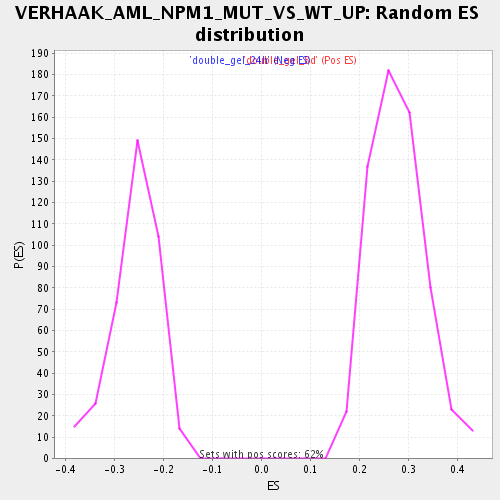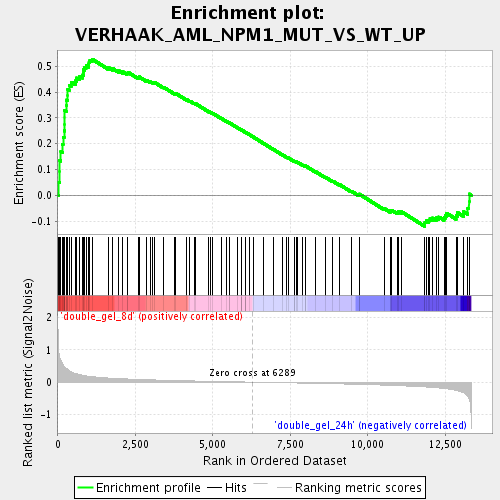
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | VERHAAK_AML_NPM1_MUT_VS_WT_UP |
| Enrichment Score (ES) | 0.52740556 |
| Normalized Enrichment Score (NES) | 1.8989547 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.012839542 |
| FWER p-Value | 0.538 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 27 | 0.963 | 0.0513 | Yes |
| 2 | S100A6 | S100A6 Entrez, Source | S100 calcium binding protein A6 | 55 | 0.785 | 0.0928 | Yes |
| 3 | VNN1 | VNN1 Entrez, Source | vanin 1 | 58 | 0.766 | 0.1351 | Yes |
| 4 | LGALS2 | LGALS2 Entrez, Source | lectin, galactoside-binding, soluble, 2 (galectin 2) | 95 | 0.672 | 0.1697 | Yes |
| 5 | BCL2A1 | BCL2A1 Entrez, Source | BCL2-related protein A1 | 147 | 0.568 | 0.1973 | Yes |
| 6 | TMEM176B | TMEM176B Entrez, Source | transmembrane protein 176B | 168 | 0.535 | 0.2254 | Yes |
| 7 | FBP1 | FBP1 Entrez, Source | fructose-1,6-bisphosphatase 1 | 203 | 0.484 | 0.2497 | Yes |
| 8 | PRKAR2B | PRKAR2B Entrez, Source | protein kinase, cAMP-dependent, regulatory, type II, beta | 214 | 0.474 | 0.2752 | Yes |
| 9 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 217 | 0.472 | 0.3012 | Yes |
| 10 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 218 | 0.471 | 0.3273 | Yes |
| 11 | GGT1 | GGT1 Entrez, Source | gamma-glutamyltransferase 1 | 271 | 0.423 | 0.3469 | Yes |
| 12 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 279 | 0.415 | 0.3693 | Yes |
| 13 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 315 | 0.388 | 0.3882 | Yes |
| 14 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 318 | 0.386 | 0.4094 | Yes |
| 15 | PF4 | PF4 Entrez, Source | platelet factor 4 (chemokine (C-X-C motif) ligand 4) | 364 | 0.357 | 0.4258 | Yes |
| 16 | DEFB1 | DEFB1 Entrez, Source | defensin, beta 1 | 429 | 0.322 | 0.4388 | Yes |
| 17 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 565 | 0.265 | 0.4433 | Yes |
| 18 | FGR | FGR Entrez, Source | Gardner-Rasheed feline sarcoma viral (v-fgr) oncogene homolog | 605 | 0.256 | 0.4546 | Yes |
| 19 | C1QA | C1QA Entrez, Source | complement component 1, q subcomponent, A chain | 699 | 0.236 | 0.4606 | Yes |
| 20 | C1QB | C1QB Entrez, Source | complement component 1, q subcomponent, B chain | 786 | 0.217 | 0.4662 | Yes |
| 21 | SEPP1 | SEPP1 Entrez, Source | selenoprotein P, plasma, 1 | 823 | 0.209 | 0.4750 | Yes |
| 22 | NGFRAP1 | NGFRAP1 Entrez, Source | nerve growth factor receptor (TNFRSF16) associated protein 1 | 830 | 0.208 | 0.4861 | Yes |
| 23 | MAFB | MAFB Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog B (avian) | 856 | 0.203 | 0.4955 | Yes |
| 24 | IER3 | IER3 Entrez, Source | immediate early response 3 | 908 | 0.195 | 0.5024 | Yes |
| 25 | SORT1 | SORT1 Entrez, Source | sortilin 1 | 989 | 0.183 | 0.5065 | Yes |
| 26 | QPRT | QPRT Entrez, Source | quinolinate phosphoribosyltransferase (nicotinate-nucleotide pyrophosphorylase (carboxylating)) | 1002 | 0.182 | 0.5157 | Yes |
| 27 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 1031 | 0.178 | 0.5235 | Yes |
| 28 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 1105 | 0.170 | 0.5274 | Yes |
| 29 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 1627 | 0.127 | 0.4951 | No |
| 30 | CTSD | CTSD Entrez, Source | cathepsin D (lysosomal aspartyl peptidase) | 1766 | 0.118 | 0.4912 | No |
| 31 | EREG | EREG Entrez, Source | epiregulin | 1968 | 0.109 | 0.4821 | No |
| 32 | TBXAS1 | TBXAS1 Entrez, Source | thromboxane A synthase 1 (platelet, cytochrome P450, family 5, subfamily A) | 2089 | 0.104 | 0.4788 | No |
| 33 | TLR4 | TLR4 Entrez, Source | toll-like receptor 4 | 2232 | 0.098 | 0.4735 | No |
| 34 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 2261 | 0.097 | 0.4767 | No |
| 35 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 2595 | 0.084 | 0.4562 | No |
| 36 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 2618 | 0.083 | 0.4592 | No |
| 37 | DPEP2 | DPEP2 Entrez, Source | dipeptidase 2 | 2868 | 0.075 | 0.4445 | No |
| 38 | HNMT | HNMT Entrez, Source | histamine N-methyltransferase | 2986 | 0.072 | 0.4396 | No |
| 39 | CAST | CAST Entrez, Source | calpastatin | 3052 | 0.070 | 0.4386 | No |
| 40 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 3128 | 0.068 | 0.4367 | No |
| 41 | SNX10 | SNX10 Entrez, Source | sorting nexin 10 | 3407 | 0.060 | 0.4190 | No |
| 42 | SMPDL3A | SMPDL3A Entrez, Source | sphingomyelin phosphodiesterase, acid-like 3A | 3776 | 0.051 | 0.3940 | No |
| 43 | PLA2G4A | PLA2G4A Entrez, Source | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 3785 | 0.050 | 0.3962 | No |
| 44 | ADCY2 | ADCY2 Entrez, Source | adenylate cyclase 2 (brain) | 4158 | 0.041 | 0.3704 | No |
| 45 | CAMK1 | CAMK1 Entrez, Source | calcium/calmodulin-dependent protein kinase I | 4262 | 0.038 | 0.3647 | No |
| 46 | NFKB2 | NFKB2 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 2 (p49/p100) | 4395 | 0.036 | 0.3567 | No |
| 47 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 4454 | 0.035 | 0.3543 | No |
| 48 | C3AR1 | C3AR1 Entrez, Source | complement component 3a receptor 1 | 4875 | 0.027 | 0.3240 | No |
| 49 | SCPEP1 | SCPEP1 Entrez, Source | serine carboxypeptidase 1 | 4916 | 0.026 | 0.3224 | No |
| 50 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 5002 | 0.024 | 0.3174 | No |
| 51 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 5285 | 0.019 | 0.2971 | No |
| 52 | CLU | CLU Entrez, Source | clusterin | 5443 | 0.016 | 0.2861 | No |
| 53 | LGALS3BP | LGALS3BP Entrez, Source | lectin, galactoside-binding, soluble, 3 binding protein | 5525 | 0.014 | 0.2808 | No |
| 54 | MMP2 | MMP2 Entrez, Source | matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) | 5540 | 0.014 | 0.2805 | No |
| 55 | PELI1 | PELI1 Entrez, Source | pellino homolog 1 (Drosophila) | 5792 | 0.009 | 0.2621 | No |
| 56 | PPBP | PPBP Entrez, Source | pro-platelet basic protein (chemokine (C-X-C motif) ligand 7) | 5909 | 0.007 | 0.2537 | No |
| 57 | CX3CR1 | CX3CR1 Entrez, Source | chemokine (C-X3-C motif) receptor 1 | 5914 | 0.007 | 0.2538 | No |
| 58 | AQP9 | AQP9 Entrez, Source | aquaporin 9 | 5937 | 0.006 | 0.2525 | No |
| 59 | KCNJ2 | KCNJ2 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 2 | 6048 | 0.004 | 0.2444 | No |
| 60 | MVP | MVP Entrez, Source | major vault protein | 6049 | 0.004 | 0.2446 | No |
| 61 | SERPINA1 | SERPINA1 Entrez, Source | serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 1 | 6063 | 0.004 | 0.2438 | No |
| 62 | TNF | TNF Entrez, Source | tumor necrosis factor (TNF superfamily, member 2) | 6181 | 0.001 | 0.2351 | No |
| 63 | C5AR1 | C5AR1 Entrez, Source | complement component 5a receptor 1 | 6304 | -0.000 | 0.2259 | No |
| 64 | SERPINB2 | SERPINB2 Entrez, Source | serpin peptidase inhibitor, clade B (ovalbumin), member 2 | 6630 | -0.006 | 0.2017 | No |
| 65 | PLEK | PLEK Entrez, Source | pleckstrin | 6970 | -0.012 | 0.1767 | No |
| 66 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 7236 | -0.017 | 0.1576 | No |
| 67 | CTSL | CTSL Entrez, Source | cathepsin L | 7390 | -0.020 | 0.1472 | No |
| 68 | HIP1 | HIP1 Entrez, Source | huntingtin interacting protein 1 | 7430 | -0.020 | 0.1453 | No |
| 69 | PTAFR | PTAFR Entrez, Source | platelet-activating factor receptor | 7629 | -0.024 | 0.1317 | No |
| 70 | TRIB1 | TRIB1 Entrez, Source | tribbles homolog 1 (Drosophila) | 7632 | -0.024 | 0.1329 | No |
| 71 | CD86 | CD86 Entrez, Source | CD86 molecule | 7686 | -0.025 | 0.1303 | No |
| 72 | PRKCD | PRKCD Entrez, Source | protein kinase C, delta | 7739 | -0.026 | 0.1278 | No |
| 73 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 7897 | -0.029 | 0.1176 | No |
| 74 | PDGFD | PDGFD Entrez, Source | platelet derived growth factor D | 7976 | -0.031 | 0.1134 | No |
| 75 | SECTM1 | SECTM1 Entrez, Source | secreted and transmembrane 1 | 7983 | -0.031 | 0.1146 | No |
| 76 | CARD9 | CARD9 Entrez, Source | caspase recruitment domain family, member 9 | 8328 | -0.038 | 0.0908 | No |
| 77 | HCK | HCK Entrez, Source | hemopoietic cell kinase | 8632 | -0.044 | 0.0703 | No |
| 78 | CAT | CAT Entrez, Source | catalase | 8873 | -0.049 | 0.0549 | No |
| 79 | HOXA4 | HOXA4 Entrez, Source | homeobox A4 | 9080 | -0.054 | 0.0423 | No |
| 80 | IGF2R | IGF2R Entrez, Source | insulin-like growth factor 2 receptor | 9465 | -0.062 | 0.0167 | No |
| 81 | SLC15A3 | SLC15A3 Entrez, Source | solute carrier family 15, member 3 | 9719 | -0.069 | 0.0014 | No |
| 82 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 9721 | -0.069 | 0.0052 | No |
| 83 | C2 | C2 Entrez, Source | complement component 2 | 10542 | -0.091 | -0.0517 | No |
| 84 | LAPTM4B | LAPTM4B Entrez, Source | lysosomal associated protein transmembrane 4 beta | 10724 | -0.098 | -0.0600 | No |
| 85 | EMR1 | EMR1 Entrez, Source | egf-like module containing, mucin-like, hormone receptor-like 1 | 10780 | -0.100 | -0.0586 | No |
| 86 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 10956 | -0.107 | -0.0659 | No |
| 87 | RHAG | RHAG Entrez, Source | Rh-associated glycoprotein | 10996 | -0.108 | -0.0628 | No |
| 88 | CCL4 | CCL4 Entrez, Source | chemokine (C-C motif) ligand 4 | 11087 | -0.111 | -0.0634 | No |
| 89 | TCIRG1 | TCIRG1 Entrez, Source | T-cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 subunit A3 | 11833 | -0.147 | -0.1116 | No |
| 90 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 11843 | -0.147 | -0.1041 | No |
| 91 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 11883 | -0.149 | -0.0988 | No |
| 92 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 11961 | -0.154 | -0.0961 | No |
| 93 | SLC16A3 | SLC16A3 Entrez, Source | solute carrier family 16, member 3 (monocarboxylic acid transporter 4) | 12008 | -0.158 | -0.0908 | No |
| 94 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 12088 | -0.163 | -0.0877 | No |
| 95 | HOMER3 | HOMER3 Entrez, Source | homer homolog 3 (Drosophila) | 12210 | -0.172 | -0.0873 | No |
| 96 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 12298 | -0.179 | -0.0839 | No |
| 97 | CD14 | CD14 Entrez, Source | CD14 molecule | 12462 | -0.194 | -0.0855 | No |
| 98 | LMNA | LMNA Entrez, Source | lamin A/C | 12500 | -0.199 | -0.0772 | No |
| 99 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 12547 | -0.204 | -0.0694 | No |
| 100 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 12851 | -0.254 | -0.0782 | No |
| 101 | NINJ1 | NINJ1 Entrez, Source | ninjurin 1 | 12892 | -0.265 | -0.0665 | No |
| 102 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 13097 | -0.326 | -0.0639 | No |
| 103 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 13228 | -0.440 | -0.0493 | No |
| 104 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 13267 | -0.519 | -0.0234 | No |
| 105 | ABCC4 | ABCC4 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 4 | 13272 | -0.523 | 0.0053 | No |