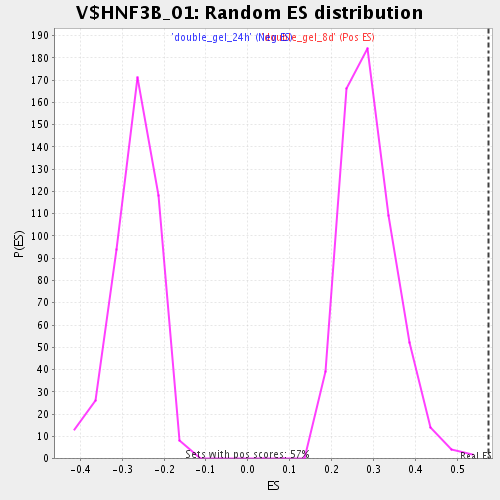
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$HNF3B_01 |
| Enrichment Score (ES) | 0.5735075 |
| Normalized Enrichment Score (NES) | 1.981983 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0067341975 |
| FWER p-Value | 0.204 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 25 | 0.995 | 0.0924 | Yes |
| 2 | WNT4 | WNT4 Entrez, Source | wingless-type MMTV integration site family, member 4 | 42 | 0.852 | 0.1718 | Yes |
| 3 | GPM6A | GPM6A Entrez, Source | glycoprotein M6A | 165 | 0.540 | 0.2138 | Yes |
| 4 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 312 | 0.390 | 0.2397 | Yes |
| 5 | RBP4 | RBP4 Entrez, Source | retinol binding protein 4, plasma | 323 | 0.378 | 0.2748 | Yes |
| 6 | PLA1A | PLA1A Entrez, Source | phospholipase A1 member A | 435 | 0.317 | 0.2964 | Yes |
| 7 | SERPIND1 | SERPIND1 Entrez, Source | serpin peptidase inhibitor, clade D (heparin cofactor), member 1 | 504 | 0.284 | 0.3182 | Yes |
| 8 | FUT8 | FUT8 Entrez, Source | fucosyltransferase 8 (alpha (1,6) fucosyltransferase) | 567 | 0.265 | 0.3386 | Yes |
| 9 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 597 | 0.258 | 0.3608 | Yes |
| 10 | ENPEP | ENPEP Entrez, Source | glutamyl aminopeptidase (aminopeptidase A) | 658 | 0.244 | 0.3793 | Yes |
| 11 | BAIAP2 | BAIAP2 Entrez, Source | BAI1-associated protein 2 | 768 | 0.222 | 0.3921 | Yes |
| 12 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 793 | 0.215 | 0.4107 | Yes |
| 13 | TTR | TTR Entrez, Source | transthyretin (prealbumin, amyloidosis type I) | 798 | 0.213 | 0.4306 | Yes |
| 14 | NCDN | NCDN Entrez, Source | neurochondrin | 809 | 0.212 | 0.4499 | Yes |
| 15 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 824 | 0.209 | 0.4686 | Yes |
| 16 | SULF2 | SULF2 Entrez, Source | sulfatase 2 | 885 | 0.197 | 0.4828 | Yes |
| 17 | IER3 | IER3 Entrez, Source | immediate early response 3 | 908 | 0.195 | 0.4996 | Yes |
| 18 | GFRA1 | GFRA1 Entrez, Source | GDNF family receptor alpha 1 | 947 | 0.188 | 0.5146 | Yes |
| 19 | RAB3IP | RAB3IP Entrez, Source | RAB3A interacting protein (rabin3) | 964 | 0.187 | 0.5310 | Yes |
| 20 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 1062 | 0.175 | 0.5403 | Yes |
| 21 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1066 | 0.174 | 0.5566 | Yes |
| 22 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 1115 | 0.169 | 0.5690 | Yes |
| 23 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 1266 | 0.154 | 0.5723 | Yes |
| 24 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 1427 | 0.140 | 0.5735 | Yes |
| 25 | ATP2A3 | ATP2A3 Entrez, Source | ATPase, Ca++ transporting, ubiquitous | 1837 | 0.115 | 0.5536 | No |
| 26 | LCP2 | LCP2 Entrez, Source | lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa) | 1877 | 0.113 | 0.5613 | No |
| 27 | TNMD | TNMD Entrez, Source | tenomodulin | 2550 | 0.086 | 0.5187 | No |
| 28 | FOXA2 | FOXA2 Entrez, Source | forkhead box A2 | 2583 | 0.084 | 0.5243 | No |
| 29 | ELMO3 | ELMO3 Entrez, Source | engulfment and cell motility 3 | 2651 | 0.082 | 0.5270 | No |
| 30 | TWIST2 | TWIST2 Entrez, Source | twist homolog 2 (Drosophila) | 2757 | 0.078 | 0.5264 | No |
| 31 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 2831 | 0.076 | 0.5282 | No |
| 32 | PDCD4 | PDCD4 Entrez, Source | programmed cell death 4 (neoplastic transformation inhibitor) | 3035 | 0.070 | 0.5195 | No |
| 33 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 3037 | 0.070 | 0.5261 | No |
| 34 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 3286 | 0.063 | 0.5133 | No |
| 35 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 3506 | 0.057 | 0.5022 | No |
| 36 | ATP1B4 | ATP1B4 Entrez, Source | ATPase, (Na+)/K+ transporting, beta 4 polypeptide | 3612 | 0.054 | 0.4995 | No |
| 37 | FLOT1 | FLOT1 Entrez, Source | flotillin 1 | 3693 | 0.052 | 0.4984 | No |
| 38 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 3725 | 0.052 | 0.5010 | No |
| 39 | POU3F4 | POU3F4 Entrez, Source | POU domain, class 3, transcription factor 4 | 3843 | 0.049 | 0.4968 | No |
| 40 | LHX5 | LHX5 Entrez, Source | LIM homeobox 5 | 3894 | 0.047 | 0.4975 | No |
| 41 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 4035 | 0.044 | 0.4911 | No |
| 42 | SSH3 | SSH3 Entrez, Source | slingshot homolog 3 (Drosophila) | 4110 | 0.042 | 0.4895 | No |
| 43 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 4246 | 0.039 | 0.4829 | No |
| 44 | SLC6A4 | SLC6A4 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, serotonin), member 4 | 4511 | 0.034 | 0.4662 | No |
| 45 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 4797 | 0.028 | 0.4474 | No |
| 46 | CYP26B1 | CYP26B1 Entrez, Source | cytochrome P450, family 26, subfamily B, polypeptide 1 | 4829 | 0.027 | 0.4476 | No |
| 47 | PVRL1 | PVRL1 Entrez, Source | poliovirus receptor-related 1 (herpesvirus entry mediator C; nectin) | 5060 | 0.023 | 0.4324 | No |
| 48 | CALD1 | CALD1 Entrez, Source | caldesmon 1 | 5081 | 0.023 | 0.4331 | No |
| 49 | IBSP | IBSP Entrez, Source | integrin-binding sialoprotein (bone sialoprotein, bone sialoprotein II) | 5170 | 0.021 | 0.4284 | No |
| 50 | PLCB1 | PLCB1 Entrez, Source | phospholipase C, beta 1 (phosphoinositide-specific) | 5171 | 0.021 | 0.4304 | No |
| 51 | OPRM1 | OPRM1 Entrez, Source | opioid receptor, mu 1 | 5387 | 0.017 | 0.4158 | No |
| 52 | MCART1 | MCART1 Entrez, Source | mitochondrial carrier triple repeat 1 | 5537 | 0.014 | 0.4058 | No |
| 53 | PHEX | PHEX Entrez, Source | phosphate regulating endopeptidase homolog, X-linked (hypophosphatemia, vitamin D resistant rickets) | 5619 | 0.012 | 0.4009 | No |
| 54 | SLC16A6 | SLC16A6 Entrez, Source | solute carrier family 16, member 6 (monocarboxylic acid transporter 7) | 6019 | 0.005 | 0.3713 | No |
| 55 | IL5 | IL5 Entrez, Source | interleukin 5 (colony-stimulating factor, eosinophil) | 6076 | 0.004 | 0.3674 | No |
| 56 | TNF | TNF Entrez, Source | tumor necrosis factor (TNF superfamily, member 2) | 6181 | 0.001 | 0.3597 | No |
| 57 | TAC1 | TAC1 Entrez, Source | tachykinin, precursor 1 (substance K, substance P, neurokinin 1, neurokinin 2, neuromedin L, neurokinin alpha, neuropeptide K, neuropeptide gamma) | 6693 | -0.007 | 0.3218 | No |
| 58 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 6785 | -0.009 | 0.3157 | No |
| 59 | PNMA1 | PNMA1 Entrez, Source | paraneoplastic antigen MA1 | 6990 | -0.012 | 0.3015 | No |
| 60 | DNAJA4 | DNAJA4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 4 | 7050 | -0.013 | 0.2983 | No |
| 61 | FGF13 | FGF13 Entrez, Source | fibroblast growth factor 13 | 7222 | -0.016 | 0.2870 | No |
| 62 | HAS2 | HAS2 Entrez, Source | hyaluronan synthase 2 | 7235 | -0.017 | 0.2876 | No |
| 63 | ITPKB | ITPKB Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase B | 7323 | -0.018 | 0.2828 | No |
| 64 | ORC4L | ORC4L Entrez, Source | origin recognition complex, subunit 4-like (yeast) | 7357 | -0.019 | 0.2821 | No |
| 65 | CACNA1H | CACNA1H Entrez, Source | calcium channel, voltage-dependent, alpha 1H subunit | 7393 | -0.020 | 0.2813 | No |
| 66 | NFATC4 | NFATC4 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 | 7625 | -0.024 | 0.2662 | No |
| 67 | GABRA1 | GABRA1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 1 | 7649 | -0.025 | 0.2667 | No |
| 68 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 7768 | -0.027 | 0.2604 | No |
| 69 | PPP1CB | PPP1CB Entrez, Source | protein phosphatase 1, catalytic subunit, beta isoform | 7860 | -0.028 | 0.2562 | No |
| 70 | LTA | LTA Entrez, Source | lymphotoxin alpha (TNF superfamily, member 1) | 8246 | -0.036 | 0.2306 | No |
| 71 | CHN2 | CHN2 Entrez, Source | chimerin (chimaerin) 2 | 8642 | -0.044 | 0.2050 | No |
| 72 | VAX1 | VAX1 Entrez, Source | ventral anterior homeobox 1 | 8915 | -0.050 | 0.1892 | No |
| 73 | NEUROD2 | NEUROD2 Entrez, Source | neurogenic differentiation 2 | 9017 | -0.052 | 0.1866 | No |
| 74 | HOXA4 | HOXA4 Entrez, Source | homeobox A4 | 9080 | -0.054 | 0.1870 | No |
| 75 | EPHB1 | EPHB1 Entrez, Source | EPH receptor B1 | 9688 | -0.068 | 0.1476 | No |
| 76 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 9896 | -0.074 | 0.1389 | No |
| 77 | ABT1 | ABT1 Entrez, Source | activator of basal transcription 1 | 10127 | -0.080 | 0.1291 | No |
| 78 | ACCN5 | ACCN5 Entrez, Source | amiloride-sensitive cation channel 5, intestinal | 10267 | -0.083 | 0.1266 | No |
| 79 | DRD3 | DRD3 Entrez, Source | dopamine receptor D3 | 10311 | -0.085 | 0.1314 | No |
| 80 | PPP1R10 | PPP1R10 Entrez, Source | protein phosphatase 1, regulatory subunit 10 | 10924 | -0.106 | 0.0952 | No |
| 81 | CYLN2 | CYLN2 Entrez, Source | cytoplasmic linker 2 | 11077 | -0.111 | 0.0943 | No |
| 82 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 11229 | -0.117 | 0.0940 | No |
| 83 | MRPS18B | MRPS18B Entrez, Source | mitochondrial ribosomal protein S18B | 11505 | -0.129 | 0.0854 | No |
| 84 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 11943 | -0.153 | 0.0670 | No |
| 85 | NFIX | NFIX Entrez, Source | nuclear factor I/X (CCAAT-binding transcription factor) | 12029 | -0.159 | 0.0756 | No |
| 86 | GTF2A1 | GTF2A1 Entrez, Source | general transcription factor IIA, 1, 19/37kDa | 12822 | -0.246 | 0.0392 | No |