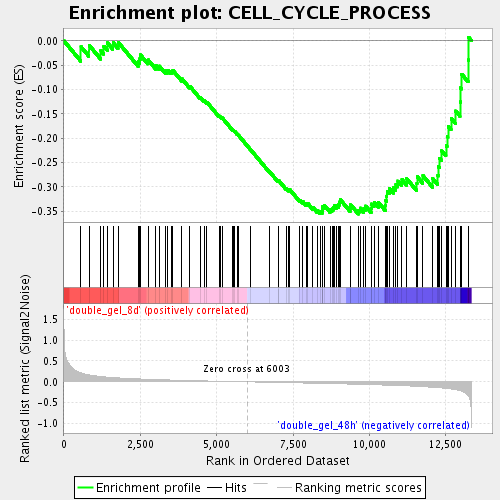
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | CELL_CYCLE_PROCESS |
| Enrichment Score (ES) | -0.35593423 |
| Normalized Enrichment Score (NES) | -1.3880615 |
| Nominal p-value | 0.032258064 |
| FDR q-value | 0.3490879 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 558 | 0.215 | -0.0122 | No |
| 2 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 821 | 0.166 | -0.0087 | No |
| 3 | PTPRC | PTPRC Entrez, Source | protein tyrosine phosphatase, receptor type, C | 1207 | 0.127 | -0.0201 | No |
| 4 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 1305 | 0.119 | -0.0107 | No |
| 5 | NPM2 | NPM2 Entrez, Source | nucleophosmin/nucleoplasmin, 2 | 1422 | 0.112 | -0.0039 | No |
| 6 | APBB1 | APBB1 Entrez, Source | amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) | 1606 | 0.102 | -0.0034 | No |
| 7 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 1773 | 0.094 | -0.0028 | No |
| 8 | TGFA | TGFA Entrez, Source | transforming growth factor, alpha | 2430 | 0.070 | -0.0425 | No |
| 9 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 2485 | 0.069 | -0.0370 | No |
| 10 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 2499 | 0.068 | -0.0285 | No |
| 11 | STAG3 | STAG3 Entrez, Source | stromal antigen 3 | 2752 | 0.061 | -0.0390 | No |
| 12 | EREG | EREG Entrez, Source | epiregulin | 3011 | 0.054 | -0.0509 | No |
| 13 | RAD17 | RAD17 Entrez, Source | RAD17 homolog (S. pombe) | 3116 | 0.051 | -0.0516 | No |
| 14 | CCNA1 | CCNA1 Entrez, Source | cyclin A1 | 3333 | 0.046 | -0.0614 | No |
| 15 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 3401 | 0.045 | -0.0602 | No |
| 16 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 3506 | 0.043 | -0.0621 | No |
| 17 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 3563 | 0.042 | -0.0605 | No |
| 18 | GFI1 | GFI1 Entrez, Source | growth factor independent 1 | 3862 | 0.035 | -0.0781 | No |
| 19 | USH1C | USH1C Entrez, Source | Usher syndrome 1C (autosomal recessive, severe) | 4117 | 0.030 | -0.0930 | No |
| 20 | DUSP13 | DUSP13 Entrez, Source | dual specificity phosphatase 13 | 4468 | 0.024 | -0.1161 | No |
| 21 | CDKN2B | CDKN2B Entrez, Source | cyclin-dependent kinase inhibitor 2B (p15, inhibits CDK4) | 4592 | 0.022 | -0.1223 | No |
| 22 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 4680 | 0.020 | -0.1260 | No |
| 23 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 5081 | 0.014 | -0.1543 | No |
| 24 | PPP5C | PPP5C Entrez, Source | protein phosphatase 5, catalytic subunit | 5131 | 0.013 | -0.1562 | No |
| 25 | CDKN2D | CDKN2D Entrez, Source | cyclin-dependent kinase inhibitor 2D (p19, inhibits CDK4) | 5179 | 0.012 | -0.1581 | No |
| 26 | SYCP1 | SYCP1 Entrez, Source | synaptonemal complex protein 1 | 5530 | 0.007 | -0.1835 | No |
| 27 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 5540 | 0.007 | -0.1833 | No |
| 28 | NBN | NBN Entrez, Source | nibrin | 5580 | 0.006 | -0.1854 | No |
| 29 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 5669 | 0.005 | -0.1914 | No |
| 30 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 5704 | 0.004 | -0.1933 | No |
| 31 | CHFR | CHFR Entrez, Source | checkpoint with forkhead and ring finger domains | 6104 | -0.002 | -0.2232 | No |
| 32 | CDK10 | CDK10 Entrez, Source | cyclin-dependent kinase (CDC2-like) 10 | 6722 | -0.011 | -0.2682 | No |
| 33 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 7011 | -0.015 | -0.2879 | No |
| 34 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 7015 | -0.015 | -0.2860 | No |
| 35 | DCTN2 | DCTN2 Entrez, Source | dynactin 2 (p50) | 7286 | -0.019 | -0.3036 | No |
| 36 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 7356 | -0.020 | -0.3060 | No |
| 37 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 7386 | -0.021 | -0.3053 | No |
| 38 | CENTD2 | CENTD2 Entrez, Source | centaurin, delta 2 | 7725 | -0.026 | -0.3272 | No |
| 39 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 7811 | -0.028 | -0.3297 | No |
| 40 | SMC1A | SMC1A Entrez, Source | structural maintenance of chromosomes 1A | 7928 | -0.030 | -0.3344 | No |
| 41 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 7971 | -0.030 | -0.3333 | No |
| 42 | ZWINT | ZWINT Entrez, Source | ZW10 interactor | 8148 | -0.033 | -0.3420 | No |
| 43 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 8298 | -0.036 | -0.3483 | No |
| 44 | SPDYA | SPDYA Entrez, Source | speedy homolog A (Drosophila) | 8392 | -0.037 | -0.3501 | Yes |
| 45 | TTN | TTN Entrez, Source | titin | 8450 | -0.038 | -0.3491 | Yes |
| 46 | SC65 | SC65 Entrez, Source | - | 8463 | -0.038 | -0.3446 | Yes |
| 47 | RAD50 | RAD50 Entrez, Source | RAD50 homolog (S. cerevisiae) | 8469 | -0.038 | -0.3397 | Yes |
| 48 | TBRG4 | TBRG4 Entrez, Source | transforming growth factor beta regulator 4 | 8526 | -0.039 | -0.3384 | Yes |
| 49 | CDC2L5 | CDC2L5 Entrez, Source | cell division cycle 2-like 5 (cholinesterase-related cell division controller) | 8711 | -0.042 | -0.3464 | Yes |
| 50 | NEK6 | NEK6 Entrez, Source | NIMA (never in mitosis gene a)-related kinase 6 | 8774 | -0.043 | -0.3450 | Yes |
| 51 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 8812 | -0.044 | -0.3417 | Yes |
| 52 | ANAPC4 | ANAPC4 Entrez, Source | anaphase promoting complex subunit 4 | 8855 | -0.045 | -0.3386 | Yes |
| 53 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 8927 | -0.046 | -0.3375 | Yes |
| 54 | NUMA1 | NUMA1 Entrez, Source | nuclear mitotic apparatus protein 1 | 8995 | -0.047 | -0.3360 | Yes |
| 55 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 9017 | -0.048 | -0.3310 | Yes |
| 56 | CETN3 | CETN3 Entrez, Source | centrin, EF-hand protein, 3 (CDC31 homolog, yeast) | 9041 | -0.048 | -0.3260 | Yes |
| 57 | CDC23 | CDC23 Entrez, Source | CDC23 (cell division cycle 23, yeast, homolog) | 9367 | -0.054 | -0.3431 | Yes |
| 58 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 9384 | -0.054 | -0.3368 | Yes |
| 59 | NDE1 | NDE1 Entrez, Source | nudE nuclear distribution gene E homolog 1 (A. nidulans) | 9639 | -0.058 | -0.3478 | Yes |
| 60 | BRCA2 | BRCA2 Entrez, Source | breast cancer 2, early onset | 9692 | -0.060 | -0.3434 | Yes |
| 61 | ZW10 | ZW10 Entrez, Source | ZW10, kinetochore associated, homolog (Drosophila) | 9814 | -0.062 | -0.3438 | Yes |
| 62 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 9872 | -0.064 | -0.3392 | Yes |
| 63 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 10056 | -0.067 | -0.3436 | Yes |
| 64 | SMC3 | SMC3 Entrez, Source | structural maintenance of chromosomes 3 | 10069 | -0.068 | -0.3351 | Yes |
| 65 | SUGT1 | SUGT1 Entrez, Source | SGT1, suppressor of G2 allele of SKP1 (S. cerevisiae) | 10151 | -0.069 | -0.3315 | Yes |
| 66 | CKAP5 | CKAP5 Entrez, Source | cytoskeleton associated protein 5 | 10289 | -0.072 | -0.3318 | Yes |
| 67 | CDC16 | CDC16 Entrez, Source | CDC16 cell division cycle 16 homolog (S. cerevisiae) | 10523 | -0.078 | -0.3384 | Yes |
| 68 | GSPT1 | GSPT1 Entrez, Source | G1 to S phase transition 1 | 10533 | -0.079 | -0.3282 | Yes |
| 69 | KHDRBS1 | KHDRBS1 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 1 | 10557 | -0.079 | -0.3189 | Yes |
| 70 | AKAP8 | AKAP8 Entrez, Source | A kinase (PRKA) anchor protein 8 | 10573 | -0.079 | -0.3089 | Yes |
| 71 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 10649 | -0.081 | -0.3033 | Yes |
| 72 | CETN1 | CETN1 Entrez, Source | centrin, EF-hand protein, 1 | 10779 | -0.085 | -0.3012 | Yes |
| 73 | LIG3 | LIG3 Entrez, Source | ligase III, DNA, ATP-dependent | 10864 | -0.087 | -0.2954 | Yes |
| 74 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 10930 | -0.089 | -0.2879 | Yes |
| 75 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 11062 | -0.092 | -0.2850 | Yes |
| 76 | CIT | CIT Entrez, Source | citron (rho-interacting, serine/threonine kinase 21) | 11218 | -0.096 | -0.2832 | Yes |
| 77 | PPP6C | PPP6C Entrez, Source | protein phosphatase 6, catalytic subunit | 11550 | -0.107 | -0.2932 | Yes |
| 78 | RAN | RAN Entrez, Source | RAN, member RAS oncogene family | 11559 | -0.108 | -0.2788 | Yes |
| 79 | HSPA2 | HSPA2 Entrez, Source | heat shock 70kDa protein 2 | 11748 | -0.115 | -0.2770 | Yes |
| 80 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 12066 | -0.130 | -0.2828 | Yes |
| 81 | CD28 | CD28 Entrez, Source | CD28 molecule | 12234 | -0.138 | -0.2761 | Yes |
| 82 | MAD2L2 | MAD2L2 Entrez, Source | MAD2 mitotic arrest deficient-like 2 (yeast) | 12256 | -0.140 | -0.2581 | Yes |
| 83 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 12292 | -0.142 | -0.2410 | Yes |
| 84 | SMC4 | SMC4 Entrez, Source | structural maintenance of chromosomes 4 | 12364 | -0.147 | -0.2259 | Yes |
| 85 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 12505 | -0.155 | -0.2149 | Yes |
| 86 | KATNA1 | KATNA1 Entrez, Source | katanin p60 (ATPase-containing) subunit A 1 | 12541 | -0.158 | -0.1955 | Yes |
| 87 | CDC27 | CDC27 Entrez, Source | cell division cycle 27 | 12571 | -0.160 | -0.1754 | Yes |
| 88 | AURKA | AURKA Entrez, Source | aurora kinase A | 12673 | -0.170 | -0.1593 | Yes |
| 89 | EGF | EGF Entrez, Source | epidermal growth factor (beta-urogastrone) | 12819 | -0.188 | -0.1440 | Yes |
| 90 | CUL5 | CUL5 Entrez, Source | cullin 5 | 12964 | -0.212 | -0.1252 | Yes |
| 91 | TIMELESS | TIMELESS Entrez, Source | timeless homolog (Drosophila) | 12978 | -0.214 | -0.0964 | Yes |
| 92 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 13015 | -0.221 | -0.0682 | Yes |
| 93 | MDM4 | MDM4 Entrez, Source | Mdm4, transformed 3T3 cell double minute 4, p53 binding protein (mouse) | 13232 | -0.326 | -0.0390 | Yes |
| 94 | MRE11A | MRE11A Entrez, Source | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | 13244 | -0.339 | 0.0074 | Yes |

