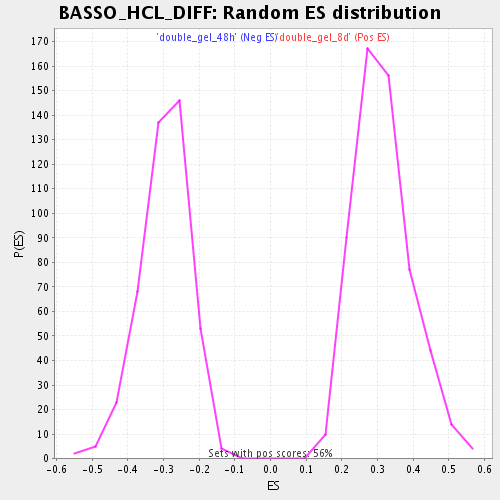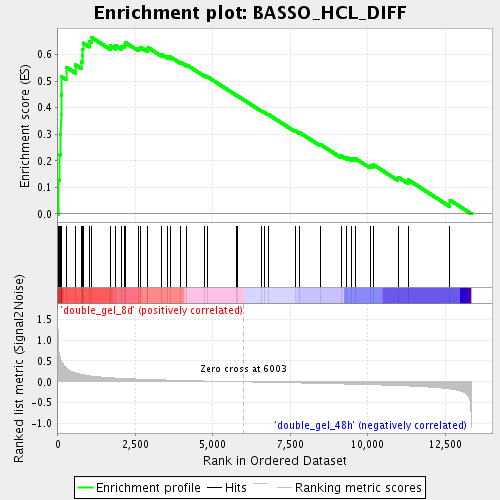
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | BASSO_HCL_DIFF |
| Enrichment Score (ES) | 0.6656083 |
| Normalized Enrichment Score (NES) | 2.1207333 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0010423323 |
| FWER p-Value | 0.023 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 11 | 0.884 | 0.1285 | Yes |
| 2 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 43 | 0.675 | 0.2249 | Yes |
| 3 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 89 | 0.531 | 0.2992 | Yes |
| 4 | TACC1 | TACC1 Entrez, Source | transforming, acidic coiled-coil containing protein 1 | 99 | 0.518 | 0.3743 | Yes |
| 5 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 101 | 0.510 | 0.4488 | Yes |
| 6 | DPYD | DPYD Entrez, Source | dihydropyrimidine dehydrogenase | 123 | 0.473 | 0.5165 | Yes |
| 7 | PRSS23 | PRSS23 Entrez, Source | protease, serine, 23 | 273 | 0.325 | 0.5528 | Yes |
| 8 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 571 | 0.212 | 0.5614 | Yes |
| 9 | TIMP1 | TIMP1 Entrez, Source | TIMP metallopeptidase inhibitor 1 | 759 | 0.176 | 0.5731 | Yes |
| 10 | SPRY2 | SPRY2 Entrez, Source | sprouty homolog 2 (Drosophila) | 784 | 0.172 | 0.5965 | Yes |
| 11 | TRIM2 | TRIM2 Entrez, Source | tripartite motif-containing 2 | 802 | 0.169 | 0.6200 | Yes |
| 12 | PTTG1IP | PTTG1IP Entrez, Source | pituitary tumor-transforming 1 interacting protein | 811 | 0.168 | 0.6440 | Yes |
| 13 | IL3RA | IL3RA Entrez, Source | interleukin 3 receptor, alpha (low affinity) | 1013 | 0.146 | 0.6502 | Yes |
| 14 | PDE4DIP | PDE4DIP Entrez, Source | phosphodiesterase 4D interacting protein (myomegalin) | 1077 | 0.138 | 0.6656 | Yes |
| 15 | GABARAPL2 | GABARAPL2 Entrez, Source | GABA(A) receptor-associated protein-like 2 | 1687 | 0.098 | 0.6341 | No |
| 16 | SMPDL3A | SMPDL3A Entrez, Source | sphingomyelin phosphodiesterase, acid-like 3A | 1863 | 0.090 | 0.6341 | No |
| 17 | PAWR | PAWR Entrez, Source | PRKC, apoptosis, WT1, regulator | 2056 | 0.083 | 0.6318 | No |
| 18 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 2137 | 0.079 | 0.6374 | No |
| 19 | MAPK10 | MAPK10 Entrez, Source | mitogen-activated protein kinase 10 | 2176 | 0.078 | 0.6460 | No |
| 20 | MYF6 | MYF6 Entrez, Source | myogenic factor 6 (herculin) | 2600 | 0.065 | 0.6237 | No |
| 21 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 2668 | 0.063 | 0.6279 | No |
| 22 | HBB | HBB Entrez, Source | hemoglobin, beta | 2898 | 0.057 | 0.6191 | No |
| 23 | VAMP3 | VAMP3 Entrez, Source | vesicle-associated membrane protein 3 (cellubrevin) | 2906 | 0.057 | 0.6269 | No |
| 24 | IL18 | IL18 Entrez, Source | interleukin 18 (interferon-gamma-inducing factor) | 3347 | 0.046 | 0.6006 | No |
| 25 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 3540 | 0.042 | 0.5923 | No |
| 26 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 3628 | 0.040 | 0.5916 | No |
| 27 | GAS7 | GAS7 Entrez, Source | growth arrest-specific 7 | 3956 | 0.034 | 0.5720 | No |
| 28 | RAB13 | RAB13 Entrez, Source | RAB13, member RAS oncogene family | 4159 | 0.030 | 0.5611 | No |
| 29 | SIX3 | SIX3 Entrez, Source | sine oculis homeobox homolog 3 (Drosophila) | 4743 | 0.019 | 0.5201 | No |
| 30 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 4825 | 0.018 | 0.5167 | No |
| 31 | ENG | ENG Entrez, Source | endoglin (Osler-Rendu-Weber syndrome 1) | 5761 | 0.003 | 0.4468 | No |
| 32 | B2M | B2M Entrez, Source | beta-2-microglobulin | 5795 | 0.003 | 0.4447 | No |
| 33 | BLR1 | BLR1 Entrez, Source | Burkitt lymphoma receptor 1, GTP binding protein (chemokine (C-X-C motif) receptor 5) | 6582 | -0.009 | 0.3869 | No |
| 34 | PTPRM | PTPRM Entrez, Source | protein tyrosine phosphatase, receptor type, M | 6655 | -0.010 | 0.3829 | No |
| 35 | SCN1B | SCN1B Entrez, Source | sodium channel, voltage-gated, type I, beta | 6793 | -0.012 | 0.3744 | No |
| 36 | NOVA1 | NOVA1 Entrez, Source | neuro-oncological ventral antigen 1 | 7658 | -0.025 | 0.3131 | No |
| 37 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 7788 | -0.027 | 0.3074 | No |
| 38 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 8465 | -0.038 | 0.2622 | No |
| 39 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 9141 | -0.050 | 0.2187 | No |
| 40 | SDC3 | SDC3 Entrez, Source | syndecan 3 (N-syndecan) | 9311 | -0.053 | 0.2137 | No |
| 41 | TBKBP1 | TBKBP1 Entrez, Source | TBK1 binding protein 1 | 9480 | -0.056 | 0.2092 | No |
| 42 | RPLP1 | RPLP1 Entrez, Source | ribosomal protein, large, P1 | 9602 | -0.058 | 0.2085 | No |
| 43 | CD63 | CD63 Entrez, Source | CD63 molecule | 10073 | -0.068 | 0.1831 | No |
| 44 | RTN2 | RTN2 Entrez, Source | reticulon 2 | 10170 | -0.070 | 0.1861 | No |
| 45 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 10976 | -0.090 | 0.1387 | No |
| 46 | RCBTB2 | RCBTB2 Entrez, Source | regulator of chromosome condensation (RCC1) and BTB (POZ) domain containing protein 2 | 11303 | -0.099 | 0.1286 | No |
| 47 | PPIC | PPIC Entrez, Source | peptidylprolyl isomerase C (cyclophilin C) | 12653 | -0.168 | 0.0518 | No |