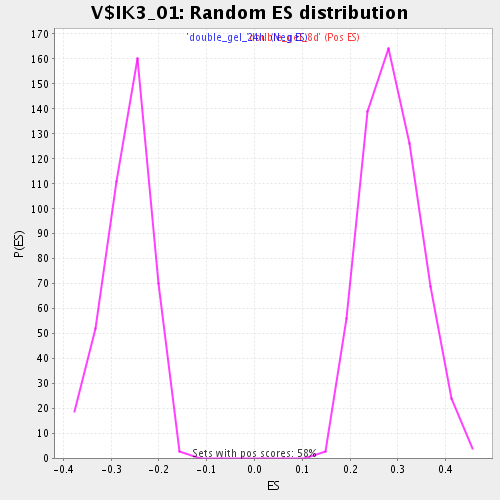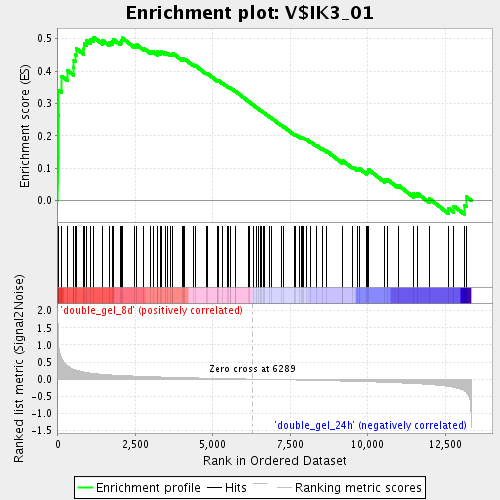
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$IK3_01 |
| Enrichment Score (ES) | 0.5044467 |
| Normalized Enrichment Score (NES) | 1.761194 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.037264533 |
| FWER p-Value | 0.99 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CYP17A1 | CYP17A1 Entrez, Source | cytochrome P450, family 17, subfamily A, polypeptide 1 | 2 | 1.623 | 0.1323 | Yes |
| 2 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 3 | 1.622 | 0.2648 | Yes |
| 3 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 27 | 0.963 | 0.3416 | Yes |
| 4 | RGS5 | RGS5 Entrez, Source | regulator of G-protein signalling 5 | 125 | 0.616 | 0.3846 | Yes |
| 5 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 318 | 0.386 | 0.4016 | Yes |
| 6 | RHBDF1 | RHBDF1 Entrez, Source | rhomboid 5 homolog 1 (Drosophila) | 506 | 0.283 | 0.4106 | Yes |
| 7 | HCST | HCST Entrez, Source | hematopoietic cell signal transducer | 517 | 0.280 | 0.4327 | Yes |
| 8 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 565 | 0.265 | 0.4508 | Yes |
| 9 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 597 | 0.258 | 0.4695 | Yes |
| 10 | ADAM15 | ADAM15 Entrez, Source | ADAM metallopeptidase domain 15 (metargidin) | 837 | 0.207 | 0.4683 | Yes |
| 11 | MKNK2 | MKNK2 Entrez, Source | MAP kinase interacting serine/threonine kinase 2 | 842 | 0.206 | 0.4848 | Yes |
| 12 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 932 | 0.190 | 0.4936 | Yes |
| 13 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 1062 | 0.175 | 0.4981 | Yes |
| 14 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 1157 | 0.164 | 0.5044 | Yes |
| 15 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 1445 | 0.139 | 0.4941 | No |
| 16 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 1666 | 0.124 | 0.4877 | No |
| 17 | PROK2 | PROK2 Entrez, Source | prokineticin 2 | 1757 | 0.119 | 0.4906 | No |
| 18 | CBX8 | CBX8 Entrez, Source | chromobox homolog 8 (Pc class homolog, Drosophila) | 1783 | 0.118 | 0.4983 | No |
| 19 | KCNA4 | KCNA4 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 4 | 2019 | 0.107 | 0.4893 | No |
| 20 | VAMP3 | VAMP3 Entrez, Source | vesicle-associated membrane protein 3 (cellubrevin) | 2057 | 0.105 | 0.4951 | No |
| 21 | KCNJ13 | KCNJ13 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 13 | 2073 | 0.105 | 0.5025 | No |
| 22 | SEMA3A | SEMA3A Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A | 2466 | 0.089 | 0.4802 | No |
| 23 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 2548 | 0.086 | 0.4811 | No |
| 24 | FGF10 | FGF10 Entrez, Source | fibroblast growth factor 10 | 2765 | 0.078 | 0.4711 | No |
| 25 | CEPT1 | CEPT1 Entrez, Source | choline/ethanolamine phosphotransferase 1 | 2998 | 0.071 | 0.4594 | No |
| 26 | RAB11B | RAB11B Entrez, Source | RAB11B, member RAS oncogene family | 3071 | 0.069 | 0.4596 | No |
| 27 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 3208 | 0.065 | 0.4547 | No |
| 28 | RBBP9 | RBBP9 Entrez, Source | retinoblastoma binding protein 9 | 3213 | 0.065 | 0.4598 | No |
| 29 | CNTN4 | CNTN4 Entrez, Source | contactin 4 | 3319 | 0.062 | 0.4569 | No |
| 30 | TRAPPC3 | TRAPPC3 Entrez, Source | trafficking protein particle complex 3 | 3333 | 0.062 | 0.4610 | No |
| 31 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 3460 | 0.059 | 0.4562 | No |
| 32 | FGF20 | FGF20 Entrez, Source | fibroblast growth factor 20 | 3530 | 0.057 | 0.4557 | No |
| 33 | KIFC3 | KIFC3 Entrez, Source | kinesin family member C3 | 3636 | 0.054 | 0.4521 | No |
| 34 | NAPG | NAPG Entrez, Source | N-ethylmaleimide-sensitive factor attachment protein, gamma | 3699 | 0.052 | 0.4517 | No |
| 35 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 3704 | 0.052 | 0.4557 | No |
| 36 | MAG | MAG Entrez, Source | myelin associated glycoprotein | 4004 | 0.044 | 0.4367 | No |
| 37 | MAP1LC3A | MAP1LC3A Entrez, Source | microtubule-associated protein 1 light chain 3 alpha | 4039 | 0.044 | 0.4377 | No |
| 38 | DHRS4 | DHRS4 Entrez, Source | dehydrogenase/reductase (SDR family) member 4 | 4095 | 0.042 | 0.4370 | No |
| 39 | ESM1 | ESM1 Entrez, Source | endothelial cell-specific molecule 1 | 4360 | 0.037 | 0.4201 | No |
| 40 | SNX16 | SNX16 Entrez, Source | sorting nexin 16 | 4434 | 0.035 | 0.4174 | No |
| 41 | KCNH2 | KCNH2 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 2 | 4785 | 0.028 | 0.3933 | No |
| 42 | RAMP2 | RAMP2 Entrez, Source | receptor (calcitonin) activity modifying protein 2 | 4826 | 0.027 | 0.3925 | No |
| 43 | ACCN2 | ACCN2 Entrez, Source | amiloride-sensitive cation channel 2, neuronal | 5147 | 0.021 | 0.3701 | No |
| 44 | C7ORF10 | C7ORF10 Entrez, Source | chromosome 7 open reading frame 10 | 5156 | 0.021 | 0.3712 | No |
| 45 | VSNL1 | VSNL1 Entrez, Source | visinin-like 1 | 5194 | 0.020 | 0.3701 | No |
| 46 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 5323 | 0.018 | 0.3620 | No |
| 47 | APBA1 | APBA1 Entrez, Source | amyloid beta (A4) precursor protein-binding, family A, member 1 (X11) | 5471 | 0.015 | 0.3521 | No |
| 48 | GABBR1 | GABBR1 Entrez, Source | gamma-aminobutyric acid (GABA) B receptor, 1 | 5513 | 0.014 | 0.3502 | No |
| 49 | SERPINI1 | SERPINI1 Entrez, Source | serpin peptidase inhibitor, clade I (neuroserpin), member 1 | 5563 | 0.013 | 0.3476 | No |
| 50 | PSD | PSD Entrez, Source | pleckstrin and Sec7 domain containing | 5571 | 0.013 | 0.3482 | No |
| 51 | SLIT3 | SLIT3 Entrez, Source | slit homolog 3 (Drosophila) | 5734 | 0.010 | 0.3368 | No |
| 52 | CDK9 | CDK9 Entrez, Source | cyclin-dependent kinase 9 (CDC2-related kinase) | 5738 | 0.010 | 0.3374 | No |
| 53 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 6151 | 0.002 | 0.3065 | No |
| 54 | DCX | DCX Entrez, Source | doublecortex; lissencephaly, X-linked (doublecortin) | 6186 | 0.001 | 0.3040 | No |
| 55 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 6299 | -0.000 | 0.2956 | No |
| 56 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 6415 | -0.002 | 0.2871 | No |
| 57 | GNAO1 | GNAO1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O | 6475 | -0.003 | 0.2829 | No |
| 58 | SIRT6 | SIRT6 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 6 (S. cerevisiae) | 6551 | -0.005 | 0.2776 | No |
| 59 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 6582 | -0.005 | 0.2757 | No |
| 60 | PRRX1 | PRRX1 Entrez, Source | paired related homeobox 1 | 6636 | -0.006 | 0.2723 | No |
| 61 | PPP2R2B | PPP2R2B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), beta isoform | 6664 | -0.007 | 0.2708 | No |
| 62 | WSB2 | WSB2 Entrez, Source | WD repeat and SOCS box-containing 2 | 6832 | -0.009 | 0.2589 | No |
| 63 | CHRNB2 | CHRNB2 Entrez, Source | cholinergic receptor, nicotinic, beta 2 (neuronal) | 6897 | -0.010 | 0.2549 | No |
| 64 | TMC4 | TMC4 Entrez, Source | transmembrane channel-like 4 | 7210 | -0.016 | 0.2327 | No |
| 65 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 7291 | -0.018 | 0.2281 | No |
| 66 | GABRB1 | GABRB1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 1 | 7638 | -0.024 | 0.2040 | No |
| 67 | CHAD | CHAD Entrez, Source | chondroadherin | 7677 | -0.025 | 0.2031 | No |
| 68 | SLC6A9 | SLC6A9 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, glycine), member 9 | 7789 | -0.027 | 0.1970 | No |
| 69 | PPP1CB | PPP1CB Entrez, Source | protein phosphatase 1, catalytic subunit, beta isoform | 7860 | -0.028 | 0.1940 | No |
| 70 | OTX1 | OTX1 Entrez, Source | orthodenticle homolog 1 (Drosophila) | 7894 | -0.029 | 0.1939 | No |
| 71 | FLNC | FLNC Entrez, Source | filamin C, gamma (actin binding protein 280) | 7939 | -0.030 | 0.1930 | No |
| 72 | SLC38A5 | SLC38A5 Entrez, Source | solute carrier family 38, member 5 | 8010 | -0.031 | 0.1903 | No |
| 73 | SMPX | SMPX Entrez, Source | small muscle protein, X-linked | 8166 | -0.035 | 0.1814 | No |
| 74 | HCN3 | HCN3 Entrez, Source | hyperpolarization activated cyclic nucleotide-gated potassium channel 3 | 8351 | -0.038 | 0.1707 | No |
| 75 | DIO2 | DIO2 Entrez, Source | deiodinase, iodothyronine, type II | 8533 | -0.042 | 0.1604 | No |
| 76 | AMBN | AMBN Entrez, Source | ameloblastin (enamel matrix protein) | 8671 | -0.045 | 0.1538 | No |
| 77 | ACE2 | ACE2 Entrez, Source | angiotensin I converting enzyme (peptidyl-dipeptidase A) 2 | 9170 | -0.055 | 0.1207 | No |
| 78 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 9200 | -0.056 | 0.1231 | No |
| 79 | SLC30A3 | SLC30A3 Entrez, Source | solute carrier family 30 (zinc transporter), member 3 | 9521 | -0.063 | 0.1041 | No |
| 80 | ASCL3 | ASCL3 Entrez, Source | achaete-scute complex (Drosophila) homolog-like 3 | 9680 | -0.067 | 0.0977 | No |
| 81 | SYNPR | SYNPR Entrez, Source | synaptoporin | 9740 | -0.069 | 0.0988 | No |
| 82 | LIFR | LIFR Entrez, Source | leukemia inhibitory factor receptor alpha | 9964 | -0.075 | 0.0882 | No |
| 83 | TNNI2 | TNNI2 Entrez, Source | troponin I type 2 (skeletal, fast) | 10004 | -0.077 | 0.0915 | No |
| 84 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 10009 | -0.077 | 0.0975 | No |
| 85 | C2 | C2 Entrez, Source | complement component 2 | 10542 | -0.091 | 0.0648 | No |
| 86 | FGF17 | FGF17 Entrez, Source | fibroblast growth factor 17 | 10631 | -0.095 | 0.0659 | No |
| 87 | TCF8 | TCF8 Entrez, Source | transcription factor 8 (represses interleukin 2 expression) | 10979 | -0.107 | 0.0484 | No |
| 88 | UBXD2 | UBXD2 Entrez, Source | UBX domain containing 2 | 11474 | -0.128 | 0.0216 | No |
| 89 | RBX1 | RBX1 Entrez, Source | ring-box 1 | 11606 | -0.134 | 0.0226 | No |
| 90 | EIF4A2 | EIF4A2 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 2 | 11996 | -0.157 | 0.0061 | No |
| 91 | MECP2 | MECP2 Entrez, Source | methyl CpG binding protein 2 (Rett syndrome) | 12609 | -0.212 | -0.0228 | No |
| 92 | RHOQ | RHOQ Entrez, Source | ras homolog gene family, member Q | 12780 | -0.238 | -0.0162 | No |
| 93 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 13111 | -0.334 | -0.0139 | No |
| 94 | IGFALS | IGFALS Entrez, Source | insulin-like growth factor binding protein, acid labile subunit | 13179 | -0.383 | 0.0123 | No |