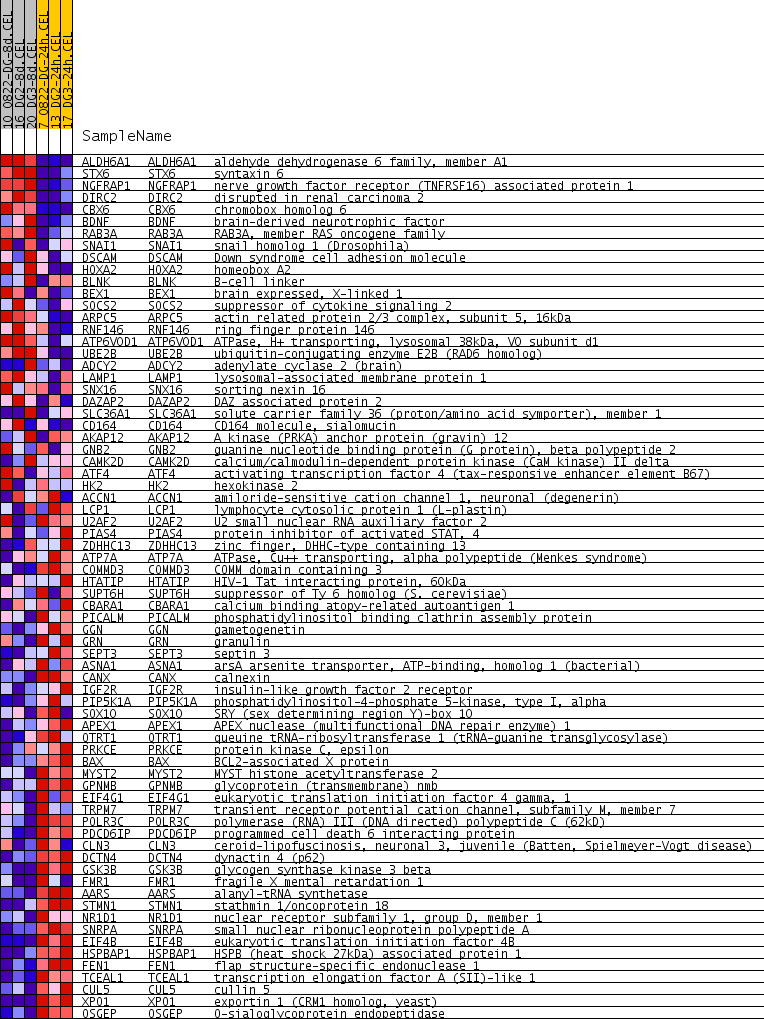
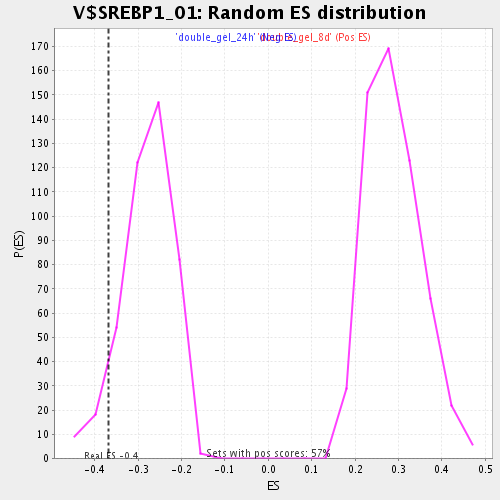
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_24h |
| GeneSet | V$SREBP1_01 |
| Enrichment Score (ES) | -0.3694123 |
| Normalized Enrichment Score (NES) | -1.3265922 |
| Nominal p-value | 0.06912442 |
| FDR q-value | 0.37840334 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 300 | 0.402 | 0.0286 | No |
| 2 | STX6 | STX6 Entrez, Source | syntaxin 6 | 766 | 0.222 | 0.0218 | No |
| 3 | NGFRAP1 | NGFRAP1 Entrez, Source | nerve growth factor receptor (TNFRSF16) associated protein 1 | 830 | 0.208 | 0.0434 | No |
| 4 | DIRC2 | DIRC2 Entrez, Source | disrupted in renal carcinoma 2 | 926 | 0.191 | 0.0606 | No |
| 5 | CBX6 | CBX6 Entrez, Source | chromobox homolog 6 | 1181 | 0.163 | 0.0622 | No |
| 6 | BDNF | BDNF Entrez, Source | brain-derived neurotrophic factor | 1218 | 0.159 | 0.0797 | No |
| 7 | RAB3A | RAB3A Entrez, Source | RAB3A, member RAS oncogene family | 1306 | 0.150 | 0.0923 | No |
| 8 | SNAI1 | SNAI1 Entrez, Source | snail homolog 1 (Drosophila) | 2341 | 0.094 | 0.0263 | No |
| 9 | DSCAM | DSCAM Entrez, Source | Down syndrome cell adhesion molecule | 2385 | 0.092 | 0.0348 | No |
| 10 | HOXA2 | HOXA2 Entrez, Source | homeobox A2 | 2403 | 0.092 | 0.0452 | No |
| 11 | BLNK | BLNK Entrez, Source | B-cell linker | 2743 | 0.079 | 0.0296 | No |
| 12 | BEX1 | BEX1 Entrez, Source | brain expressed, X-linked 1 | 3234 | 0.064 | 0.0009 | No |
| 13 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 3280 | 0.063 | 0.0056 | No |
| 14 | ARPC5 | ARPC5 Entrez, Source | actin related protein 2/3 complex, subunit 5, 16kDa | 3675 | 0.053 | -0.0174 | No |
| 15 | RNF146 | RNF146 Entrez, Source | ring finger protein 146 | 3779 | 0.051 | -0.0187 | No |
| 16 | ATP6V0D1 | ATP6V0D1 Entrez, Source | ATPase, H+ transporting, lysosomal 38kDa, V0 subunit d1 | 3820 | 0.050 | -0.0154 | No |
| 17 | UBE2B | UBE2B Entrez, Source | ubiquitin-conjugating enzyme E2B (RAD6 homolog) | 3964 | 0.045 | -0.0204 | No |
| 18 | ADCY2 | ADCY2 Entrez, Source | adenylate cyclase 2 (brain) | 4158 | 0.041 | -0.0298 | No |
| 19 | LAMP1 | LAMP1 Entrez, Source | lysosomal-associated membrane protein 1 | 4337 | 0.037 | -0.0385 | No |
| 20 | SNX16 | SNX16 Entrez, Source | sorting nexin 16 | 4434 | 0.035 | -0.0413 | No |
| 21 | DAZAP2 | DAZAP2 Entrez, Source | DAZ associated protein 2 | 5107 | 0.022 | -0.0891 | No |
| 22 | SLC36A1 | SLC36A1 Entrez, Source | solute carrier family 36 (proton/amino acid symporter), member 1 | 5193 | 0.020 | -0.0929 | No |
| 23 | CD164 | CD164 Entrez, Source | CD164 molecule, sialomucin | 5509 | 0.014 | -0.1148 | No |
| 24 | AKAP12 | AKAP12 Entrez, Source | A kinase (PRKA) anchor protein (gravin) 12 | 5716 | 0.011 | -0.1290 | No |
| 25 | GNB2 | GNB2 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 2 | 5801 | 0.009 | -0.1341 | No |
| 26 | CAMK2D | CAMK2D Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II delta | 5975 | 0.006 | -0.1465 | No |
| 27 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 6295 | -0.000 | -0.1705 | No |
| 28 | HK2 | HK2 Entrez, Source | hexokinase 2 | 6340 | -0.001 | -0.1737 | No |
| 29 | ACCN1 | ACCN1 Entrez, Source | amiloride-sensitive cation channel 1, neuronal (degenerin) | 7079 | -0.014 | -0.2276 | No |
| 30 | LCP1 | LCP1 Entrez, Source | lymphocyte cytosolic protein 1 (L-plastin) | 7308 | -0.018 | -0.2425 | No |
| 31 | U2AF2 | U2AF2 Entrez, Source | U2 small nuclear RNA auxiliary factor 2 | 7498 | -0.022 | -0.2540 | No |
| 32 | PIAS4 | PIAS4 Entrez, Source | protein inhibitor of activated STAT, 4 | 7603 | -0.024 | -0.2588 | No |
| 33 | ZDHHC13 | ZDHHC13 Entrez, Source | zinc finger, DHHC-type containing 13 | 7981 | -0.031 | -0.2833 | No |
| 34 | ATP7A | ATP7A Entrez, Source | ATPase, Cu++ transporting, alpha polypeptide (Menkes syndrome) | 8081 | -0.033 | -0.2866 | No |
| 35 | COMMD3 | COMMD3 Entrez, Source | COMM domain containing 3 | 8085 | -0.033 | -0.2826 | No |
| 36 | HTATIP | HTATIP Entrez, Source | HIV-1 Tat interacting protein, 60kDa | 8212 | -0.035 | -0.2875 | No |
| 37 | SUPT6H | SUPT6H Entrez, Source | suppressor of Ty 6 homolog (S. cerevisiae) | 8289 | -0.037 | -0.2885 | No |
| 38 | CBARA1 | CBARA1 Entrez, Source | calcium binding atopy-related autoantigen 1 | 8308 | -0.038 | -0.2851 | No |
| 39 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 8462 | -0.041 | -0.2915 | No |
| 40 | GGN | GGN Entrez, Source | gametogenetin | 8728 | -0.046 | -0.3056 | No |
| 41 | GRN | GRN Entrez, Source | granulin | 8784 | -0.047 | -0.3037 | No |
| 42 | SEPT3 | SEPT3 Entrez, Source | septin 3 | 9018 | -0.052 | -0.3146 | No |
| 43 | ASNA1 | ASNA1 Entrez, Source | arsA arsenite transporter, ATP-binding, homolog 1 (bacterial) | 9112 | -0.054 | -0.3147 | No |
| 44 | CANX | CANX Entrez, Source | calnexin | 9223 | -0.057 | -0.3158 | No |
| 45 | IGF2R | IGF2R Entrez, Source | insulin-like growth factor 2 receptor | 9465 | -0.062 | -0.3261 | No |
| 46 | PIP5K1A | PIP5K1A Entrez, Source | phosphatidylinositol-4-phosphate 5-kinase, type I, alpha | 9484 | -0.063 | -0.3195 | No |
| 47 | SOX10 | SOX10 Entrez, Source | SRY (sex determining region Y)-box 10 | 10051 | -0.078 | -0.3522 | No |
| 48 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 10112 | -0.079 | -0.3466 | No |
| 49 | QTRT1 | QTRT1 Entrez, Source | queuine tRNA-ribosyltransferase 1 (tRNA-guanine transglycosylase) | 10396 | -0.087 | -0.3569 | Yes |
| 50 | PRKCE | PRKCE Entrez, Source | protein kinase C, epsilon | 10468 | -0.089 | -0.3508 | Yes |
| 51 | BAX | BAX Entrez, Source | BCL2-associated X protein | 10485 | -0.090 | -0.3406 | Yes |
| 52 | MYST2 | MYST2 Entrez, Source | MYST histone acetyltransferase 2 | 10762 | -0.099 | -0.3488 | Yes |
| 53 | GPNMB | GPNMB Entrez, Source | glycoprotein (transmembrane) nmb | 10958 | -0.107 | -0.3499 | Yes |
| 54 | EIF4G1 | EIF4G1 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 1 | 10984 | -0.108 | -0.3381 | Yes |
| 55 | TRPM7 | TRPM7 Entrez, Source | transient receptor potential cation channel, subfamily M, member 7 | 11259 | -0.119 | -0.3436 | Yes |
| 56 | POLR3C | POLR3C Entrez, Source | polymerase (RNA) III (DNA directed) polypeptide C (62kD) | 11398 | -0.125 | -0.3381 | Yes |
| 57 | PDCD6IP | PDCD6IP Entrez, Source | programmed cell death 6 interacting protein | 11814 | -0.145 | -0.3509 | Yes |
| 58 | CLN3 | CLN3 Entrez, Source | ceroid-lipofuscinosis, neuronal 3, juvenile (Batten, Spielmeyer-Vogt disease) | 11907 | -0.150 | -0.3387 | Yes |
| 59 | DCTN4 | DCTN4 Entrez, Source | dynactin 4 (p62) | 11929 | -0.152 | -0.3210 | Yes |
| 60 | GSK3B | GSK3B Entrez, Source | glycogen synthase kinase 3 beta | 11989 | -0.156 | -0.3055 | Yes |
| 61 | FMR1 | FMR1 Entrez, Source | fragile X mental retardation 1 | 12198 | -0.171 | -0.2994 | Yes |
| 62 | AARS | AARS Entrez, Source | alanyl-tRNA synthetase | 12234 | -0.174 | -0.2799 | Yes |
| 63 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 12244 | -0.175 | -0.2583 | Yes |
| 64 | NR1D1 | NR1D1 Entrez, Source | nuclear receptor subfamily 1, group D, member 1 | 12353 | -0.184 | -0.2430 | Yes |
| 65 | SNRPA | SNRPA Entrez, Source | small nuclear ribonucleoprotein polypeptide A | 12756 | -0.234 | -0.2435 | Yes |
| 66 | EIF4B | EIF4B Entrez, Source | eukaryotic translation initiation factor 4B | 12823 | -0.247 | -0.2171 | Yes |
| 67 | HSPBAP1 | HSPBAP1 Entrez, Source | HSPB (heat shock 27kDa) associated protein 1 | 12882 | -0.262 | -0.1882 | Yes |
| 68 | FEN1 | FEN1 Entrez, Source | flap structure-specific endonuclease 1 | 12971 | -0.283 | -0.1588 | Yes |
| 69 | TCEAL1 | TCEAL1 Entrez, Source | transcription elongation factor A (SII)-like 1 | 13002 | -0.293 | -0.1238 | Yes |
| 70 | CUL5 | CUL5 Entrez, Source | cullin 5 | 13074 | -0.316 | -0.0890 | Yes |
| 71 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 13111 | -0.334 | -0.0492 | Yes |
| 72 | OSGEP | OSGEP Entrez, Source | O-sialoglycoprotein endopeptidase | 13271 | -0.523 | 0.0054 | Yes |