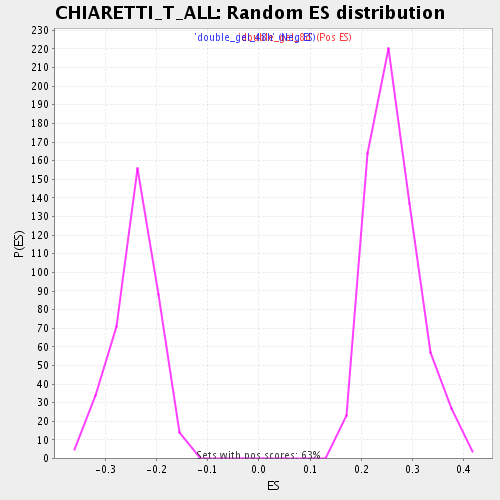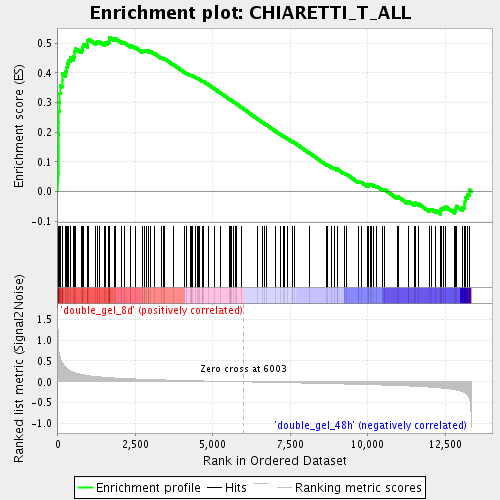
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | CHIARETTI_T_ALL |
| Enrichment Score (ES) | 0.5204245 |
| Normalized Enrichment Score (NES) | 1.983898 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.005297919 |
| FWER p-Value | 0.249 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGFBP7 | IGFBP7 Entrez, Source | insulin-like growth factor binding protein 7 | 2 | 1.254 | 0.0615 | Yes |
| 2 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 9 | 0.936 | 0.1071 | Yes |
| 3 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 10 | 0.885 | 0.1506 | Yes |
| 4 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 11 | 0.884 | 0.1940 | Yes |
| 5 | ARMCX2 | ARMCX2 Entrez, Source | armadillo repeat containing, X-linked 2 | 20 | 0.787 | 0.2320 | Yes |
| 6 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 25 | 0.771 | 0.2697 | Yes |
| 7 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 43 | 0.675 | 0.3015 | Yes |
| 8 | LGALS1 | LGALS1 Entrez, Source | lectin, galactoside-binding, soluble, 1 (galectin 1) | 60 | 0.614 | 0.3305 | Yes |
| 9 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 75 | 0.565 | 0.3572 | Yes |
| 10 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 138 | 0.458 | 0.3750 | Yes |
| 11 | SERPINE2 | SERPINE2 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 2 | 140 | 0.455 | 0.3973 | Yes |
| 12 | AHNAK | AHNAK Entrez, Source | AHNAK nucleoprotein (desmoyokin) | 255 | 0.338 | 0.4053 | Yes |
| 13 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 286 | 0.317 | 0.4186 | Yes |
| 14 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 310 | 0.299 | 0.4315 | Yes |
| 15 | CCL3 | CCL3 Entrez, Source | chemokine (C-C motif) ligand 3 | 353 | 0.281 | 0.4421 | Yes |
| 16 | PTGER4 | PTGER4 Entrez, Source | prostaglandin E receptor 4 (subtype EP4) | 402 | 0.258 | 0.4512 | Yes |
| 17 | PF4 | PF4 Entrez, Source | platelet factor 4 (chemokine (C-X-C motif) ligand 4) | 506 | 0.226 | 0.4545 | Yes |
| 18 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 538 | 0.219 | 0.4629 | Yes |
| 19 | ALCAM | ALCAM Entrez, Source | activated leukocyte cell adhesion molecule | 539 | 0.219 | 0.4737 | Yes |
| 20 | PLAGL1 | PLAGL1 Entrez, Source | pleiomorphic adenoma gene-like 1 | 576 | 0.211 | 0.4813 | Yes |
| 21 | EMP3 | EMP3 Entrez, Source | epithelial membrane protein 3 | 755 | 0.176 | 0.4765 | Yes |
| 22 | SPRY2 | SPRY2 Entrez, Source | sprouty homolog 2 (Drosophila) | 784 | 0.172 | 0.4828 | Yes |
| 23 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 808 | 0.169 | 0.4894 | Yes |
| 24 | PTTG1IP | PTTG1IP Entrez, Source | pituitary tumor-transforming 1 interacting protein | 811 | 0.168 | 0.4975 | Yes |
| 25 | PDLIM1 | PDLIM1 Entrez, Source | PDZ and LIM domain 1 (elfin) | 957 | 0.151 | 0.4939 | Yes |
| 26 | S100A8 | S100A8 Entrez, Source | S100 calcium binding protein A8 | 960 | 0.151 | 0.5012 | Yes |
| 27 | S100A4 | S100A4 Entrez, Source | S100 calcium binding protein A4 | 962 | 0.151 | 0.5085 | Yes |
| 28 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 999 | 0.147 | 0.5130 | Yes |
| 29 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 1218 | 0.126 | 0.5027 | Yes |
| 30 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 1261 | 0.123 | 0.5055 | Yes |
| 31 | NID2 | NID2 Entrez, Source | nidogen 2 (osteonidogen) | 1349 | 0.116 | 0.5047 | Yes |
| 32 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 1504 | 0.108 | 0.4983 | Yes |
| 33 | PLK2 | PLK2 Entrez, Source | polo-like kinase 2 (Drosophila) | 1523 | 0.107 | 0.5022 | Yes |
| 34 | TES | TES Entrez, Source | testis derived transcript (3 LIM domains) | 1616 | 0.102 | 0.5002 | Yes |
| 35 | ADFP | ADFP Entrez, Source | adipose differentiation-related protein | 1626 | 0.101 | 0.5045 | Yes |
| 36 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 1657 | 0.100 | 0.5071 | Yes |
| 37 | PRKCA | PRKCA Entrez, Source | protein kinase C, alpha | 1663 | 0.099 | 0.5116 | Yes |
| 38 | PTPRE | PTPRE Entrez, Source | protein tyrosine phosphatase, receptor type, E | 1672 | 0.099 | 0.5159 | Yes |
| 39 | HES1 | HES1 Entrez, Source | hairy and enhancer of split 1, (Drosophila) | 1677 | 0.098 | 0.5204 | Yes |
| 40 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 1841 | 0.091 | 0.5125 | No |
| 41 | CXADR | CXADR Entrez, Source | coxsackie virus and adenovirus receptor | 1853 | 0.090 | 0.5162 | No |
| 42 | ALAS2 | ALAS2 Entrez, Source | aminolevulinate, delta-, synthase 2 (sideroblastic/hypochromic anemia) | 2058 | 0.083 | 0.5048 | No |
| 43 | CTBP2 | CTBP2 Entrez, Source | C-terminal binding protein 2 | 2137 | 0.079 | 0.5028 | No |
| 44 | SNF1LK | SNF1LK Entrez, Source | SNF1-like kinase | 2345 | 0.073 | 0.4907 | No |
| 45 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 2355 | 0.072 | 0.4935 | No |
| 46 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 2499 | 0.068 | 0.4860 | No |
| 47 | CTSH | CTSH Entrez, Source | cathepsin H | 2723 | 0.062 | 0.4722 | No |
| 48 | DUSP2 | DUSP2 Entrez, Source | dual specificity phosphatase 2 | 2724 | 0.062 | 0.4752 | No |
| 49 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 2779 | 0.061 | 0.4741 | No |
| 50 | BIRC3 | BIRC3 Entrez, Source | baculoviral IAP repeat-containing 3 | 2803 | 0.060 | 0.4753 | No |
| 51 | CEBPB | CEBPB Entrez, Source | CCAAT/enhancer binding protein (C/EBP), beta | 2848 | 0.059 | 0.4749 | No |
| 52 | MINPP1 | MINPP1 Entrez, Source | multiple inositol polyphosphate histidine phosphatase, 1 | 2918 | 0.057 | 0.4725 | No |
| 53 | ITPR3 | ITPR3 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 3 | 2923 | 0.057 | 0.4750 | No |
| 54 | S100A10 | S100A10 Entrez, Source | S100 calcium binding protein A10 | 2981 | 0.055 | 0.4733 | No |
| 55 | MAL | MAL Entrez, Source | mal, T-cell differentiation protein | 3112 | 0.052 | 0.4660 | No |
| 56 | GAB2 | GAB2 Entrez, Source | GRB2-associated binding protein 2 | 3334 | 0.046 | 0.4516 | No |
| 57 | FRMD4B | FRMD4B Entrez, Source | FERM domain containing 4B | 3422 | 0.045 | 0.4472 | No |
| 58 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 3428 | 0.044 | 0.4490 | No |
| 59 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 3732 | 0.038 | 0.4279 | No |
| 60 | CD69 | CD69 Entrez, Source | CD69 molecule | 4095 | 0.031 | 0.4020 | No |
| 61 | BCL11A | BCL11A Entrez, Source | B-cell CLL/lymphoma 11A (zinc finger protein) | 4147 | 0.030 | 0.3996 | No |
| 62 | FHL1 | FHL1 Entrez, Source | four and a half LIM domains 1 | 4269 | 0.028 | 0.3918 | No |
| 63 | EPHB6 | EPHB6 Entrez, Source | EPH receptor B6 | 4295 | 0.027 | 0.3913 | No |
| 64 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 4301 | 0.027 | 0.3923 | No |
| 65 | CD97 | CD97 Entrez, Source | CD97 molecule | 4342 | 0.026 | 0.3905 | No |
| 66 | HIST1H2BL | HIST1H2BL Entrez, Source | histone cluster 1, H2bl | 4424 | 0.025 | 0.3856 | No |
| 67 | IL4R | IL4R Entrez, Source | interleukin 4 receptor | 4494 | 0.024 | 0.3815 | No |
| 68 | WASF1 | WASF1 Entrez, Source | WAS protein family, member 1 | 4524 | 0.023 | 0.3805 | No |
| 69 | STK17B | STK17B Entrez, Source | serine/threonine kinase 17b (apoptosis-inducing) | 4570 | 0.022 | 0.3781 | No |
| 70 | ALOX5 | ALOX5 Entrez, Source | arachidonate 5-lipoxygenase | 4658 | 0.021 | 0.3726 | No |
| 71 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 4680 | 0.020 | 0.3720 | No |
| 72 | NOTCH3 | NOTCH3 Entrez, Source | Notch homolog 3 (Drosophila) | 4689 | 0.020 | 0.3724 | No |
| 73 | TSHR | TSHR Entrez, Source | thyroid stimulating hormone receptor | 4846 | 0.018 | 0.3614 | No |
| 74 | ZBTB16 | ZBTB16 Entrez, Source | zinc finger and BTB domain containing 16 | 5042 | 0.014 | 0.3474 | No |
| 75 | GUCY1A3 | GUCY1A3 Entrez, Source | guanylate cyclase 1, soluble, alpha 3 | 5232 | 0.011 | 0.3336 | No |
| 76 | CDC42 | CDC42 Entrez, Source | cell division cycle 42 (GTP binding protein, 25kDa) | 5548 | 0.007 | 0.3101 | No |
| 77 | PRDX2 | PRDX2 Entrez, Source | peroxiredoxin 2 | 5562 | 0.006 | 0.3094 | No |
| 78 | CASP8 | CASP8 Entrez, Source | caspase 8, apoptosis-related cysteine peptidase | 5583 | 0.006 | 0.3082 | No |
| 79 | SCG2 | SCG2 Entrez, Source | secretogranin II (chromogranin C) | 5614 | 0.005 | 0.3062 | No |
| 80 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 5682 | 0.005 | 0.3013 | No |
| 81 | HPGD | HPGD Entrez, Source | hydroxyprostaglandin dehydrogenase 15-(NAD) | 5728 | 0.004 | 0.2981 | No |
| 82 | SLC4A1 | SLC4A1 Entrez, Source | solute carrier family 4, anion exchanger, member 1 (erythrocyte membrane protein band 3, Diego blood group) | 5762 | 0.003 | 0.2958 | No |
| 83 | FXYD2 | FXYD2 Entrez, Source | FXYD domain containing ion transport regulator 2 | 5938 | 0.001 | 0.2826 | No |
| 84 | AQP3 | AQP3 Entrez, Source | aquaporin 3 (Gill blood group) | 6430 | -0.007 | 0.2457 | No |
| 85 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 6595 | -0.009 | 0.2337 | No |
| 86 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 6682 | -0.010 | 0.2277 | No |
| 87 | ARID5A | ARID5A Entrez, Source | AT rich interactive domain 5A (MRF1-like) | 6737 | -0.011 | 0.2242 | No |
| 88 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 7011 | -0.015 | 0.2043 | No |
| 89 | MX1 | MX1 Entrez, Source | myxovirus (influenza virus) resistance 1, interferon-inducible protein p78 (mouse) | 7189 | -0.018 | 0.1917 | No |
| 90 | PPBP | PPBP Entrez, Source | pro-platelet basic protein (chemokine (C-X-C motif) ligand 7) | 7274 | -0.019 | 0.1863 | No |
| 91 | CD2 | CD2 Entrez, Source | CD2 molecule | 7323 | -0.020 | 0.1837 | No |
| 92 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 7402 | -0.021 | 0.1788 | No |
| 93 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 7557 | -0.023 | 0.1683 | No |
| 94 | CD200 | CD200 Entrez, Source | CD200 molecule | 7585 | -0.024 | 0.1674 | No |
| 95 | IL12A | IL12A Entrez, Source | interleukin 12A (natural killer cell stimulatory factor 1, cytotoxic lymphocyte maturation factor 1, p35) | 7646 | -0.025 | 0.1641 | No |
| 96 | SPON1 | SPON1 Entrez, Source | spondin 1, extracellular matrix protein | 8114 | -0.032 | 0.1303 | No |
| 97 | HNRPDL | HNRPDL Entrez, Source | heterogeneous nuclear ribonucleoprotein D-like | 8663 | -0.041 | 0.0908 | No |
| 98 | LMO2 | LMO2 Entrez, Source | LIM domain only 2 (rhombotin-like 1) | 8708 | -0.042 | 0.0896 | No |
| 99 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 8829 | -0.044 | 0.0827 | No |
| 100 | CCL5 | CCL5 Entrez, Source | chemokine (C-C motif) ligand 5 | 8913 | -0.046 | 0.0786 | No |
| 101 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 9007 | -0.047 | 0.0739 | No |
| 102 | RAP2B | RAP2B Entrez, Source | RAP2B, member of RAS oncogene family | 9009 | -0.047 | 0.0762 | No |
| 103 | HHEX | HHEX Entrez, Source | homeobox, hematopoietically expressed | 9234 | -0.051 | 0.0617 | No |
| 104 | ST18 | ST18 Entrez, Source | suppression of tumorigenicity 18 (breast carcinoma) (zinc finger protein) | 9316 | -0.053 | 0.0582 | No |
| 105 | SELL | SELL Entrez, Source | selectin L (lymphocyte adhesion molecule 1) | 9693 | -0.060 | 0.0327 | No |
| 106 | AHR | AHR Entrez, Source | aryl hydrocarbon receptor | 9715 | -0.060 | 0.0340 | No |
| 107 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 9785 | -0.062 | 0.0318 | No |
| 108 | PRG1 | PRG1 Entrez, Source | proteoglycan 1, secretory granule | 10004 | -0.066 | 0.0186 | No |
| 109 | IGJ | IGJ Entrez, Source | immunoglobulin J polypeptide, linker protein for immunoglobulin alpha and mu polypeptides | 10022 | -0.067 | 0.0206 | No |
| 110 | GARNL1 | GARNL1 Entrez, Source | GTPase activating Rap/RanGAP domain-like 1 | 10028 | -0.067 | 0.0235 | No |
| 111 | CD52 | CD52 Entrez, Source | CD52 molecule | 10093 | -0.068 | 0.0220 | No |
| 112 | ARF6 | ARF6 Entrez, Source | ADP-ribosylation factor 6 | 10133 | -0.069 | 0.0224 | No |
| 113 | RASGRP1 | RASGRP1 Entrez, Source | RAS guanyl releasing protein 1 (calcium and DAG-regulated) | 10196 | -0.070 | 0.0212 | No |
| 114 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 10294 | -0.072 | 0.0174 | No |
| 115 | CLEC11A | CLEC11A Entrez, Source | C-type lectin domain family 11, member A | 10483 | -0.077 | 0.0070 | No |
| 116 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 10547 | -0.079 | 0.0061 | No |
| 117 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 10960 | -0.089 | -0.0207 | No |
| 118 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 10976 | -0.090 | -0.0175 | No |
| 119 | RCBTB2 | RCBTB2 Entrez, Source | regulator of chromosome condensation (RCC1) and BTB (POZ) domain containing protein 2 | 11303 | -0.099 | -0.0373 | No |
| 120 | LAT | LAT Entrez, Source | linker for activation of T cells | 11315 | -0.099 | -0.0332 | No |
| 121 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 11513 | -0.106 | -0.0429 | No |
| 122 | CD8A | CD8A Entrez, Source | CD8a molecule | 11524 | -0.106 | -0.0385 | No |
| 123 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 11645 | -0.111 | -0.0421 | No |
| 124 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 11986 | -0.126 | -0.0617 | No |
| 125 | SKAP2 | SKAP2 Entrez, Source | src kinase associated phosphoprotein 2 | 12042 | -0.128 | -0.0595 | No |
| 126 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 12183 | -0.136 | -0.0635 | No |
| 127 | QKI | QKI Entrez, Source | quaking homolog, KH domain RNA binding (mouse) | 12354 | -0.146 | -0.0692 | No |
| 128 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 12360 | -0.146 | -0.0624 | No |
| 129 | CSDA | CSDA Entrez, Source | cold shock domain protein A | 12392 | -0.148 | -0.0574 | No |
| 130 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 12453 | -0.151 | -0.0545 | No |
| 131 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 12523 | -0.156 | -0.0521 | No |
| 132 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 12802 | -0.185 | -0.0640 | No |
| 133 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 12841 | -0.191 | -0.0575 | No |
| 134 | TAP1 | TAP1 Entrez, Source | transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) | 12854 | -0.192 | -0.0490 | No |
| 135 | PSPH | PSPH Entrez, Source | phosphoserine phosphatase | 13062 | -0.238 | -0.0529 | No |
| 136 | ARNTL | ARNTL Entrez, Source | aryl hydrocarbon receptor nuclear translocator-like | 13133 | -0.262 | -0.0454 | No |
| 137 | G0S2 | G0S2 Entrez, Source | G0/G1switch 2 | 13137 | -0.265 | -0.0326 | No |
| 138 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 13158 | -0.275 | -0.0206 | No |
| 139 | SFPQ | SFPQ Entrez, Source | splicing factor proline/glutamine-rich (polypyrimidine tract binding protein associated) | 13213 | -0.310 | -0.0094 | No |
| 140 | CLIC4 | CLIC4 Entrez, Source | chloride intracellular channel 4 | 13274 | -0.389 | 0.0052 | No |