entrez, Source | BarH-like 1 (Drosophila) | 4685 | 0.030 | -0.1009 | No |
| 24 | PVRL1 | PVRL1 Entrez, Source | poliovirus receptor-related 1 (herpesvirus entry mediator C; nectin) | 5060 | 0.023 | -0.1271 | No |
| 25 | HTR2C | HTR2C Entrez, Source | 5-hydroxytryptamine (serotonin) receptor 2C | 5202 | 0.020 | -0.1359 | No |
| 26 | TLE3 | TLE3 Entrez, Source | transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) | 5643 | 0.012 | -0.1681 | No |
| 27 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 5869 | 0.008 | -0.1844 | No |
| 28 | ACBD6 | ACBD6 Entrez, Source | acyl-Coenzyme A binding domain containing 6 | 6149 | 0.002 | -0.2052 | No |
| 29 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 6380 | -0.001 | -0.2224 | No |
| 30 | GRIA4 | GRIA4 Entrez, Source | glutamate receptor, ionotrophic, AMPA 4 | 6422 | -0.002 | -0.2253 | No |
| 31 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 7291 | -0.018 | -0.2893 | No |
| 32 | FBXL20 | FBXL20 Entrez, Source | F-box and leucine-rich repeat protein 20 | 7404 | -0.020 | -0.2960 | No |
| 33 | CACNA1G | CACNA1G Entrez, Source | calcium channel, voltage-dependent, alpha 1G subunit | 7509 | -0.022 | -0.3019 | No |
| 34 | KCND2 | KCND2 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 2 | 7584 | -0.023 | -0.3054 | No |
| 35 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 7606 | -0.024 | -0.3049 | No |
| 36 | NDUFA11 | NDUFA11 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 11, 14.7kDa | 7683 | -0.025 | -0.3085 | No |
| 37 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 8374 | -0.039 | -0.3571 | No |
| 38 | DNMT1 | DNMT1 Entrez, Source | DNA (cytosine-5-)-methyltransferase 1 | 8474 | -0.041 | -0.3609 | No |
| 39 | ACIN1 | ACIN1 Entrez, Source | apoptotic chromatin condensation inducer 1 | 8577 | -0.043 | -0.3648 | No |
| 40 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 8685 | -0.045 | -0.3689 | No |
| 41 | NRP2 | NRP2 Entrez, Source | neuropilin 2 | 8752 | -0.047 | -0.3698 | No |
| 42 | SNCB | SNCB Entrez, Source | synuclein, beta | 8772 | -0.047 | -0.3671 | No |
| 43 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 9009 | -0.052 | -0.3803 | No |
| 44 | FMO4 | FMO4 Entrez, Source | flavin containing monooxygenase 4 | 9249 | -0.057 | -0.3933 | No |
| 45 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 9382 | -0.060 | -0.3979 | No |
| 46 | TIAM1 | TIAM1 Entrez, Source | T-cell lymphoma invasion and metastasis 1 | 9463 | -0.062 | -0.3985 | No |
| 47 | RALY | RALY Entrez, Source | RNA binding protein, autoantigenic (hnRNP-associated with lethal yellow homolog (mouse)) | 9531 | -0.064 | -0.3980 | No |
| 48 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 9688 | -0.068 | -0.4038 | No |
| 49 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 9713 | -0.068 | -0.3996 | No |
| 50 | KLHDC3 | KLHDC3 Entrez, Source | kelch domain containing 3 | 9714 | -0.068 | -0.3935 | No |
| 51 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 9896 | -0.074 | -0.4007 | No |
| 52 | EIF5A | EIF5A Entrez, Source | eukaryotic translation initiation factor 5A | 10178 | -0.081 | -0.4147 | No |
| 53 | MXD3 | MXD3 Entrez, Source | MAX dimerization protein 3 | 10205 | -0.082 | -0.4095 | No |
| 54 | HMGN2 | HMGN2 Entrez, Source | high-mobility group nucleosomal binding domain 2 | 10359 | -0.086 | -0.4134 | No |
| 55 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10438 | -0.088 | -0.4115 | No |
| 56 | SIAHBP1 | SIAHBP1 Entrez, Source | - | 10678 | -0.096 | -0.4211 | No |
| 57 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 11050 | -0.110 | -0.4394 | Yes |
| 58 | SFMBT1 | SFMBT1 Entrez, Source | Scm-like with four mbt domains 1 | 11109 | -0.112 | -0.4338 | Yes |
| 59 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 11114 | -0.113 | -0.4242 | Yes |
| 60 | RPS19 | RPS19 Entrez, Source | ribosomal protein S19 | 11310 | -0.121 | -0.4282 | Yes |
| 61 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 11449 | -0.127 | -0.4275 | Yes |
| 62 | NUTF2 | NUTF2 Entrez, Source | nuclear transport factor 2 | 11453 | -0.127 | -0.4165 | Yes |
| 63 | BMP7 | BMP7 Entrez, Source | bone morphogenetic protein 7 (osteogenic protein 1) | 11644 | -0.136 | -0.4188 | Yes |
| 64 | GLRA3 | GLRA3 Entrez, Source | glycine receptor, alpha 3 | 11725 | -0.140 | -0.4126 | Yes |
| 65 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 11948 | -0.154 | -0.4158 | Yes |
| 66 | UFD1L | UFD1L Entrez, Source | ubiquitin fusion degradation 1 like (yeast) | 11964 | -0.154 | -0.4033 | Yes |
| 67 | HTF9C | HTF9C Entrez, Source | - | 11970 | -0.155 | -0.3900 | Yes |
| 68 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 12039 | -0.160 | -0.3810 | Yes |
| 69 | XTP3TPA | XTP3TPA Entrez, Source | - | 12079 | -0.163 | -0.3696 | Yes |
| 70 | SFRS12 | SFRS12 Entrez, Source | splicing factor, arginine/serine-rich 12 | 12159 | -0.169 | -0.3607 | Yes |
| 71 | UCHL1 | UCHL1 Entrez, Source | ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | 12209 | -0.172 | -0.3492 | Yes |
| 72 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 12244 | -0.175 | -0.3363 | Yes |
| 73 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 12249 | -0.175 | -0.3211 | Yes |
| 74 | KPNB1 | KPNB1 Entrez, Source | karyopherin (importin) beta 1 | 12477 | -0.196 | -0.3210 | Yes |
| 75 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 12830 | -0.249 | -0.3256 | Yes |
| 76 | NCL | NCL Entrez, Source | nucleolin | 12857 | -0.256 | -0.3050 | Yes |
| 77 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 12917 | -0.270 | -0.2856 | Yes |
| 78 | PTMA | PTMA Entrez, Source | prothymosin, alpha (gene sequence 28) | 12945 | -0.275 | -0.2634 | Yes |
| 79 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 12994 | -0.291 | -0.2413 | Yes |
| 80 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 13080 | -0.318 | -0.2197 | Yes |
| 81 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13101 | -0.328 | -0.1923 | Yes |
| 82 | CASP2 | CASP2 Entrez, Source | caspase 2, apoptosis-related cysteine peptidase (neural precursor cell expressed, developmentally down-regulated 2) | 13151 | -0.360 | -0.1642 | Yes |
| 83 | DCK | DCK Entrez, Source | deoxycytidine kinase | 13222 | -0.428 | -0.1318 | Yes |
| 84 | HNRPD | HNRPD Entrez, Source | heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) | 13227 | -0.440 | -0.0933 | Yes |
| 85 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13288 | -0.553 | -0.0490 | Yes |
| 86 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 13299 | -0.601 | 0.0032 | Yes |