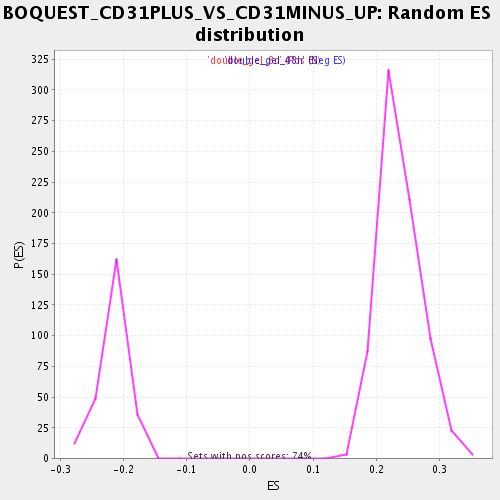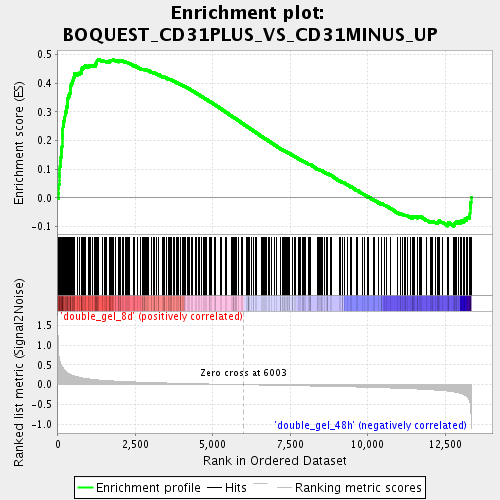
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | BOQUEST_CD31PLUS_VS_CD31MINUS_UP |
| Enrichment Score (ES) | 0.4836616 |
| Normalized Enrichment Score (NES) | 2.0459166 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0025709972 |
| FWER p-Value | 0.095 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | MAP3K8 | MAP3K8 Entrez, Source | mitogen-activated protein kinase kinase kinase 8 | 12 | 0.878 | 0.0172 | Yes |
| 2 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 25 | 0.771 | 0.0322 | Yes |
| 3 | HLA-DMA | HLA-DMA Entrez, Source | major histocompatibility complex, class II, DM alpha | 32 | 0.725 | 0.0467 | Yes |
| 4 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 43 | 0.675 | 0.0598 | Yes |
| 5 | LPL | LPL Entrez, Source | lipoprotein lipase | 45 | 0.671 | 0.0736 | Yes |
| 6 | MGP | MGP Entrez, Source | matrix Gla protein | 52 | 0.634 | 0.0862 | Yes |
| 7 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 56 | 0.628 | 0.0989 | Yes |
| 8 | RAPGEF4 | RAPGEF4 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 4 | 66 | 0.600 | 0.1106 | Yes |
| 9 | PDK4 | PDK4 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 4 | 84 | 0.549 | 0.1206 | Yes |
| 10 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 87 | 0.537 | 0.1315 | Yes |
| 11 | RNASE1 | RNASE1 Entrez, Source | ribonuclease, RNase A family, 1 (pancreatic) | 94 | 0.529 | 0.1420 | Yes |
| 12 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 103 | 0.505 | 0.1518 | Yes |
| 13 | OMD | OMD Entrez, Source | osteomodulin | 109 | 0.490 | 0.1615 | Yes |
| 14 | TNFAIP6 | TNFAIP6 Entrez, Source | tumor necrosis factor, alpha-induced protein 6 | 116 | 0.483 | 0.1710 | Yes |
| 15 | GPR64 | GPR64 Entrez, Source | G protein-coupled receptor 64 | 131 | 0.466 | 0.1795 | Yes |
| 16 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 138 | 0.458 | 0.1885 | Yes |
| 17 | AGTR1 | AGTR1 Entrez, Source | angiotensin II receptor, type 1 | 139 | 0.457 | 0.1979 | Yes |
| 18 | SERPINE2 | SERPINE2 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 | 140 | 0.455 | 0.2073 | Yes |
| 19 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 145 | 0.452 | 0.2164 | Yes |
| 20 | APOC1 | APOC1 Entrez, Source | apolipoprotein C-I | 151 | 0.445 | 0.2251 | Yes |
| 21 | LOX | LOX Entrez, Source | lysyl oxidase | 154 | 0.441 | 0.2341 | Yes |
| 22 | PRKAR2B | PRKAR2B Entrez, Source | protein kinase, cAMP-dependent, regulatory, type II, beta | 160 | 0.434 | 0.2427 | Yes |
| 23 | BMP2 | BMP2 Entrez, Source | bone morphogenetic protein 2 | 164 | 0.430 | 0.2513 | Yes |
| 24 | PGCP | PGCP Entrez, Source | - | 183 | 0.404 | 0.2582 | Yes |
| 25 | ABCG1 | ABCG1 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 1 | 187 | 0.400 | 0.2663 | Yes |
| 26 | GPX3 | GPX3 Entrez, Source | glutathione peroxidase 3 (plasma) | 199 | 0.391 | 0.2735 | Yes |
| 27 | PRELP | PRELP Entrez, Source | proline/arginine-rich end leucine-rich repeat protein | 204 | 0.383 | 0.2811 | Yes |
| 28 | CFD | CFD Entrez, Source | complement factor D (adipsin) | 228 | 0.365 | 0.2868 | Yes |
| 29 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 231 | 0.364 | 0.2942 | Yes |
| 30 | PLA1A | PLA1A Entrez, Source | phospholipase A1 member A | 236 | 0.355 | 0.3012 | Yes |
| 31 | FMO3 | FMO3 Entrez, Source | flavin containing monooxygenase 3 | 281 | 0.322 | 0.3044 | Yes |
| 32 | CD9 | CD9 Entrez, Source | CD9 molecule | 283 | 0.320 | 0.3110 | Yes |
| 33 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 286 | 0.317 | 0.3173 | Yes |
| 34 | ENPEP | ENPEP Entrez, Source | glutamyl aminopeptidase (aminopeptidase A) | 298 | 0.310 | 0.3229 | Yes |
| 35 | CLIC2 | CLIC2 Entrez, Source | chloride intracellular channel 2 | 304 | 0.304 | 0.3288 | Yes |
| 36 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 310 | 0.299 | 0.3346 | Yes |
| 37 | CBS | CBS Entrez, Source | cystathionine-beta-synthase | 313 | 0.297 | 0.3406 | Yes |
| 38 | TFPI | TFPI Entrez, Source | tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) | 316 | 0.294 | 0.3465 | Yes |
| 39 | PTPRD | PTPRD Entrez, Source | protein tyrosine phosphatase, receptor type, D | 330 | 0.290 | 0.3514 | Yes |
| 40 | SLC5A3 | SLC5A3 Entrez, Source | solute carrier family 5 (inositol transporters), member 3 | 333 | 0.289 | 0.3572 | Yes |
| 41 | RAPGEF5 | RAPGEF5 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 5 | 384 | 0.267 | 0.3589 | Yes |
| 42 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 385 | 0.267 | 0.3644 | Yes |
| 43 | PTGER4 | PTGER4 Entrez, Source | prostaglandin E receptor 4 (subtype EP4) | 402 | 0.258 | 0.3685 | Yes |
| 44 | TMEM176B | TMEM176B Entrez, Source | transmembrane protein 176B | 403 | 0.258 | 0.3738 | Yes |
| 45 | CCNL1 | CCNL1 Entrez, Source | cyclin L1 | 405 | 0.257 | 0.3790 | Yes |
| 46 | FOXO1A | FOXO1A Entrez, Source | forkhead box O1A (rhabdomyosarcoma) | 411 | 0.256 | 0.3839 | Yes |
| 47 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 417 | 0.254 | 0.3888 | Yes |
| 48 | DDAH1 | DDAH1 Entrez, Source | dimethylarginine dimethylaminohydrolase 1 | 421 | 0.254 | 0.3938 | Yes |
| 49 | RAMP3 | RAMP3 Entrez, Source | receptor (calcitonin) activity modifying protein 3 | 429 | 0.251 | 0.3984 | Yes |
| 50 | RGS4 | RGS4 Entrez, Source | regulator of G-protein signalling 4 | 454 | 0.244 | 0.4016 | Yes |
| 51 | THBD | THBD Entrez, Source | thrombomodulin | 474 | 0.236 | 0.4050 | Yes |
| 52 | GSN | GSN Entrez, Source | gelsolin (amyloidosis, Finnish type) | 483 | 0.233 | 0.4092 | Yes |
| 53 | ANGPTL4 | ANGPTL4 Entrez, Source | angiopoietin-like 4 | 493 | 0.230 | 0.4133 | Yes |
| 54 | RGS5 | RGS5 Entrez, Source | regulator of G-protein signalling 5 | 513 | 0.224 | 0.4164 | Yes |
| 55 | VCAM1 | VCAM1 Entrez, Source | vascular cell adhesion molecule 1 | 516 | 0.223 | 0.4209 | Yes |
| 56 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 523 | 0.222 | 0.4250 | Yes |
| 57 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 538 | 0.219 | 0.4284 | Yes |
| 58 | ALCAM | ALCAM Entrez, Source | activated leukocyte cell adhesion molecule | 539 | 0.219 | 0.4329 | Yes |
| 59 | SCARB1 | SCARB1 Entrez, Source | scavenger receptor class B, member 1 | 620 | 0.203 | 0.4309 | Yes |
| 60 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 627 | 0.202 | 0.4346 | Yes |
| 61 | STX11 | STX11 Entrez, Source | syntaxin 11 | 699 | 0.186 | 0.4330 | Yes |
| 62 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 709 | 0.184 | 0.4361 | Yes |
| 63 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 754 | 0.176 | 0.4363 | Yes |
| 64 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 756 | 0.176 | 0.4399 | Yes |
| 65 | IL6R | IL6R Entrez, Source | interleukin 6 receptor | 761 | 0.176 | 0.4432 | Yes |
| 66 | CD320 | CD320 Entrez, Source | CD320 molecule | 770 | 0.174 | 0.4462 | Yes |
| 67 | EMP2 | EMP2 Entrez, Source | epithelial membrane protein 2 | 773 | 0.174 | 0.4496 | Yes |
| 68 | SPRY2 | SPRY2 Entrez, Source | sprouty homolog 2 (Drosophila) | 784 | 0.172 | 0.4524 | Yes |
| 69 | FLT1 | FLT1 Entrez, Source | fms-related tyrosine kinase 1 (vascular endothelial growth factor/vascular permeability factor receptor) | 789 | 0.172 | 0.4556 | Yes |
| 70 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 837 | 0.164 | 0.4554 | Yes |
| 71 | JUNB | JUNB Entrez, Source | jun B proto-oncogene | 861 | 0.161 | 0.4569 | Yes |
| 72 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 880 | 0.159 | 0.4588 | Yes |
| 73 | FABP4 | FABP4 Entrez, Source | fatty acid binding protein 4, adipocyte | 884 | 0.158 | 0.4618 | Yes |
| 74 | PALMD | PALMD Entrez, Source | palmdelphin | 986 | 0.148 | 0.4571 | Yes |
| 75 | KLF9 | KLF9 Entrez, Source | Kruppel-like factor 9 | 998 | 0.147 | 0.4593 | Yes |
| 76 | IL3RA | IL3RA Entrez, Source | interleukin 3 receptor, alpha (low affinity) | 1013 | 0.146 | 0.4612 | Yes |
| 77 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 1041 | 0.142 | 0.4621 | Yes |
| 78 | SLC38A1 | SLC38A1 Entrez, Source | solute carrier family 38, member 1 | 1109 | 0.135 | 0.4597 | Yes |
| 79 | AGC1 | AGC1 Entrez, Source | aggrecan 1 (chondroitin sulfate proteoglycan 1, large aggregating proteoglycan, antigen identified by monoclonal antibody A0122) | 1118 | 0.135 | 0.4619 | Yes |
| 80 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 1170 | 0.131 | 0.4606 | Yes |
| 81 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 1210 | 0.127 | 0.4602 | Yes |
| 82 | KLHL24 | KLHL24 Entrez, Source | kelch-like 24 (Drosophila) | 1215 | 0.126 | 0.4625 | Yes |
| 83 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 1221 | 0.126 | 0.4648 | Yes |
| 84 | ICA1 | ICA1 Entrez, Source | islet cell autoantigen 1, 69kDa | 1223 | 0.126 | 0.4673 | Yes |
| 85 | CD302 | CD302 Entrez, Source | CD302 molecule | 1233 | 0.125 | 0.4691 | Yes |
| 86 | PSCD1 | PSCD1 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 1(cytohesin 1) | 1237 | 0.125 | 0.4715 | Yes |
| 87 | TFPI2 | TFPI2 Entrez, Source | tissue factor pathway inhibitor 2 | 1241 | 0.124 | 0.4738 | Yes |
| 88 | ZFP36 | ZFP36 Entrez, Source | zinc finger protein 36, C3H type, homolog (mouse) | 1260 | 0.123 | 0.4750 | Yes |
| 89 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 1261 | 0.123 | 0.4775 | Yes |
| 90 | CLK1 | CLK1 Entrez, Source | CDC-like kinase 1 | 1262 | 0.123 | 0.4800 | Yes |
| 91 | KLF6 | KLF6 Entrez, Source | Kruppel-like factor 6 | 1286 | 0.121 | 0.4808 | Yes |
| 92 | TSPAN8 | TSPAN8 Entrez, Source | tetraspanin 8 | 1311 | 0.119 | 0.4814 | Yes |
| 93 | SERPINA5 | SERPINA5 Entrez, Source | serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 5 | 1314 | 0.119 | 0.4837 | Yes |
| 94 | TSPAN12 | TSPAN12 Entrez, Source | tetraspanin 12 | 1431 | 0.112 | 0.4770 | No |
| 95 | PER2 | PER2 Entrez, Source | period homolog 2 (Drosophila) | 1454 | 0.110 | 0.4776 | No |
| 96 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 1504 | 0.108 | 0.4761 | No |
| 97 | RNASE4 | RNASE4 Entrez, Source | ribonuclease, RNase A family, 4 | 1525 | 0.107 | 0.4767 | No |
| 98 | PELI1 | PELI1 Entrez, Source | pellino homolog 1 (Drosophila) | 1568 | 0.104 | 0.4756 | No |
| 99 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 1582 | 0.103 | 0.4768 | No |
| 100 | TGM2 | TGM2 Entrez, Source | transglutaminase 2 (C polypeptide, protein-glutamine-gamma-glutamyltransferase) | 1650 | 0.100 | 0.4737 | No |
| 101 | GPC3 | GPC3 Entrez, Source | glypican 3 | 1656 | 0.100 | 0.4753 | No |
| 102 | PTPRE | PTPRE Entrez, Source | protein tyrosine phosphatase, receptor type, E | 1672 | 0.099 | 0.4762 | No |
| 103 | UPP1 | UPP1 Entrez, Source | uridine phosphorylase 1 | 1679 | 0.098 | 0.4778 | No |
| 104 | SLC16A3 | SLC16A3 Entrez, Source | solute carrier family 16, member 3 (monocarboxylic acid transporter 4) | 1690 | 0.098 | 0.4790 | No |
| 105 | DCX | DCX Entrez, Source | doublecortex; lissencephaly, X-linked (doublecortin) | 1708 | 0.097 | 0.4797 | No |
| 106 | AKAP13 | AKAP13 Entrez, Source | A kinase (PRKA) anchor protein 13 | 1725 | 0.096 | 0.4805 | No |
| 107 | COL11A1 | COL11A1 Entrez, Source | collagen, type XI, alpha 1 | 1749 | 0.095 | 0.4806 | No |
| 108 | ARHGDIB | ARHGDIB Entrez, Source | Rho GDP dissociation inhibitor (GDI) beta | 1774 | 0.094 | 0.4807 | No |
| 109 | FXYD1 | FXYD1 Entrez, Source | FXYD domain containing ion transport regulator 1 (phospholemman) | 1781 | 0.094 | 0.4822 | No |
| 110 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 1849 | 0.091 | 0.4789 | No |
| 111 | CXCL14 | CXCL14 Entrez, Source | chemokine (C-X-C motif) ligand 14 | 1856 | 0.090 | 0.4803 | No |
| 112 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 1942 | 0.087 | 0.4756 | No |
| 113 | SULF1 | SULF1 Entrez, Source | sulfatase 1 | 1955 | 0.087 | 0.4764 | No |
| 114 | LCP2 | LCP2 Entrez, Source | lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa) | 1958 | 0.087 | 0.4781 | No |
| 115 | DSCR1 | DSCR1 Entrez, Source | Down syndrome critical region gene 1 | 1990 | 0.085 | 0.4774 | No |
| 116 | ST6GAL1 | ST6GAL1 Entrez, Source | ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 | 2013 | 0.085 | 0.4775 | No |
| 117 | PTGER2 | PTGER2 Entrez, Source | prostaglandin E receptor 2 (subtype EP2), 53kDa | 2027 | 0.084 | 0.4782 | No |
| 118 | GLIPR1 | GLIPR1 Entrez, Source | GLI pathogenesis-related 1 (glioma) | 2028 | 0.084 | 0.4799 | No |
| 119 | MYRIP | MYRIP Entrez, Source | myosin VIIA and Rab interacting protein | 2080 | 0.082 | 0.4777 | No |
| 120 | APOE | APOE Entrez, Source | apolipoprotein E | 2123 | 0.080 | 0.4761 | No |
| 121 | CHRDL1 | CHRDL1 Entrez, Source | chordin-like 1 | 2170 | 0.078 | 0.4742 | No |
| 122 | TPM4 | TPM4 Entrez, Source | tropomyosin 4 | 2205 | 0.077 | 0.4731 | No |
| 123 | SDPR | SDPR Entrez, Source | serum deprivation response (phosphatidylserine binding protein) | 2231 | 0.076 | 0.4728 | No |
| 124 | WISP2 | WISP2 Entrez, Source | WNT1 inducible signaling pathway protein 2 | 2285 | 0.074 | 0.4702 | No |
| 125 | NELL2 | NELL2 Entrez, Source | NEL-like 2 (chicken) | 2324 | 0.073 | 0.4688 | No |
| 126 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 2451 | 0.069 | 0.4605 | No |
| 127 | TNMD | TNMD Entrez, Source | tenomodulin | 2474 | 0.069 | 0.4603 | No |
| 128 | CSF3 | CSF3 Entrez, Source | colony stimulating factor 3 (granulocyte) | 2483 | 0.069 | 0.4610 | No |
| 129 | CLIC3 | CLIC3 Entrez, Source | chloride intracellular channel 3 | 2562 | 0.066 | 0.4564 | No |
| 130 | SLC19A2 | SLC19A2 Entrez, Source | solute carrier family 19 (thiamine transporter), member 2 | 2667 | 0.063 | 0.4497 | No |
| 131 | FN1 | FN1 Entrez, Source | fibronectin 1 | 2715 | 0.062 | 0.4473 | No |
| 132 | CTSH | CTSH Entrez, Source | cathepsin H | 2723 | 0.062 | 0.4481 | No |
| 133 | LMO7 | LMO7 Entrez, Source | LIM domain 7 | 2745 | 0.062 | 0.4477 | No |
| 134 | TIE1 | TIE1 Entrez, Source | tyrosine kinase with immunoglobulin-like and EGF-like domains 1 | 2769 | 0.061 | 0.4472 | No |
| 135 | SELP | SELP Entrez, Source | selectin P (granule membrane protein 140kDa, antigen CD62) | 2789 | 0.060 | 0.4470 | No |
| 136 | TRIB1 | TRIB1 Entrez, Source | tribbles homolog 1 (Drosophila) | 2792 | 0.060 | 0.4481 | No |
| 137 | ING1 | ING1 Entrez, Source | inhibitor of growth family, member 1 | 2807 | 0.060 | 0.4482 | No |
| 138 | BAALC | BAALC Entrez, Source | brain and acute leukemia, cytoplasmic | 2819 | 0.060 | 0.4486 | No |
| 139 | TPST2 | TPST2 Entrez, Source | tyrosylprotein sulfotransferase 2 | 2856 | 0.059 | 0.4471 | No |
| 140 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 2902 | 0.057 | 0.4448 | No |
| 141 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 2935 | 0.057 | 0.4435 | No |
| 142 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 3028 | 0.054 | 0.4375 | No |
| 143 | TGFBR3 | TGFBR3 Entrez, Source | transforming growth factor, beta receptor III (betaglycan, 300kDa) | 3034 | 0.054 | 0.4382 | No |
| 144 | CTSS | CTSS Entrez, Source | cathepsin S | 3084 | 0.052 | 0.4355 | No |
| 145 | JUN | JUN Entrez, Source | jun oncogene | 3104 | 0.052 | 0.4351 | No |
| 146 | MFAP4 | MFAP4 Entrez, Source | microfibrillar-associated protein 4 | 3105 | 0.052 | 0.4362 | No |
| 147 | MAFB | MAFB Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog B (avian) | 3122 | 0.051 | 0.4360 | No |
| 148 | AP2S1 | AP2S1 Entrez, Source | adaptor-related protein complex 2, sigma 1 subunit | 3175 | 0.050 | 0.4330 | No |
| 149 | CYP26B1 | CYP26B1 Entrez, Source | cytochrome P450, family 26, subfamily B, polypeptide 1 | 3231 | 0.049 | 0.4298 | No |
| 150 | APOD | APOD Entrez, Source | apolipoprotein D | 3255 | 0.048 | 0.4290 | No |
| 151 | CA4 | CA4 Entrez, Source | carbonic anhydrase IV | 3258 | 0.048 | 0.4298 | No |
| 152 | JAG2 | JAG2 Entrez, Source | jagged 2 | 3370 | 0.046 | 0.4222 | No |
| 153 | DCN | DCN Entrez, Source | decorin | 3405 | 0.045 | 0.4205 | No |
| 154 | FRMD4B | FRMD4B Entrez, Source | FERM domain containing 4B | 3422 | 0.045 | 0.4202 | No |
| 155 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 3428 | 0.044 | 0.4208 | No |
| 156 | GREM1 | GREM1 Entrez, Source | gremlin 1, cysteine knot superfamily, homolog (Xenopus laevis) | 3429 | 0.044 | 0.4217 | No |
| 157 | MCF2L | MCF2L Entrez, Source | MCF.2 cell line derived transforming sequence-like | 3454 | 0.044 | 0.4207 | No |
| 158 | WNT11 | WNT11 Entrez, Source | wingless-type MMTV integration site family, member 11 | 3495 | 0.043 | 0.4185 | No |
| 159 | SEMA3G | SEMA3G Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3G | 3558 | 0.042 | 0.4146 | No |
| 160 | KDR | KDR Entrez, Source | kinase insert domain receptor (a type III receptor tyrosine kinase) | 3567 | 0.041 | 0.4148 | No |
| 161 | MOXD1 | MOXD1 Entrez, Source | monooxygenase, DBH-like 1 | 3603 | 0.041 | 0.4130 | No |
| 162 | SEPP1 | SEPP1 Entrez, Source | selenoprotein P, plasma, 1 | 3625 | 0.040 | 0.4122 | No |
| 163 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 3628 | 0.040 | 0.4129 | No |
| 164 | PER3 | PER3 Entrez, Source | period homolog 3 (Drosophila) | 3660 | 0.040 | 0.4113 | No |
| 165 | ASPA | ASPA Entrez, Source | aspartoacylase (Canavan disease) | 3681 | 0.039 | 0.4106 | No |
| 166 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 3732 | 0.038 | 0.4075 | No |
| 167 | SPARCL1 | SPARCL1 Entrez, Source | SPARC-like 1 (mast9, hevin) | 3771 | 0.037 | 0.4053 | No |
| 168 | F11R | F11R Entrez, Source | F11 receptor | 3837 | 0.036 | 0.4011 | No |
| 169 | JUND | JUND Entrez, Source | jun D proto-oncogene | 3846 | 0.036 | 0.4012 | No |
| 170 | SLC1A1 | SLC1A1 Entrez, Source | solute carrier family 1 (neuronal/epithelial high affinity glutamate transporter, system Xag), member 1 | 3880 | 0.035 | 0.3994 | No |
| 171 | CXCR7 | CXCR7 Entrez, Source | chemokine (C-X-C motif) receptor 7 | 3953 | 0.034 | 0.3945 | No |
| 172 | GAS7 | GAS7 Entrez, Source | growth arrest-specific 7 | 3956 | 0.034 | 0.3950 | No |
| 173 | NRN1 | NRN1 Entrez, Source | neuritin 1 | 3957 | 0.034 | 0.3957 | No |
| 174 | NOTCH4 | NOTCH4 Entrez, Source | Notch homolog 4 (Drosophila) | 3964 | 0.033 | 0.3960 | No |
| 175 | KL | KL Entrez, Source | klotho | 4012 | 0.033 | 0.3930 | No |
| 176 | ELOVL5 | ELOVL5 Entrez, Source | ELOVL family member 5, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4037 | 0.032 | 0.3918 | No |
| 177 | MARK1 | MARK1 Entrez, Source | MAP/microtubule affinity-regulating kinase 1 | 4077 | 0.031 | 0.3895 | No |
| 178 | DSP | DSP Entrez, Source | desmoplakin | 4079 | 0.031 | 0.3900 | No |
| 179 | RGC32 | RGC32 Entrez, Source | - | 4104 | 0.031 | 0.3888 | No |
| 180 | NQO1 | NQO1 Entrez, Source | NAD(P)H dehydrogenase, quinone 1 | 4179 | 0.030 | 0.3837 | No |
| 181 | SCRG1 | SCRG1 Entrez, Source | - | 4215 | 0.029 | 0.3816 | No |
| 182 | FLNC | FLNC Entrez, Source | filamin C, gamma (actin binding protein 280) | 4230 | 0.029 | 0.3811 | No |
| 183 | RND1 | RND1 Entrez, Source | Rho family GTPase 1 | 4318 | 0.027 | 0.3750 | No |
| 184 | AGTRL1 | AGTRL1 Entrez, Source | angiotensin II receptor-like 1 | 4338 | 0.027 | 0.3741 | No |
| 185 | CPE | CPE Entrez, Source | carboxypeptidase E | 4444 | 0.025 | 0.3665 | No |
| 186 | RPS6KA2 | RPS6KA2 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 2 | 4460 | 0.024 | 0.3658 | No |
| 187 | SH2B3 | SH2B3 Entrez, Source | SH2B adaptor protein 3 | 4523 | 0.023 | 0.3615 | No |
| 188 | GDF10 | GDF10 Entrez, Source | growth differentiation factor 10 | 4564 | 0.022 | 0.3589 | No |
| 189 | HLF | HLF Entrez, Source | hepatic leukemia factor | 4632 | 0.021 | 0.3542 | No |
| 190 | SERPINF1 | SERPINF1 Entrez, Source | serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 | 4697 | 0.020 | 0.3496 | No |
| 191 | ADRB2 | ADRB2 Entrez, Source | adrenergic, beta-2-, receptor, surface | 4733 | 0.020 | 0.3473 | No |
| 192 | GGTLA1 | GGTLA1 Entrez, Source | gamma-glutamyltransferase-like activity 1 | 4761 | 0.019 | 0.3457 | No |
| 193 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 4794 | 0.019 | 0.3436 | No |
| 194 | SYNPO | SYNPO Entrez, Source | synaptopodin | 4898 | 0.017 | 0.3360 | No |
| 195 | KIAA0101 | KIAA0101 Entrez, Source | KIAA0101 | 4903 | 0.017 | 0.3360 | No |
| 196 | ATF5 | ATF5 Entrez, Source | activating transcription factor 5 | 4937 | 0.016 | 0.3338 | No |
| 197 | PRKCH | PRKCH Entrez, Source | protein kinase C, eta | 4972 | 0.016 | 0.3315 | No |
| 198 | ZBTB16 | ZBTB16 Entrez, Source | zinc finger and BTB domain containing 16 | 5042 | 0.014 | 0.3265 | No |
| 199 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 5081 | 0.014 | 0.3238 | No |
| 200 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 5086 | 0.014 | 0.3238 | No |
| 201 | GUCY1A3 | GUCY1A3 Entrez, Source | guanylate cyclase 1, soluble, alpha 3 | 5232 | 0.011 | 0.3129 | No |
| 202 | PDE2A | PDE2A Entrez, Source | phosphodiesterase 2A, cGMP-stimulated | 5264 | 0.011 | 0.3107 | No |
| 203 | AQP1 | AQP1 Entrez, Source | aquaporin 1 (Colton blood group) | 5400 | 0.009 | 0.3005 | No |
| 204 | ICAM2 | ICAM2 Entrez, Source | intercellular adhesion molecule 2 | 5448 | 0.008 | 0.2970 | No |
| 205 | CDK7 | CDK7 Entrez, Source | cyclin-dependent kinase 7 (MO15 homolog, Xenopus laevis, cdk-activating kinase) | 5591 | 0.006 | 0.2862 | No |
| 206 | EPHA3 | EPHA3 Entrez, Source | EPH receptor A3 | 5621 | 0.005 | 0.2840 | No |
| 207 | IL15 | IL15 Entrez, Source | interleukin 15 | 5674 | 0.005 | 0.2801 | No |
| 208 | SPATA6 | SPATA6 Entrez, Source | spermatogenesis associated 6 | 5677 | 0.005 | 0.2801 | No |
| 209 | WSB2 | WSB2 Entrez, Source | WD repeat and SOCS box-containing 2 | 5679 | 0.005 | 0.2801 | No |
| 210 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 5704 | 0.004 | 0.2783 | No |
| 211 | SMAD1 | SMAD1 Entrez, Source | SMAD, mothers against DPP homolog 1 (Drosophila) | 5731 | 0.004 | 0.2764 | No |
| 212 | LIMK2 | LIMK2 Entrez, Source | LIM domain kinase 2 | 5743 | 0.004 | 0.2756 | No |
| 213 | MAOA | MAOA Entrez, Source | monoamine oxidase A | 5751 | 0.003 | 0.2752 | No |
| 214 | ENG | ENG Entrez, Source | endoglin (Osler-Rendu-Weber syndrome 1) | 5761 | 0.003 | 0.2745 | No |
| 215 | GIMAP6 | GIMAP6 Entrez, Source | GTPase, IMAP family member 6 | 5839 | 0.002 | 0.2686 | No |
| 216 | GLS | GLS Entrez, Source | glutaminase | 5935 | 0.001 | 0.2613 | No |
| 217 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 5955 | 0.001 | 0.2599 | No |
| 218 | SLC1A4 | SLC1A4 Entrez, Source | solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 | 6096 | -0.002 | 0.2491 | No |
| 219 | CDO1 | CDO1 Entrez, Source | cysteine dioxygenase, type I | 6103 | -0.002 | 0.2487 | No |
| 220 | ACSL5 | ACSL5 Entrez, Source | acyl-CoA synthetase long-chain family member 5 | 6137 | -0.002 | 0.2462 | No |
| 221 | JAM2 | JAM2 Entrez, Source | junctional adhesion molecule 2 | 6141 | -0.002 | 0.2460 | No |
| 222 | NOX4 | NOX4 Entrez, Source | NADPH oxidase 4 | 6149 | -0.002 | 0.2455 | No |
| 223 | OXTR | OXTR Entrez, Source | oxytocin receptor | 6165 | -0.002 | 0.2444 | No |
| 224 | FMO2 | FMO2 Entrez, Source | flavin containing monooxygenase 2 (non-functional) | 6188 | -0.003 | 0.2428 | No |
| 225 | CFI | CFI Entrez, Source | complement factor I | 6257 | -0.004 | 0.2376 | No |
| 226 | ESR1 | ESR1 Entrez, Source | estrogen receptor 1 | 6300 | -0.005 | 0.2345 | No |
| 227 | RRAD | RRAD Entrez, Source | Ras-related associated with diabetes | 6316 | -0.005 | 0.2334 | No |
| 228 | ADAM10 | ADAM10 Entrez, Source | ADAM metallopeptidase domain 10 | 6375 | -0.006 | 0.2291 | No |
| 229 | ACACB | ACACB Entrez, Source | acetyl-Coenzyme A carboxylase beta | 6400 | -0.006 | 0.2274 | No |
| 230 | LAPTM5 | LAPTM5 Entrez, Source | lysosomal associated multispanning membrane protein 5 | 6572 | -0.009 | 0.2144 | No |
| 231 | NINJ2 | NINJ2 Entrez, Source | ninjurin 2 | 6616 | -0.009 | 0.2112 | No |
| 232 | PLVAP | PLVAP Entrez, Source | plasmalemma vesicle associated protein | 6633 | -0.010 | 0.2102 | No |
| 233 | PTPRM | PTPRM Entrez, Source | protein tyrosine phosphatase, receptor type, M | 6655 | -0.010 | 0.2088 | No |
| 234 | C3ORF64 | C3ORF64 Entrez, Source | chromosome 3 open reading frame 64 | 6713 | -0.011 | 0.2046 | No |
| 235 | ANGPTL2 | ANGPTL2 Entrez, Source | angiopoietin-like 2 | 6736 | -0.011 | 0.2031 | No |
| 236 | EDNRB | EDNRB Entrez, Source | endothelin receptor type B | 6802 | -0.012 | 0.1984 | No |
| 237 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 6833 | -0.013 | 0.1963 | No |
| 238 | ACTC1 | ACTC1 Entrez, Source | actin, alpha, cardiac muscle 1 | 6882 | -0.013 | 0.1929 | No |
| 239 | STC1 | STC1 Entrez, Source | stanniocalcin 1 | 6986 | -0.015 | 0.1853 | No |
| 240 | HSPB7 | HSPB7 Entrez, Source | heat shock 27kDa protein family, member 7 (cardiovascular) | 7043 | -0.016 | 0.1813 | No |
| 241 | LTBP4 | LTBP4 Entrez, Source | latent transforming growth factor beta binding protein 4 | 7185 | -0.018 | 0.1708 | No |
| 242 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 7189 | -0.018 | 0.1709 | No |
| 243 | CFH | CFH Entrez, Source | complement factor H | 7191 | -0.018 | 0.1712 | No |
| 244 | DNM3 | DNM3 Entrez, Source | dynamin 3 | 7241 | -0.019 | 0.1678 | No |
| 245 | CNOT2 | CNOT2 Entrez, Source | CCR4-NOT transcription complex, subunit 2 | 7253 | -0.019 | 0.1674 | No |
| 246 | CDH2 | CDH2 Entrez, Source | cadherin 2, type 1, N-cadherin (neuronal) | 7290 | -0.019 | 0.1650 | No |
| 247 | HAPLN1 | HAPLN1 Entrez, Source | hyaluronan and proteoglycan link protein 1 | 7300 | -0.020 | 0.1647 | No |
| 248 | GATA2 | GATA2 Entrez, Source | GATA binding protein 2 | 7313 | -0.020 | 0.1642 | No |
| 249 | DNM1 | DNM1 Entrez, Source | dynamin 1 | 7344 | -0.020 | 0.1623 | No |
| 250 | COL15A1 | COL15A1 Entrez, Source | collagen, type XV, alpha 1 | 7379 | -0.021 | 0.1601 | No |
| 251 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 7396 | -0.021 | 0.1593 | No |
| 252 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 7402 | -0.021 | 0.1593 | No |
| 253 | EMCN | EMCN Entrez, Source | endomucin | 7455 | -0.022 | 0.1558 | No |
| 254 | ANGPT2 | ANGPT2 Entrez, Source | angiopoietin 2 | 7458 | -0.022 | 0.1561 | No |
| 255 | MTUS1 | MTUS1 Entrez, Source | mitochondrial tumor suppressor 1 | 7482 | -0.022 | 0.1548 | No |
| 256 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 7557 | -0.023 | 0.1495 | No |
| 257 | PDGFRL | PDGFRL Entrez, Source | platelet-derived growth factor receptor-like | 7568 | -0.024 | 0.1493 | No |
| 258 | DMN | DMN Entrez, Source | desmuslin | 7569 | -0.024 | 0.1497 | No |
| 259 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 7649 | -0.025 | 0.1442 | No |
| 260 | NOVA1 | NOVA1 Entrez, Source | neuro-oncological ventral antigen 1 | 7658 | -0.025 | 0.1441 | No |
| 261 | ACLY | ACLY Entrez, Source | ATP citrate lyase | 7774 | -0.027 | 0.1358 | No |
| 262 | SOCS3 | SOCS3 Entrez, Source | suppressor of cytokine signaling 3 | 7799 | -0.027 | 0.1345 | No |
| 263 | C6 | C6 Entrez, Source | complement component 6 | 7841 | -0.028 | 0.1319 | No |
| 264 | F10 | F10 Entrez, Source | coagulation factor X | 7895 | -0.029 | 0.1284 | No |
| 265 | NEDD9 | NEDD9 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 9 | 7899 | -0.029 | 0.1288 | No |
| 266 | MCFD2 | MCFD2 Entrez, Source | multiple coagulation factor deficiency 2 | 7936 | -0.030 | 0.1266 | No |
| 267 | PFKFB3 | PFKFB3 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 3 | 7937 | -0.030 | 0.1272 | No |
| 268 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 7950 | -0.030 | 0.1269 | No |
| 269 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 7989 | -0.031 | 0.1246 | No |
| 270 | NPY1R | NPY1R Entrez, Source | neuropeptide Y receptor Y1 | 8092 | -0.032 | 0.1174 | No |
| 271 | CTSK | CTSK Entrez, Source | cathepsin K (pycnodysostosis) | 8117 | -0.032 | 0.1162 | No |
| 272 | COMP | COMP Entrez, Source | cartilage oligomeric matrix protein | 8125 | -0.033 | 0.1164 | No |
| 273 | KCNK2 | KCNK2 Entrez, Source | potassium channel, subfamily K, member 2 | 8139 | -0.033 | 0.1161 | No |
| 274 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 8146 | -0.033 | 0.1163 | No |
| 275 | PIK3R3 | PIK3R3 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 3 (p55, gamma) | 8161 | -0.033 | 0.1159 | No |
| 276 | VWF | VWF Entrez, Source | von Willebrand factor | 8376 | -0.037 | 0.1001 | No |
| 277 | CDC37 | CDC37 Entrez, Source | CDC37 cell division cycle 37 homolog (S. cerevisiae) | 8391 | -0.037 | 0.0998 | No |
| 278 | ELTD1 | ELTD1 Entrez, Source | EGF, latrophilin and seven transmembrane domain containing 1 | 8400 | -0.037 | 0.1000 | No |
| 279 | KCNAB1 | KCNAB1 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, beta member 1 | 8440 | -0.038 | 0.0978 | No |
| 280 | FGF2 | FGF2 Entrez, Source | fibroblast growth factor 2 (basic) | 8465 | -0.038 | 0.0967 | No |
| 281 | CAPN6 | CAPN6 Entrez, Source | calpain 6 | 8491 | -0.039 | 0.0956 | No |
| 282 | SCG5 | SCG5 Entrez, Source | secretogranin V (7B2 protein) | 8494 | -0.039 | 0.0962 | No |
| 283 | IER5 | IER5 Entrez, Source | immediate early response 5 | 8527 | -0.039 | 0.0946 | No |
| 284 | ASNS | ASNS Entrez, Source | asparagine synthetase | 8596 | -0.040 | 0.0902 | No |
| 285 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 8667 | -0.041 | 0.0856 | No |
| 286 | NR5A2 | NR5A2 Entrez, Source | nuclear receptor subfamily 5, group A, member 2 | 8704 | -0.042 | 0.0837 | No |
| 287 | CD14 | CD14 Entrez, Source | CD14 molecule | 8706 | -0.042 | 0.0845 | No |
| 288 | LMO2 | LMO2 Entrez, Source | LIM domain only 2 (rhombotin-like 1) | 8708 | -0.042 | 0.0853 | No |
| 289 | SLCO4A1 | SLCO4A1 Entrez, Source | solute carrier organic anion transporter family, member 4A1 | 8782 | -0.043 | 0.0806 | No |
| 290 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 8829 | -0.044 | 0.0779 | No |
| 291 | TRPC4 | TRPC4 Entrez, Source | transient receptor potential cation channel, subfamily C, member 4 | 9085 | -0.049 | 0.0593 | No |
| 292 | GIMAP4 | GIMAP4 Entrez, Source | GTPase, IMAP family member 4 | 9106 | -0.049 | 0.0587 | No |
| 293 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 9173 | -0.050 | 0.0547 | No |
| 294 | TMPO | TMPO Entrez, Source | thymopoietin | 9189 | -0.050 | 0.0546 | No |
| 295 | GPNMB | GPNMB Entrez, Source | glycoprotein (transmembrane) nmb | 9192 | -0.050 | 0.0555 | No |
| 296 | HHEX | HHEX Entrez, Source | homeobox, hematopoietically expressed | 9234 | -0.051 | 0.0534 | No |
| 297 | CD93 | CD93 Entrez, Source | CD93 molecule | 9341 | -0.053 | 0.0463 | No |
| 298 | SRD5A1 | SRD5A1 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 1 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 1) | 9458 | -0.055 | 0.0385 | No |
| 299 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 9459 | -0.055 | 0.0396 | No |
| 300 | CLDN5 | CLDN5 Entrez, Source | claudin 5 (transmembrane protein deleted in velocardiofacial syndrome) | 9647 | -0.059 | 0.0264 | No |
| 301 | EIF5 | EIF5 Entrez, Source | eukaryotic translation initiation factor 5 | 9653 | -0.059 | 0.0272 | No |
| 302 | FRZB | FRZB Entrez, Source | frizzled-related protein | 9829 | -0.063 | 0.0150 | No |
| 303 | PGK1 | PGK1 Entrez, Source | phosphoglycerate kinase 1 | 9880 | -0.064 | 0.0125 | No |
| 304 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 10004 | -0.066 | 0.0044 | No |
| 305 | KCNS3 | KCNS3 Entrez, Source | potassium voltage-gated channel, delayed-rectifier, subfamily S, member 3 | 10019 | -0.067 | 0.0047 | No |
| 306 | MYOC | MYOC Entrez, Source | myocilin, trabecular meshwork inducible glucocorticoid response | 10189 | -0.070 | -0.0069 | No |
| 307 | COLEC12 | COLEC12 Entrez, Source | collectin sub-family member 12 | 10217 | -0.071 | -0.0075 | No |
| 308 | SRPX | SRPX Entrez, Source | sushi-repeat-containing protein, X-linked | 10360 | -0.074 | -0.0169 | No |
| 309 | AOC3 | AOC3 Entrez, Source | amine oxidase, copper containing 3 (vascular adhesion protein 1) | 10438 | -0.076 | -0.0213 | No |
| 310 | TNFRSF11B | TNFRSF11B Entrez, Source | tumor necrosis factor receptor superfamily, member 11b (osteoprotegerin) | 10443 | -0.076 | -0.0200 | No |
| 311 | SELE | SELE Entrez, Source | selectin E (endothelial adhesion molecule 1) | 10446 | -0.076 | -0.0186 | No |
| 312 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 10547 | -0.079 | -0.0247 | No |
| 313 | BAX | BAX Entrez, Source | BCL2-associated X protein | 10615 | -0.080 | -0.0282 | No |
| 314 | EIF2S2 | EIF2S2 Entrez, Source | eukaryotic translation initiation factor 2, subunit 2 beta, 38kDa | 10731 | -0.084 | -0.0354 | No |
| 315 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 10960 | -0.089 | -0.0511 | No |
| 316 | UCHL1 | UCHL1 Entrez, Source | ubiquitin carboxyl-terminal esterase L1 (ubiquitin thiolesterase) | 11042 | -0.092 | -0.0555 | No |
| 317 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 11066 | -0.092 | -0.0553 | No |
| 318 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 11118 | -0.094 | -0.0573 | No |
| 319 | GMFG | GMFG Entrez, Source | glia maturation factor, gamma | 11174 | -0.095 | -0.0596 | No |
| 320 | SLC31A1 | SLC31A1 Entrez, Source | solute carrier family 31 (copper transporters), member 1 | 11210 | -0.096 | -0.0603 | No |
| 321 | WNT5A | WNT5A Entrez, Source | wingless-type MMTV integration site family, member 5A | 11232 | -0.097 | -0.0600 | No |
| 322 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 11272 | -0.098 | -0.0609 | No |
| 323 | NTRK2 | NTRK2 Entrez, Source | neurotrophic tyrosine kinase, receptor, type 2 | 11369 | -0.101 | -0.0663 | No |
| 324 | WARS | WARS Entrez, Source | tryptophanyl-tRNA synthetase | 11445 | -0.104 | -0.0699 | No |
| 325 | RAI14 | RAI14 Entrez, Source | retinoic acid induced 14 | 11447 | -0.104 | -0.0678 | No |
| 326 | GFPT1 | GFPT1 Entrez, Source | glutamine-fructose-6-phosphate transaminase 1 | 11452 | -0.104 | -0.0660 | No |
| 327 | IRF1 | IRF1 Entrez, Source | interferon regulatory factor 1 | 11466 | -0.105 | -0.0648 | No |
| 328 | SHMT2 | SHMT2 Entrez, Source | serine hydroxymethyltransferase 2 (mitochondrial) | 11474 | -0.105 | -0.0632 | No |
| 329 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 11513 | -0.106 | -0.0639 | No |
| 330 | PRKD1 | PRKD1 Entrez, Source | protein kinase D1 | 11594 | -0.109 | -0.0679 | No |
| 331 | TRIB3 | TRIB3 Entrez, Source | tribbles homolog 3 (Drosophila) | 11605 | -0.110 | -0.0664 | No |
| 332 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 11607 | -0.110 | -0.0642 | No |
| 333 | EIF1 | EIF1 Entrez, Source | eukaryotic translation initiation factor 1 | 11662 | -0.112 | -0.0660 | No |
| 334 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 11710 | -0.114 | -0.0673 | No |
| 335 | TOMM40 | TOMM40 Entrez, Source | translocase of outer mitochondrial membrane 40 homolog (yeast) | 11720 | -0.114 | -0.0657 | No |
| 336 | PDLIM5 | PDLIM5 Entrez, Source | PDZ and LIM domain 5 | 11910 | -0.122 | -0.0777 | No |
| 337 | GPR56 | GPR56 Entrez, Source | G protein-coupled receptor 56 | 12030 | -0.128 | -0.0843 | No |
| 338 | A2M | A2M Entrez, Source | alpha-2-macroglobulin | 12069 | -0.130 | -0.0845 | No |
| 339 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 12092 | -0.131 | -0.0835 | No |
| 340 | EFNA1 | EFNA1 Entrez, Source | ephrin-A1 | 12095 | -0.131 | -0.0810 | No |
| 341 | NDRG2 | NDRG2 Entrez, Source | NDRG family member 2 | 12174 | -0.135 | -0.0842 | No |
| 342 | TSPAN6 | TSPAN6 Entrez, Source | tetraspanin 6 | 12255 | -0.140 | -0.0875 | No |
| 343 | PODXL | PODXL Entrez, Source | podocalyxin-like | 12264 | -0.140 | -0.0852 | No |
| 344 | PSAT1 | PSAT1 Entrez, Source | phosphoserine aminotransferase 1 | 12281 | -0.141 | -0.0835 | No |
| 345 | BCHE | BCHE Entrez, Source | butyrylcholinesterase | 12286 | -0.141 | -0.0809 | No |
| 346 | ARTS-1 | ARTS-1 Entrez, Source | - | 12308 | -0.143 | -0.0796 | No |
| 347 | IFI44 | IFI44 Entrez, Source | interferon-induced protein 44 | 12405 | -0.149 | -0.0839 | No |
| 348 | USP3 | USP3 Entrez, Source | ubiquitin specific peptidase 3 | 12588 | -0.162 | -0.0946 | No |
| 349 | GIMAP5 | GIMAP5 Entrez, Source | GTPase, IMAP family member 5 | 12589 | -0.162 | -0.0912 | No |
| 350 | SLC7A1 | SLC7A1 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 | 12592 | -0.163 | -0.0880 | No |
| 351 | MET | MET Entrez, Source | met proto-oncogene (hepatocyte growth factor receptor) | 12611 | -0.164 | -0.0860 | No |
| 352 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 12775 | -0.181 | -0.0949 | No |
| 353 | DOCK9 | DOCK9 Entrez, Source | dedicator of cytokinesis 9 | 12801 | -0.185 | -0.0930 | No |
| 354 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 12802 | -0.185 | -0.0891 | No |
| 355 | CYP4B1 | CYP4B1 Entrez, Source | cytochrome P450, family 4, subfamily B, polypeptide 1 | 12818 | -0.188 | -0.0864 | No |
| 356 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 12857 | -0.193 | -0.0854 | No |
| 357 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 12876 | -0.196 | -0.0827 | No |
| 358 | GBP2 | GBP2 Entrez, Source | guanylate binding protein 2, interferon-inducible | 12940 | -0.207 | -0.0833 | No |
| 359 | GNG11 | GNG11 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 11 | 13005 | -0.219 | -0.0837 | No |
| 360 | ZBTB10 | ZBTB10 Entrez, Source | zinc finger and BTB domain containing 10 | 13012 | -0.220 | -0.0796 | No |
| 361 | HNRPM | HNRPM Entrez, Source | heterogeneous nuclear ribonucleoprotein M | 13088 | -0.247 | -0.0803 | No |
| 362 | FMO1 | FMO1 Entrez, Source | flavin containing monooxygenase 1 | 13123 | -0.259 | -0.0776 | No |
| 363 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 13137 | -0.265 | -0.0732 | No |
| 364 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 13201 | -0.300 | -0.0718 | No |
| 365 | BST2 | BST2 Entrez, Source | bone marrow stromal cell antigen 2 | 13220 | -0.317 | -0.0667 | No |
| 366 | CES1 | CES1 Entrez, Source | carboxylesterase 1 (monocyte/macrophage serine esterase 1) | 13281 | -0.399 | -0.0631 | No |
| 367 | ABCC4 | ABCC4 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 4 | 13299 | -0.446 | -0.0552 | No |
| 368 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 13301 | -0.450 | -0.0460 | No |
| 369 | GADD45G | GADD45G Entrez, Source | growth arrest and DNA-damage-inducible, gamma | 13302 | -0.454 | -0.0366 | No |
| 370 | PPP1R3C | PPP1R3C Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 3C | 13309 | -0.475 | -0.0273 | No |
| 371 | CD74 | CD74 Entrez, Source | CD74 molecule, major histocompatibility complex, class II invariant chain | 13325 | -0.617 | -0.0157 | No |
| 372 | EDG1 | EDG1 Entrez, Source | endothelial differentiation, sphingolipid G-protein-coupled receptor, 1 | 13334 | -0.822 | 0.0006 | No |