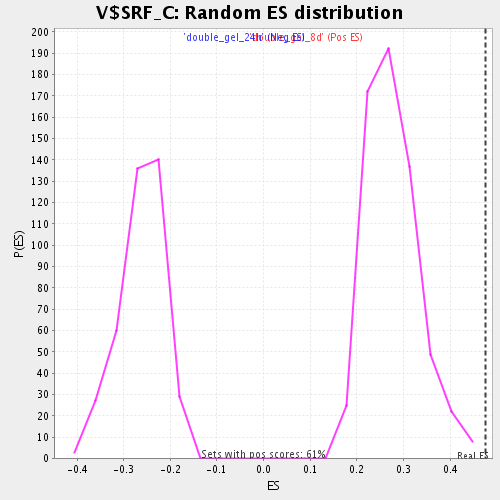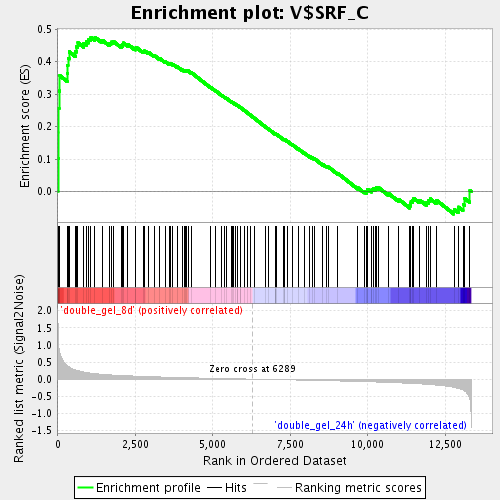
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$SRF_C |
| Enrichment Score (ES) | 0.47507045 |
| Normalized Enrichment Score (NES) | 1.7253336 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.047012076 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 3 | 1.622 | 0.1024 | Yes |
| 2 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 12 | 1.254 | 0.1812 | Yes |
| 3 | SLC25A4 | SLC25A4 Entrez, Source | solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 4 | 13 | 1.195 | 0.2568 | Yes |
| 4 | TAGLN | TAGLN Entrez, Source | transgelin | 40 | 0.874 | 0.3101 | Yes |
| 5 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 60 | 0.762 | 0.3569 | Yes |
| 6 | RGS3 | RGS3 Entrez, Source | regulator of G-protein signalling 3 | 301 | 0.402 | 0.3642 | Yes |
| 7 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 312 | 0.390 | 0.3881 | Yes |
| 8 | POPDC2 | POPDC2 Entrez, Source | popeye domain containing 2 | 327 | 0.374 | 0.4107 | Yes |
| 9 | RAB30 | RAB30 Entrez, Source | RAB30, member RAS oncogene family | 363 | 0.357 | 0.4306 | Yes |
| 10 | RNF39 | RNF39 Entrez, Source | ring finger protein 39 | 568 | 0.265 | 0.4320 | Yes |
| 11 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 603 | 0.257 | 0.4456 | Yes |
| 12 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 645 | 0.246 | 0.4581 | Yes |
| 13 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 833 | 0.207 | 0.4571 | Yes |
| 14 | ACTG2 | ACTG2 Entrez, Source | actin, gamma 2, smooth muscle, enteric | 919 | 0.192 | 0.4628 | Yes |
| 15 | PLN | PLN Entrez, Source | phospholamban | 994 | 0.183 | 0.4688 | Yes |
| 16 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 1062 | 0.175 | 0.4748 | Yes |
| 17 | ITGB6 | ITGB6 Entrez, Source | integrin, beta 6 | 1195 | 0.161 | 0.4751 | Yes |
| 18 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 1427 | 0.140 | 0.4665 | No |
| 19 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 1666 | 0.124 | 0.4564 | No |
| 20 | ANXA6 | ANXA6 Entrez, Source | annexin A6 | 1714 | 0.122 | 0.4606 | No |
| 21 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 1785 | 0.118 | 0.4627 | No |
| 22 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 2048 | 0.106 | 0.4496 | No |
| 23 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 2090 | 0.104 | 0.4531 | No |
| 24 | MYO1E | MYO1E Entrez, Source | myosin IE | 2117 | 0.103 | 0.4576 | No |
| 25 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 2261 | 0.097 | 0.4530 | No |
| 26 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 2490 | 0.088 | 0.4413 | No |
| 27 | IER2 | IER2 Entrez, Source | immediate early response 2 | 2518 | 0.087 | 0.4448 | No |
| 28 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 2752 | 0.078 | 0.4321 | No |
| 29 | TGFB1I1 | TGFB1I1 Entrez, Source | transforming growth factor beta 1 induced transcript 1 | 2804 | 0.077 | 0.4331 | No |
| 30 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 2930 | 0.073 | 0.4283 | No |
| 31 | GPR20 | GPR20 Entrez, Source | G protein-coupled receptor 20 | 3108 | 0.068 | 0.4193 | No |
| 32 | DAPK3 | DAPK3 Entrez, Source | death-associated protein kinase 3 | 3290 | 0.063 | 0.4096 | No |
| 33 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 3460 | 0.059 | 0.4005 | No |
| 34 | MAPK14 | MAPK14 Entrez, Source | mitogen-activated protein kinase 14 | 3602 | 0.055 | 0.3933 | No |
| 35 | EGR4 | EGR4 Entrez, Source | early growth response 4 | 3630 | 0.054 | 0.3947 | No |
| 36 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 3704 | 0.052 | 0.3925 | No |
| 37 | POU3F4 | POU3F4 Entrez, Source | POU domain, class 3, transcription factor 4 | 3843 | 0.049 | 0.3852 | No |
| 38 | GFPT2 | GFPT2 Entrez, Source | glutamine-fructose-6-phosphate transaminase 2 | 4020 | 0.044 | 0.3746 | No |
| 39 | MYO1B | MYO1B Entrez, Source | myosin IB | 4074 | 0.043 | 0.3734 | No |
| 40 | SSH3 | SSH3 Entrez, Source | slingshot homolog 3 (Drosophila) | 4110 | 0.042 | 0.3734 | No |
| 41 | TNNC1 | TNNC1 Entrez, Source | troponin C type 1 (slow) | 4157 | 0.041 | 0.3725 | No |
| 42 | DSTN | DSTN Entrez, Source | destrin (actin depolymerizing factor) | 4198 | 0.040 | 0.3720 | No |
| 43 | RHOJ | RHOJ Entrez, Source | ras homolog gene family, member J | 4296 | 0.038 | 0.3670 | No |
| 44 | IL17B | IL17B Entrez, Source | interleukin 17B | 4928 | 0.025 | 0.3210 | No |
| 45 | CALD1 | CALD1 Entrez, Source | caldesmon 1 | 5081 | 0.023 | 0.3109 | No |
| 46 | SLC2A4 | SLC2A4 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 4 | 5283 | 0.019 | 0.2969 | No |
| 47 | FGFRL1 | FGFRL1 Entrez, Source | fibroblast growth factor receptor-like 1 | 5385 | 0.017 | 0.2904 | No |
| 48 | CACNA1B | CACNA1B Entrez, Source | calcium channel, voltage-dependent, L type, alpha 1B subunit | 5446 | 0.016 | 0.2868 | No |
| 49 | AOC3 | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 5594 | 0.013 | 0.2765 | No |
| 50 | TLE3 | TLE3 Entrez, Source | transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) | 5643 | 0.012 | 0.2737 | No |
| 51 | PLCB3 | PLCB3 Entrez, Source | phospholipase C, beta 3 (phosphatidylinositol-specific) | 5675 | 0.011 | 0.2721 | No |
| 52 | AKAP12 | AKAP12 Entrez, Source | A kinase (PRKA) anchor protein (gravin) 12 | 5716 | 0.011 | 0.2697 | No |
| 53 | ACTB | ACTB Entrez, Source | actin, beta | 5788 | 0.009 | 0.2649 | No |
| 54 | PADI2 | PADI2 Entrez, Source | peptidyl arginine deiminase, type II | 5804 | 0.009 | 0.2644 | No |
| 55 | KCNK3 | KCNK3 Entrez, Source | potassium channel, subfamily K, member 3 | 5898 | 0.007 | 0.2578 | No |
| 56 | NR2F1 | NR2F1 Entrez, Source | nuclear receptor subfamily 2, group F, member 1 | 5904 | 0.007 | 0.2579 | No |
| 57 | GRK6 | GRK6 Entrez, Source | G protein-coupled receptor kinase 6 | 6012 | 0.005 | 0.2501 | No |
| 58 | SRD5A2 | SRD5A2 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 2 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 2) | 6110 | 0.003 | 0.2430 | No |
| 59 | MCAM | MCAM Entrez, Source | melanoma cell adhesion molecule | 6217 | 0.001 | 0.2350 | No |
| 60 | MRGPRF | MRGPRF Entrez, Source | MAS-related GPR, member F | 6355 | -0.001 | 0.2247 | No |
| 61 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 6711 | -0.007 | 0.1984 | No |
| 62 | PDLIM4 | PDLIM4 Entrez, Source | PDZ and LIM domain 4 | 6797 | -0.009 | 0.1925 | No |
| 63 | DLL1 | DLL1 Entrez, Source | delta-like 1 (Drosophila) | 7008 | -0.012 | 0.1774 | No |
| 64 | CNN1 | CNN1 Entrez, Source | calponin 1, basic, smooth muscle | 7011 | -0.012 | 0.1781 | No |
| 65 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 7017 | -0.013 | 0.1785 | No |
| 66 | JPH2 | JPH2 Entrez, Source | junctophilin 2 | 7046 | -0.013 | 0.1772 | No |
| 67 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 7291 | -0.018 | 0.1599 | No |
| 68 | ITGA7 | ITGA7 Entrez, Source | integrin, alpha 7 | 7298 | -0.018 | 0.1605 | No |
| 69 | LCP1 | LCP1 Entrez, Source | lymphocyte cytosolic protein 1 (L-plastin) | 7308 | -0.018 | 0.1610 | No |
| 70 | FOXE3 | FOXE3 Entrez, Source | forkhead box E3 | 7406 | -0.020 | 0.1549 | No |
| 71 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 7565 | -0.023 | 0.1444 | No |
| 72 | KCNMB1 | KCNMB1 Entrez, Source | potassium large conductance calcium-activated channel, subfamily M, beta member 1 | 7758 | -0.027 | 0.1316 | No |
| 73 | TCF2 | TCF2 Entrez, Source | transcription factor 2, hepatic; LF-B3; variant hepatic nuclear factor | 7953 | -0.030 | 0.1189 | No |
| 74 | YWHAZ | YWHAZ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide | 8135 | -0.034 | 0.1074 | No |
| 75 | MAP1A | MAP1A Entrez, Source | microtubule-associated protein 1A | 8202 | -0.035 | 0.1046 | No |
| 76 | MYH11 | MYH11 Entrez, Source | myosin, heavy chain 11, smooth muscle | 8278 | -0.037 | 0.1013 | No |
| 77 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 8551 | -0.042 | 0.0834 | No |
| 78 | TITF1 | TITF1 Entrez, Source | thyroid transcription factor 1 | 8658 | -0.045 | 0.0782 | No |
| 79 | GGN | GGN Entrez, Source | gametogenetin | 8728 | -0.046 | 0.0759 | No |
| 80 | RASD2 | RASD2 Entrez, Source | RASD family, member 2 | 9030 | -0.052 | 0.0565 | No |
| 81 | PRKAB2 | PRKAB2 Entrez, Source | protein kinase, AMP-activated, beta 2 non-catalytic subunit | 9678 | -0.067 | 0.0119 | No |
| 82 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 9896 | -0.074 | 0.0001 | No |
| 83 | CRK | CRK Entrez, Source | v-crk sarcoma virus CT10 oncogene homolog (avian) | 9974 | -0.076 | -0.0009 | No |
| 84 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 9978 | -0.076 | 0.0037 | No |
| 85 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 9991 | -0.076 | 0.0076 | No |
| 86 | MATR3 | MATR3 Entrez, Source | matrin 3 | 10114 | -0.079 | 0.0034 | No |
| 87 | PFN1 | PFN1 Entrez, Source | profilin 1 | 10135 | -0.080 | 0.0070 | No |
| 88 | RBBP7 | RBBP7 Entrez, Source | retinoblastoma binding protein 7 | 10181 | -0.081 | 0.0087 | No |
| 89 | SLC7A1 | SLC7A1 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 | 10239 | -0.083 | 0.0096 | No |
| 90 | TPM3 | TPM3 Entrez, Source | tropomyosin 3 | 10270 | -0.084 | 0.0126 | No |
| 91 | PPP1R12A | PPP1R12A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12A | 10350 | -0.086 | 0.0121 | No |
| 92 | ACTR3 | ACTR3 Entrez, Source | ARP3 actin-related protein 3 homolog (yeast) | 10680 | -0.096 | -0.0067 | No |
| 93 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 11004 | -0.108 | -0.0242 | No |
| 94 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 11359 | -0.123 | -0.0432 | No |
| 95 | CFL1 | CFL1 Entrez, Source | cofilin 1 (non-muscle) | 11382 | -0.124 | -0.0370 | No |
| 96 | CKM | CKM Entrez, Source | creatine kinase, muscle | 11394 | -0.125 | -0.0300 | No |
| 97 | ROCK2 | ROCK2 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 2 | 11450 | -0.127 | -0.0261 | No |
| 98 | DIXDC1 | DIXDC1 Entrez, Source | DIX domain containing 1 | 11483 | -0.128 | -0.0204 | No |
| 99 | MUS81 | MUS81 Entrez, Source | MUS81 endonuclease homolog (S. cerevisiae) | 11677 | -0.137 | -0.0263 | No |
| 100 | PDLIM7 | PDLIM7 Entrez, Source | PDZ and LIM domain 7 (enigma) | 11900 | -0.150 | -0.0336 | No |
| 101 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 11961 | -0.154 | -0.0284 | No |
| 102 | ASB2 | ASB2 Entrez, Source | ankyrin repeat and SOCS box-containing 2 | 12011 | -0.158 | -0.0221 | No |
| 103 | PNKP | PNKP Entrez, Source | polynucleotide kinase 3'-phosphatase | 12212 | -0.172 | -0.0263 | No |
| 104 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 12783 | -0.238 | -0.0543 | No |
| 105 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 12920 | -0.270 | -0.0474 | No |
| 106 | STAT5B | STAT5B Entrez, Source | signal transducer and activator of transcription 5B | 13076 | -0.317 | -0.0391 | No |
| 107 | CA3 | CA3 Entrez, Source | carbonic anhydrase III, muscle specific | 13129 | -0.343 | -0.0213 | No |
| 108 | GADD45G | GADD45G Entrez, Source | growth arrest and DNA-damage-inducible, gamma | 13297 | -0.589 | 0.0034 | No |