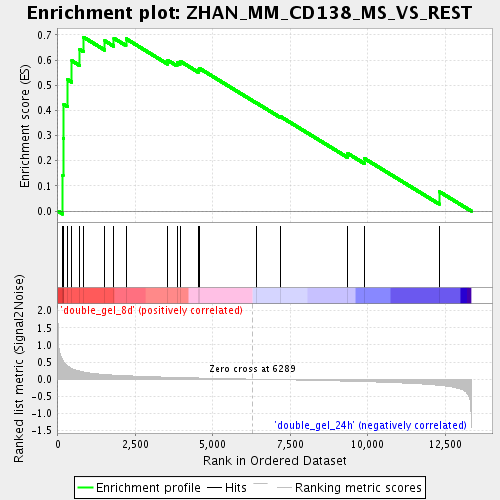
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_24h.class.cls #double_gel_8d_versus_double_gel_24h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_24h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | ZHAN_MM_CD138_MS_VS_REST |
| Enrichment Score (ES) | 0.69038284 |
| Normalized Enrichment Score (NES) | 1.803949 |
| Nominal p-value | 0.0055452865 |
| FDR q-value | 0.02761913 |
| FWER p-Value | 0.932 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 156 | 0.554 | 0.1412 | Yes |
| 2 | TMEM176B | TMEM176B Entrez, Source | transmembrane protein 176B | 168 | 0.535 | 0.2880 | Yes |
| 3 | SERPINE2 | SERPINE2 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 | 190 | 0.499 | 0.4240 | Yes |
| 4 | SPRY2 | SPRY2 Entrez, Source | sprouty homolog 2 (Drosophila) | 314 | 0.388 | 0.5219 | Yes |
| 5 | KLHL13 | KLHL13 Entrez, Source | kelch-like 13 (Drosophila) | 449 | 0.309 | 0.5970 | Yes |
| 6 | JAM3 | JAM3 Entrez, Source | junctional adhesion molecule 3 | 704 | 0.234 | 0.6425 | Yes |
| 7 | NGFRAP1 | NGFRAP1 Entrez, Source | nerve growth factor receptor (TNFRSF16) associated protein 1 | 830 | 0.208 | 0.6904 | Yes |
| 8 | CST3 | CST3 Entrez, Source | cystatin C (amyloid angiopathy and cerebral hemorrhage) | 1513 | 0.134 | 0.6762 | No |
| 9 | CLIC6 | CLIC6 Entrez, Source | chloride intracellular channel 6 | 1807 | 0.117 | 0.6864 | No |
| 10 | RNF32 | RNF32 Entrez, Source | ring finger protein 32 | 2198 | 0.100 | 0.6846 | No |
| 11 | SLC22A17 | SLC22A17 Entrez, Source | solute carrier family 22 (organic cation transporter), member 17 | 3545 | 0.056 | 0.5990 | No |
| 12 | MYRIP | MYRIP Entrez, Source | myosin VIIA and Rab interacting protein | 3846 | 0.049 | 0.5900 | No |
| 13 | MARK1 | MARK1 Entrez, Source | MAP/microtubule affinity-regulating kinase 1 | 3942 | 0.046 | 0.5955 | No |
| 14 | FZD2 | FZD2 Entrez, Source | frizzled homolog 2 (Drosophila) | 4530 | 0.033 | 0.5607 | No |
| 15 | TEAD1 | TEAD1 Entrez, Source | TEA domain family member 1 (SV40 transcriptional enhancer factor) | 4567 | 0.033 | 0.5670 | No |
| 16 | BACE2 | BACE2 Entrez, Source | beta-site APP-cleaving enzyme 2 | 6423 | -0.002 | 0.4283 | No |
| 17 | CLEC11A | CLEC11A Entrez, Source | C-type lectin domain family 11, member A | 7186 | -0.016 | 0.3754 | No |
| 18 | PPM1H | PPM1H Entrez, Source | protein phosphatase 1H (PP2C domain containing) | 9352 | -0.059 | 0.2293 | No |
| 19 | PCSK5 | PCSK5 Entrez, Source | proprotein convertase subtilisin/kexin type 5 | 9890 | -0.073 | 0.2093 | No |
| 20 | NAP1L3 | NAP1L3 Entrez, Source | nucleosome assembly protein 1-like 3 | 12307 | -0.180 | 0.0777 | No |

