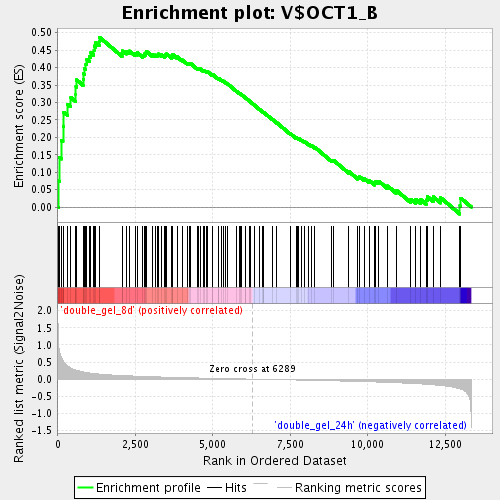
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$OCT1_B |
| Enrichment Score (ES) | 0.48713198 |
| Normalized Enrichment Score (NES) | 1.7603867 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.037113436 |
| FWER p-Value | 0.991 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 31 | 0.935 | 0.0749 | Yes |
| 2 | PFKFB1 | PFKFB1 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 | 47 | 0.829 | 0.1423 | Yes |
| 3 | LPL | LPL Entrez, Source | lipoprotein lipase | 107 | 0.646 | 0.1912 | Yes |
| 4 | GPM6A | GPM6A Entrez, Source | glycoprotein M6A | 165 | 0.540 | 0.2316 | Yes |
| 5 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 185 | 0.510 | 0.2723 | Yes |
| 6 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 312 | 0.390 | 0.2950 | Yes |
| 7 | CDH13 | CDH13 Entrez, Source | cadherin 13, H-cadherin (heart) | 417 | 0.328 | 0.3143 | Yes |
| 8 | GPC4 | GPC4 Entrez, Source | glypican 4 | 577 | 0.262 | 0.3239 | Yes |
| 9 | NFIA | NFIA Entrez, Source | nuclear factor I/A | 581 | 0.262 | 0.3453 | Yes |
| 10 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 603 | 0.257 | 0.3649 | Yes |
| 11 | TWIST1 | TWIST1 Entrez, Source | twist homolog 1 (acrocephalosyndactyly 3; Saethre-Chotzen syndrome) (Drosophila) | 813 | 0.211 | 0.3666 | Yes |
| 12 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 833 | 0.207 | 0.3823 | Yes |
| 13 | MRAS | MRAS Entrez, Source | muscle RAS oncogene homolog | 863 | 0.201 | 0.3968 | Yes |
| 14 | ADORA1 | ADORA1 Entrez, Source | adenosine A1 receptor | 901 | 0.195 | 0.4101 | Yes |
| 15 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 932 | 0.190 | 0.4236 | Yes |
| 16 | CPD | CPD Entrez, Source | carboxypeptidase D | 1024 | 0.179 | 0.4315 | Yes |
| 17 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 1062 | 0.175 | 0.4432 | Yes |
| 18 | DPYSL3 | DPYSL3 Entrez, Source | dihydropyrimidinase-like 3 | 1163 | 0.164 | 0.4492 | Yes |
| 19 | CDX2 | CDX2 Entrez, Source | caudal type homeobox transcription factor 2 | 1170 | 0.163 | 0.4622 | Yes |
| 20 | BDNF | BDNF Entrez, Source | brain-derived neurotrophic factor | 1218 | 0.159 | 0.4718 | Yes |
| 21 | FGF12 | FGF12 Entrez, Source | fibroblast growth factor 12 | 1334 | 0.148 | 0.4754 | Yes |
| 22 | POLM | POLM Entrez, Source | polymerase (DNA directed), mu | 1341 | 0.148 | 0.4871 | Yes |
| 23 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 2072 | 0.105 | 0.4406 | No |
| 24 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 2089 | 0.104 | 0.4480 | No |
| 25 | SLC24A1 | SLC24A1 Entrez, Source | solute carrier family 24 (sodium/potassium/calcium exchanger), member 1 | 2210 | 0.099 | 0.4471 | No |
| 26 | CSF3 | CSF3 Entrez, Source | colony stimulating factor 3 (granulocyte) | 2306 | 0.095 | 0.4478 | No |
| 27 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 2490 | 0.088 | 0.4413 | No |
| 28 | ITPR3 | ITPR3 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 3 | 2561 | 0.085 | 0.4430 | No |
| 29 | BLNK | BLNK Entrez, Source | B-cell linker | 2743 | 0.079 | 0.4358 | No |
| 30 | PPARGC1B | PPARGC1B Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, beta | 2782 | 0.077 | 0.4393 | No |
| 31 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 2831 | 0.076 | 0.4420 | No |
| 32 | GNB3 | GNB3 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 3 | 2846 | 0.076 | 0.4472 | No |
| 33 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 3037 | 0.070 | 0.4387 | No |
| 34 | TMOD2 | TMOD2 Entrez, Source | tropomodulin 2 (neuronal) | 3142 | 0.067 | 0.4364 | No |
| 35 | SLC1A2 | SLC1A2 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 2 | 3219 | 0.065 | 0.4360 | No |
| 36 | SFRP2 | SFRP2 Entrez, Source | secreted frizzled-related protein 2 | 3247 | 0.064 | 0.4393 | No |
| 37 | NFYB | NFYB Entrez, Source | nuclear transcription factor Y, beta | 3342 | 0.061 | 0.4372 | No |
| 38 | EAF2 | EAF2 Entrez, Source | ELL associated factor 2 | 3455 | 0.059 | 0.4336 | No |
| 39 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 3479 | 0.058 | 0.4367 | No |
| 40 | POU2F3 | POU2F3 Entrez, Source | POU domain, class 2, transcription factor 3 | 3508 | 0.057 | 0.4393 | No |
| 41 | ARPC5 | ARPC5 Entrez, Source | actin related protein 2/3 complex, subunit 5, 16kDa | 3675 | 0.053 | 0.4312 | No |
| 42 | STAT4 | STAT4 Entrez, Source | signal transducer and activator of transcription 4 | 3697 | 0.052 | 0.4339 | No |
| 43 | CRISP1 | CRISP1 Entrez, Source | cysteine-rich secretory protein 1 | 3706 | 0.052 | 0.4376 | No |
| 44 | POU3F4 | POU3F4 Entrez, Source | POU domain, class 3, transcription factor 4 | 3843 | 0.049 | 0.4314 | No |
| 45 | SH3GL3 | SH3GL3 Entrez, Source | SH3-domain GRB2-like 3 | 4015 | 0.044 | 0.4221 | No |
| 46 | BCL2 | BCL2 Entrez, Source | B-cell CLL/lymphoma 2 | 4187 | 0.040 | 0.4125 | No |
| 47 | HPCAL4 | HPCAL4 Entrez, Source | hippocalcin like 4 | 4243 | 0.039 | 0.4115 | No |
| 48 | GAB2 | GAB2 Entrez, Source | GRB2-associated binding protein 2 | 4277 | 0.038 | 0.4121 | No |
| 49 | MAP2K6 | MAP2K6 Entrez, Source | mitogen-activated protein kinase kinase 6 | 4510 | 0.034 | 0.3974 | No |
| 50 | FZD2 | FZD2 Entrez, Source | frizzled homolog 2 (Drosophila) | 4530 | 0.033 | 0.3987 | No |
| 51 | PTHLH | PTHLH Entrez, Source | parathyroid hormone-like hormone | 4595 | 0.032 | 0.3965 | No |
| 52 | RHOBTB2 | RHOBTB2 Entrez, Source | Rho-related BTB domain containing 2 | 4699 | 0.030 | 0.3912 | No |
| 53 | SLC6A15 | SLC6A15 Entrez, Source | solute carrier family 6, member 15 | 4739 | 0.029 | 0.3907 | No |
| 54 | ARHGAP4 | ARHGAP4 Entrez, Source | Rho GTPase activating protein 4 | 4793 | 0.028 | 0.3890 | No |
| 55 | RAB26 | RAB26 Entrez, Source | RAB26, member RAS oncogene family | 4823 | 0.028 | 0.3891 | No |
| 56 | THRA | THRA Entrez, Source | thyroid hormone receptor, alpha (erythroblastic leukemia viral (v-erb-a) oncogene homolog, avian) | 4984 | 0.024 | 0.3790 | No |
| 57 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 4992 | 0.024 | 0.3805 | No |
| 58 | ACBD4 | ACBD4 Entrez, Source | acyl-Coenzyme A binding domain containing 4 | 5190 | 0.020 | 0.3673 | No |
| 59 | VSNL1 | VSNL1 Entrez, Source | visinin-like 1 | 5194 | 0.020 | 0.3688 | No |
| 60 | JUND | JUND Entrez, Source | jun D proto-oncogene | 5289 | 0.019 | 0.3632 | No |
| 61 | COL23A1 | COL23A1 Entrez, Source | collagen, type XXIII, alpha 1 | 5329 | 0.018 | 0.3618 | No |
| 62 | GPR4 | GPR4 Entrez, Source | G protein-coupled receptor 4 | 5412 | 0.016 | 0.3570 | No |
| 63 | NEUROG1 | NEUROG1 Entrez, Source | neurogenin 1 | 5488 | 0.015 | 0.3525 | No |
| 64 | NRXN3 | NRXN3 Entrez, Source | neurexin 3 | 5770 | 0.010 | 0.3321 | No |
| 65 | CD79B | CD79B Entrez, Source | CD79b molecule, immunoglobulin-associated beta | 5871 | 0.008 | 0.3252 | No |
| 66 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 5890 | 0.007 | 0.3244 | No |
| 67 | SLC39A13 | SLC39A13 Entrez, Source | solute carrier family 39 (zinc transporter), member 13 | 5912 | 0.007 | 0.3234 | No |
| 68 | TRPM1 | TRPM1 Entrez, Source | transient receptor potential cation channel, subfamily M, member 1 | 6043 | 0.004 | 0.3139 | No |
| 69 | LRRN1 | LRRN1 Entrez, Source | leucine rich repeat neuronal 1 | 6069 | 0.004 | 0.3124 | No |
| 70 | KCNN3 | KCNN3 Entrez, Source | potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 | 6198 | 0.001 | 0.3028 | No |
| 71 | PCYT1B | PCYT1B Entrez, Source | phosphate cytidylyltransferase 1, choline, beta | 6229 | 0.001 | 0.3006 | No |
| 72 | IFIT2 | IFIT2 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 2 | 6336 | -0.001 | 0.2926 | No |
| 73 | NR2C2 | NR2C2 Entrez, Source | nuclear receptor subfamily 2, group C, member 2 | 6511 | -0.004 | 0.2798 | No |
| 74 | NRXN1 | NRXN1 Entrez, Source | neurexin 1 | 6602 | -0.006 | 0.2735 | No |
| 75 | PRRX1 | PRRX1 Entrez, Source | paired related homeobox 1 | 6636 | -0.006 | 0.2715 | No |
| 76 | CXXC4 | CXXC4 Entrez, Source | CXXC finger 4 | 6637 | -0.006 | 0.2720 | No |
| 77 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 6918 | -0.011 | 0.2518 | No |
| 78 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 7067 | -0.014 | 0.2417 | No |
| 79 | MYBPC1 | MYBPC1 Entrez, Source | myosin binding protein C, slow type | 7507 | -0.022 | 0.2103 | No |
| 80 | CD86 | CD86 Entrez, Source | CD86 molecule | 7686 | -0.025 | 0.1989 | No |
| 81 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 7732 | -0.026 | 0.1977 | No |
| 82 | DLGAP4 | DLGAP4 Entrez, Source | discs, large (Drosophila) homolog-associated protein 4 | 7775 | -0.027 | 0.1967 | No |
| 83 | ALK | ALK Entrez, Source | anaplastic lymphoma kinase (Ki-1) | 7870 | -0.029 | 0.1920 | No |
| 84 | TCF2 | TCF2 Entrez, Source | transcription factor 2, hepatic; LF-B3; variant hepatic nuclear factor | 7953 | -0.030 | 0.1883 | No |
| 85 | ESRRA | ESRRA Entrez, Source | estrogen-related receptor alpha | 8092 | -0.033 | 0.1806 | No |
| 86 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 8169 | -0.035 | 0.1777 | No |
| 87 | FGF14 | FGF14 Entrez, Source | fibroblast growth factor 14 | 8295 | -0.037 | 0.1714 | No |
| 88 | NXPH1 | NXPH1 Entrez, Source | neurexophilin 1 | 8833 | -0.049 | 0.1348 | No |
| 89 | ADORA2A | ADORA2A Entrez, Source | adenosine A2a receptor | 8894 | -0.050 | 0.1344 | No |
| 90 | H3F3B | H3F3B Entrez, Source | H3 histone, family 3B (H3.3B) | 9390 | -0.060 | 0.1020 | No |
| 91 | ASCL3 | ASCL3 Entrez, Source | achaete-scute complex (Drosophila) homolog-like 3 | 9680 | -0.067 | 0.0857 | No |
| 92 | SYNPR | SYNPR Entrez, Source | synaptoporin | 9740 | -0.069 | 0.0870 | No |
| 93 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 9896 | -0.074 | 0.0813 | No |
| 94 | CRYZL1 | CRYZL1 Entrez, Source | crystallin, zeta (quinone reductase)-like 1 | 10057 | -0.078 | 0.0757 | No |
| 95 | TSGA14 | TSGA14 Entrez, Source | testis specific, 14 | 10225 | -0.082 | 0.0699 | No |
| 96 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 10258 | -0.083 | 0.0743 | No |
| 97 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 10356 | -0.086 | 0.0741 | No |
| 98 | BIN3 | BIN3 Entrez, Source | bridging integrator 3 | 10638 | -0.095 | 0.0608 | No |
| 99 | ITSN1 | ITSN1 Entrez, Source | intersectin 1 (SH3 domain protein) | 10914 | -0.105 | 0.0487 | No |
| 100 | TAS2R13 | TAS2R13 Entrez, Source | taste receptor, type 2, member 13 | 11390 | -0.124 | 0.0231 | No |
| 101 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 11550 | -0.131 | 0.0219 | No |
| 102 | CSNK1E | CSNK1E Entrez, Source | casein kinase 1, epsilon | 11711 | -0.139 | 0.0213 | No |
| 103 | ABTB2 | ABTB2 Entrez, Source | ankyrin repeat and BTB (POZ) domain containing 2 | 11886 | -0.149 | 0.0205 | No |
| 104 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 11925 | -0.152 | 0.0301 | No |
| 105 | BAT2 | BAT2 Entrez, Source | HLA-B associated transcript 2 | 12112 | -0.166 | 0.0297 | No |
| 106 | NR1D1 | NR1D1 Entrez, Source | nuclear receptor subfamily 1, group D, member 1 | 12353 | -0.184 | 0.0268 | No |
| 107 | NDNL2 | NDNL2 Entrez, Source | necdin-like 2 | 12975 | -0.285 | 0.0034 | No |
| 108 | TCEAL1 | TCEAL1 Entrez, Source | transcription elongation factor A (SII)-like 1 | 13002 | -0.293 | 0.0257 | No |

