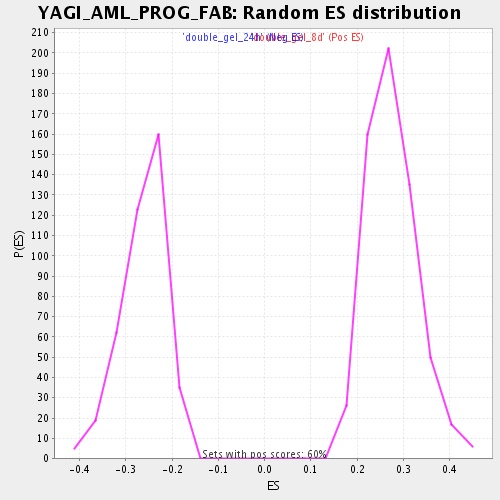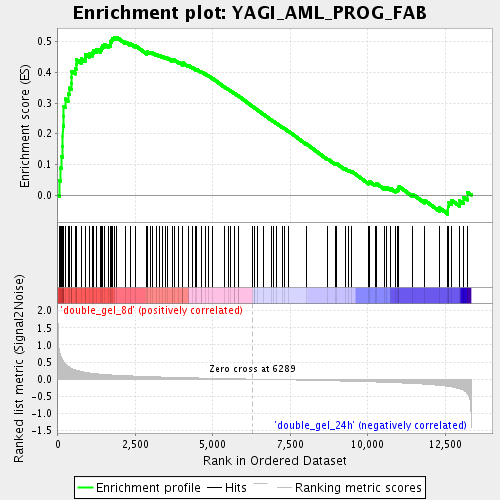
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | YAGI_AML_PROG_FAB |
| Enrichment Score (ES) | 0.51457185 |
| Normalized Enrichment Score (NES) | 1.8721703 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.016760241 |
| FWER p-Value | 0.679 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TIMP2 | TIMP2 Entrez, Source | TIMP metallopeptidase inhibitor 2 | 49 | 0.810 | 0.0475 | Yes |
| 2 | LGALS1 | LGALS1 Entrez, Source | lectin, galactoside-binding, soluble, 1 (galectin 1) | 82 | 0.687 | 0.0885 | Yes |
| 3 | F2R | F2R Entrez, Source | coagulation factor II (thrombin) receptor | 114 | 0.625 | 0.1257 | Yes |
| 4 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 156 | 0.554 | 0.1576 | Yes |
| 5 | SFXN3 | SFXN3 Entrez, Source | sideroflexin 3 | 162 | 0.546 | 0.1917 | Yes |
| 6 | AHNAK | AHNAK Entrez, Source | AHNAK nucleoprotein (desmoyokin) | 164 | 0.541 | 0.2259 | Yes |
| 7 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 185 | 0.510 | 0.2566 | Yes |
| 8 | SERPINE2 | SERPINE2 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 | 190 | 0.499 | 0.2878 | Yes |
| 9 | SERPINH1 | SERPINH1 Entrez, Source | serpin peptidase inhibitor, clade H (heat shock protein 47), member 1, (collagen binding protein 1) | 233 | 0.457 | 0.3135 | Yes |
| 10 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 330 | 0.373 | 0.3299 | Yes |
| 11 | LAMA5 | LAMA5 Entrez, Source | laminin, alpha 5 | 368 | 0.352 | 0.3493 | Yes |
| 12 | ELOVL6 | ELOVL6 Entrez, Source | ELOVL family member 6, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 433 | 0.319 | 0.3646 | Yes |
| 13 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 442 | 0.313 | 0.3838 | Yes |
| 14 | SQLE | SQLE Entrez, Source | squalene epoxidase | 446 | 0.311 | 0.4033 | Yes |
| 15 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 561 | 0.267 | 0.4115 | Yes |
| 16 | PKIA | PKIA Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor alpha | 588 | 0.260 | 0.4260 | Yes |
| 17 | ALDH5A1 | ALDH5A1 Entrez, Source | aldehyde dehydrogenase 5 family, member A1 (succinate-semialdehyde dehydrogenase) | 595 | 0.258 | 0.4418 | Yes |
| 18 | IFNGR1 | IFNGR1 Entrez, Source | interferon gamma receptor 1 | 750 | 0.226 | 0.4445 | Yes |
| 19 | IL2RG | IL2RG Entrez, Source | interleukin 2 receptor, gamma (severe combined immunodeficiency) | 879 | 0.198 | 0.4473 | Yes |
| 20 | SCP2 | SCP2 Entrez, Source | sterol carrier protein 2 | 890 | 0.197 | 0.4590 | Yes |
| 21 | HDGFRP3 | HDGFRP3 Entrez, Source | - | 1011 | 0.181 | 0.4614 | Yes |
| 22 | PKIG | PKIG Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor gamma | 1121 | 0.169 | 0.4638 | Yes |
| 23 | PCCB | PCCB Entrez, Source | propionyl Coenzyme A carboxylase, beta polypeptide | 1152 | 0.165 | 0.4720 | Yes |
| 24 | CXADR | CXADR Entrez, Source | coxsackie virus and adenovirus receptor | 1241 | 0.157 | 0.4753 | Yes |
| 25 | ANXA5 | ANXA5 Entrez, Source | annexin A5 | 1367 | 0.146 | 0.4750 | Yes |
| 26 | RP11-54H7.1 | RP11-54H7.1 Entrez, Source | - | 1418 | 0.141 | 0.4802 | Yes |
| 27 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 1445 | 0.139 | 0.4870 | Yes |
| 28 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 1507 | 0.134 | 0.4909 | Yes |
| 29 | CYB5B | CYB5B Entrez, Source | cytochrome b5 type B (outer mitochondrial membrane) | 1646 | 0.126 | 0.4884 | Yes |
| 30 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 1691 | 0.123 | 0.4928 | Yes |
| 31 | DEAF1 | DEAF1 Entrez, Source | deformed epidermal autoregulatory factor 1 (Drosophila) | 1704 | 0.122 | 0.4997 | Yes |
| 32 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 1732 | 0.121 | 0.5053 | Yes |
| 33 | MANBA | MANBA Entrez, Source | mannosidase, beta A, lysosomal | 1761 | 0.119 | 0.5107 | Yes |
| 34 | TM7SF2 | TM7SF2 Entrez, Source | transmembrane 7 superfamily member 2 | 1828 | 0.115 | 0.5130 | Yes |
| 35 | PROS1 | PROS1 Entrez, Source | protein S (alpha) | 1901 | 0.112 | 0.5146 | Yes |
| 36 | NUP50 | NUP50 Entrez, Source | nucleoporin 50kDa | 2190 | 0.100 | 0.4991 | No |
| 37 | TRPC6 | TRPC6 Entrez, Source | transient receptor potential cation channel, subfamily C, member 6 | 2326 | 0.094 | 0.4949 | No |
| 38 | MAX | MAX Entrez, Source | MYC associated factor X | 2495 | 0.088 | 0.4877 | No |
| 39 | ICAM2 | ICAM2 Entrez, Source | intercellular adhesion molecule 2 | 2873 | 0.075 | 0.4640 | No |
| 40 | CAPG | CAPG Entrez, Source | capping protein (actin filament), gelsolin-like | 2891 | 0.074 | 0.4674 | No |
| 41 | COL11A1 | COL11A1 Entrez, Source | collagen, type XI, alpha 1 | 2981 | 0.072 | 0.4652 | No |
| 42 | CAST | CAST Entrez, Source | calpastatin | 3052 | 0.070 | 0.4643 | No |
| 43 | TPST2 | TPST2 Entrez, Source | tyrosylprotein sulfotransferase 2 | 3182 | 0.066 | 0.4588 | No |
| 44 | CTSW | CTSW Entrez, Source | cathepsin W (lymphopain) | 3289 | 0.063 | 0.4547 | No |
| 45 | GIPC1 | GIPC1 Entrez, Source | GIPC PDZ domain containing family, member 1 | 3372 | 0.061 | 0.4524 | No |
| 46 | ACO1 | ACO1 Entrez, Source | aconitase 1, soluble | 3462 | 0.059 | 0.4494 | No |
| 47 | GOSR2 | GOSR2 Entrez, Source | golgi SNAP receptor complex member 2 | 3541 | 0.056 | 0.4470 | No |
| 48 | GP9 | GP9 Entrez, Source | glycoprotein IX (platelet) | 3686 | 0.053 | 0.4395 | No |
| 49 | STAT4 | STAT4 Entrez, Source | signal transducer and activator of transcription 4 | 3697 | 0.052 | 0.4420 | No |
| 50 | PIR | PIR Entrez, Source | pirin (iron-binding nuclear protein) | 3758 | 0.051 | 0.4407 | No |
| 51 | PTPN4 | PTPN4 Entrez, Source | protein tyrosine phosphatase, non-receptor type 4 (megakaryocyte) | 3901 | 0.047 | 0.4329 | No |
| 52 | TRPC1 | TRPC1 Entrez, Source | transient receptor potential cation channel, subfamily C, member 1 | 4033 | 0.044 | 0.4258 | No |
| 53 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 4035 | 0.044 | 0.4285 | No |
| 54 | ELOVL5 | ELOVL5 Entrez, Source | ELOVL family member 5, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4036 | 0.044 | 0.4313 | No |
| 55 | CFH | CFH Entrez, Source | complement factor H | 4209 | 0.039 | 0.4207 | No |
| 56 | GP1BB /// SEPT5 | 4214 | 0.039 | 0.4229 | No | ||
| 57 | ELL2 | ELL2 Entrez, Source | elongation factor, RNA polymerase II, 2 | 4354 | 0.037 | 0.4147 | No |
| 58 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 4454 | 0.035 | 0.4095 | No |
| 59 | SLC25A1 | SLC25A1 Entrez, Source | solute carrier family 25 (mitochondrial carrier; citrate transporter), member 1 | 4469 | 0.035 | 0.4106 | No |
| 60 | S100A10 | S100A10 Entrez, Source | S100 calcium binding protein A10 | 4630 | 0.031 | 0.4005 | No |
| 61 | SC5DL | SC5DL Entrez, Source | sterol-C5-desaturase (ERG3 delta-5-desaturase homolog, fungal)-like | 4644 | 0.031 | 0.4015 | No |
| 62 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 4753 | 0.029 | 0.3951 | No |
| 63 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 4845 | 0.027 | 0.3900 | No |
| 64 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 5002 | 0.024 | 0.3797 | No |
| 65 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 5378 | 0.017 | 0.3524 | No |
| 66 | CD164 | CD164 Entrez, Source | CD164 molecule, sialomucin | 5509 | 0.014 | 0.3435 | No |
| 67 | SERPINI1 | SERPINI1 Entrez, Source | serpin peptidase inhibitor, clade I (neuroserpin), member 1 | 5563 | 0.013 | 0.3404 | No |
| 68 | GJA4 | GJA4 Entrez, Source | gap junction protein, alpha 4, 37kDa (connexin 37) | 5689 | 0.011 | 0.3316 | No |
| 69 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 5694 | 0.011 | 0.3320 | No |
| 70 | KIF2A | KIF2A Entrez, Source | kinesin heavy chain member 2A | 5813 | 0.009 | 0.3237 | No |
| 71 | TNFRSF4 | TNFRSF4 Entrez, Source | tumor necrosis factor receptor superfamily, member 4 | 5838 | 0.008 | 0.3224 | No |
| 72 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 6293 | -0.000 | 0.2881 | No |
| 73 | ZBTB16 | ZBTB16 Entrez, Source | zinc finger and BTB domain containing 16 | 6337 | -0.001 | 0.2849 | No |
| 74 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 6349 | -0.001 | 0.2841 | No |
| 75 | CAPN2 | CAPN2 Entrez, Source | calpain 2, (m/II) large subunit | 6436 | -0.002 | 0.2777 | No |
| 76 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 6645 | -0.006 | 0.2624 | No |
| 77 | NCAM1 | NCAM1 Entrez, Source | neural cell adhesion molecule 1 | 6887 | -0.010 | 0.2449 | No |
| 78 | GATA2 | GATA2 Entrez, Source | GATA binding protein 2 | 6947 | -0.011 | 0.2411 | No |
| 79 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 7051 | -0.013 | 0.2342 | No |
| 80 | DDB1 | DDB1 Entrez, Source | damage-specific DNA binding protein 1, 127kDa | 7256 | -0.017 | 0.2198 | No |
| 81 | LCP1 | LCP1 Entrez, Source | lymphocyte cytosolic protein 1 (L-plastin) | 7308 | -0.018 | 0.2171 | No |
| 82 | UNC119 | UNC119 Entrez, Source | unc-119 homolog (C. elegans) | 7443 | -0.021 | 0.2083 | No |
| 83 | CDC42BPA | CDC42BPA Entrez, Source | CDC42 binding protein kinase alpha (DMPK-like) | 8008 | -0.031 | 0.1676 | No |
| 84 | CTR9 | CTR9 Entrez, Source | Ctr9, Paf1/RNA polymerase II complex component, homolog (S. cerevisiae) | 8711 | -0.046 | 0.1175 | No |
| 85 | GIT2 | GIT2 Entrez, Source | G protein-coupled receptor kinase interactor 2 | 8964 | -0.051 | 0.1017 | No |
| 86 | NTRK1 | NTRK1 Entrez, Source | neurotrophic tyrosine kinase, receptor, type 1 | 8992 | -0.052 | 0.1029 | No |
| 87 | GAD1 | GAD1 Entrez, Source | glutamate decarboxylase 1 (brain, 67kDa) | 9270 | -0.057 | 0.0856 | No |
| 88 | CD200 | CD200 Entrez, Source | CD200 molecule | 9392 | -0.060 | 0.0802 | No |
| 89 | PLA2G6 | PLA2G6 Entrez, Source | phospholipase A2, group VI (cytosolic, calcium-independent) | 9476 | -0.062 | 0.0779 | No |
| 90 | TJP2 | TJP2 Entrez, Source | tight junction protein 2 (zona occludens 2) | 10032 | -0.077 | 0.0409 | No |
| 91 | MORF4L2 | MORF4L2 Entrez, Source | mortality factor 4 like 2 | 10054 | -0.078 | 0.0442 | No |
| 92 | SLC7A1 | SLC7A1 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 | 10239 | -0.083 | 0.0355 | No |
| 93 | MLLT11 | MLLT11 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 11 | 10269 | -0.083 | 0.0386 | No |
| 94 | AMD1 | AMD1 Entrez, Source | adenosylmethionine decarboxylase 1 | 10528 | -0.091 | 0.0249 | No |
| 95 | APPBP1 | APPBP1 Entrez, Source | amyloid beta precursor protein binding protein 1 | 10606 | -0.094 | 0.0250 | No |
| 96 | LAPTM4B | LAPTM4B Entrez, Source | lysosomal associated protein transmembrane 4 beta | 10724 | -0.098 | 0.0223 | No |
| 97 | GABRE | GABRE Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, epsilon | 10891 | -0.105 | 0.0164 | No |
| 98 | TGM2 | TGM2 Entrez, Source | transglutaminase 2 (C polypeptide, protein-glutamine-gamma-glutamyltransferase) | 10955 | -0.107 | 0.0184 | No |
| 99 | RHAG | RHAG Entrez, Source | Rh-associated glycoprotein | 10996 | -0.108 | 0.0222 | No |
| 100 | SNTA1 | SNTA1 Entrez, Source | syntrophin, alpha 1 (dystrophin-associated protein A1, 59kDa, acidic component) | 11006 | -0.108 | 0.0284 | No |
| 101 | PCNA | PCNA Entrez, Source | proliferating cell nuclear antigen | 11449 | -0.127 | 0.0030 | No |
| 102 | TCIRG1 | TCIRG1 Entrez, Source | T-cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 subunit A3 | 11833 | -0.147 | -0.0167 | No |
| 103 | NAP1L3 | NAP1L3 Entrez, Source | nucleosome assembly protein 1-like 3 | 12307 | -0.180 | -0.0410 | No |
| 104 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 12585 | -0.209 | -0.0488 | No |
| 105 | PRKCBP1 | PRKCBP1 Entrez, Source | protein kinase C binding protein 1 | 12589 | -0.209 | -0.0358 | No |
| 106 | GGH | GGH Entrez, Source | gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) | 12617 | -0.213 | -0.0244 | No |
| 107 | RFC3 | RFC3 Entrez, Source | replication factor C (activator 1) 3, 38kDa | 12712 | -0.226 | -0.0172 | No |
| 108 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 12967 | -0.282 | -0.0186 | No |
| 109 | PAFAH1B3 | PAFAH1B3 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, gamma subunit 29kDa | 13095 | -0.326 | -0.0076 | No |
| 110 | CD68 | CD68 Entrez, Source | CD68 molecule | 13210 | -0.414 | 0.0100 | No |