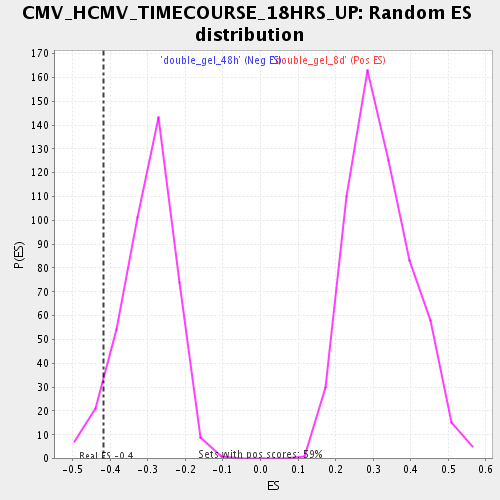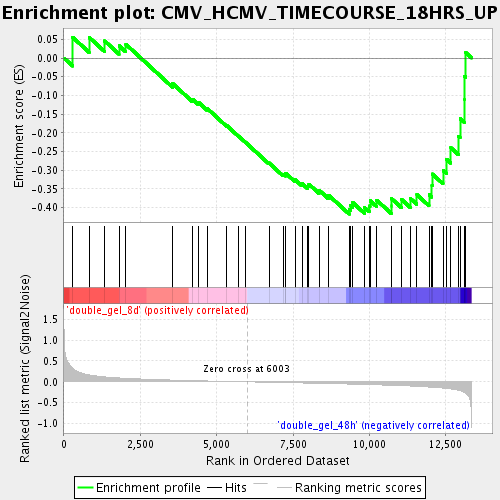
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | CMV_HCMV_TIMECOURSE_18HRS_UP |
| Enrichment Score (ES) | -0.41708538 |
| Normalized Enrichment Score (NES) | -1.3941808 |
| Nominal p-value | 0.053658538 |
| FDR q-value | 0.35252318 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 286 | 0.317 | 0.0555 | No |
| 2 | NEU1 | NEU1 Entrez, Source | sialidase 1 (lysosomal sialidase) | 827 | 0.166 | 0.0552 | No |
| 3 | WDR47 | WDR47 Entrez, Source | WD repeat domain 47 | 1333 | 0.117 | 0.0458 | No |
| 4 | BRAF | BRAF Entrez, Source | v-raf murine sarcoma viral oncogene homolog B1 | 1804 | 0.093 | 0.0330 | No |
| 5 | PTGER2 | PTGER2 Entrez, Source | prostaglandin E receptor 2 (subtype EP2), 53kDa | 2027 | 0.084 | 0.0368 | No |
| 6 | PDLIM3 | PDLIM3 Entrez, Source | PDZ and LIM domain 3 | 3560 | 0.042 | -0.0683 | No |
| 7 | OLFM1 | OLFM1 Entrez, Source | olfactomedin 1 | 4208 | 0.029 | -0.1099 | No |
| 8 | DYNC1I1 | DYNC1I1 Entrez, Source | dynein, cytoplasmic 1, intermediate chain 1 | 4395 | 0.025 | -0.1177 | No |
| 9 | RCHY1 | RCHY1 Entrez, Source | ring finger and CHY zinc finger domain containing 1 | 4687 | 0.020 | -0.1346 | No |
| 10 | SON | SON Entrez, Source | SON DNA binding protein | 5330 | 0.010 | -0.1806 | No |
| 11 | DLG1 | DLG1 Entrez, Source | discs, large homolog 1 (Drosophila) | 5729 | 0.004 | -0.2096 | No |
| 12 | RNF2 | RNF2 Entrez, Source | ring finger protein 2 | 5936 | 0.001 | -0.2248 | No |
| 13 | TMF1 | TMF1 Entrez, Source | TATA element modulatory factor 1 | 6711 | -0.011 | -0.2804 | No |
| 14 | CCNC | CCNC Entrez, Source | cyclin C | 7174 | -0.018 | -0.3108 | No |
| 15 | MEF2D | MEF2D Entrez, Source | MADS box transcription enhancer factor 2, polypeptide D (myocyte enhancer factor 2D) | 7239 | -0.019 | -0.3110 | No |
| 16 | RFK | RFK Entrez, Source | riboflavin kinase | 7256 | -0.019 | -0.3076 | No |
| 17 | DMN | DMN Entrez, Source | desmuslin | 7569 | -0.024 | -0.3253 | No |
| 18 | STAM2 | STAM2 Entrez, Source | signal transducing adaptor molecule (SH3 domain and ITAM motif) 2 | 7791 | -0.027 | -0.3352 | No |
| 19 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 7982 | -0.030 | -0.3421 | No |
| 20 | BRMS1 | BRMS1 Entrez, Source | breast cancer metastasis suppressor 1 | 8014 | -0.031 | -0.3369 | No |
| 21 | DR1 | DR1 Entrez, Source | down-regulator of transcription 1, TBP-binding (negative cofactor 2) | 8351 | -0.037 | -0.3532 | No |
| 22 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 8660 | -0.041 | -0.3663 | No |
| 23 | ISL1 | ISL1 Entrez, Source | ISL1 transcription factor, LIM/homeodomain, (islet-1) | 9336 | -0.053 | -0.4041 | Yes |
| 24 | CRLF3 | CRLF3 Entrez, Source | cytokine receptor-like factor 3 | 9382 | -0.054 | -0.3944 | Yes |
| 25 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 9451 | -0.055 | -0.3861 | Yes |
| 26 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 9836 | -0.063 | -0.3997 | Yes |
| 27 | ZBTB1 | ZBTB1 Entrez, Source | zinc finger and BTB domain containing 1 | 9994 | -0.066 | -0.3953 | Yes |
| 28 | KCNS3 | KCNS3 Entrez, Source | potassium voltage-gated channel, delayed-rectifier, subfamily S, member 3 | 10019 | -0.067 | -0.3809 | Yes |
| 29 | STCH | STCH Entrez, Source | stress 70 protein chaperone, microsome-associated, 60kDa | 10236 | -0.071 | -0.3798 | Yes |
| 30 | ATP6V1C1 | ATP6V1C1 Entrez, Source | ATPase, H+ transporting, lysosomal 42kDa, V1 subunit C1 | 10713 | -0.083 | -0.3954 | Yes |
| 31 | POLE3 | POLE3 Entrez, Source | polymerase (DNA directed), epsilon 3 (p17 subunit) | 10715 | -0.083 | -0.3753 | Yes |
| 32 | PPM1A | PPM1A Entrez, Source | protein phosphatase 1A (formerly 2C), magnesium-dependent, alpha isoform | 11053 | -0.092 | -0.3783 | Yes |
| 33 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 11338 | -0.100 | -0.3753 | Yes |
| 34 | SLC12A2 | SLC12A2 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 2 | 11551 | -0.108 | -0.3651 | Yes |
| 35 | ARHGEF5 | ARHGEF5 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 5 | 11950 | -0.124 | -0.3648 | Yes |
| 36 | RPS6KB1 | RPS6KB1 Entrez, Source | ribosomal protein S6 kinase, 70kDa, polypeptide 1 | 12044 | -0.129 | -0.3405 | Yes |
| 37 | TESK2 | TESK2 Entrez, Source | testis-specific kinase 2 | 12056 | -0.129 | -0.3098 | Yes |
| 38 | SSR1 | SSR1 Entrez, Source | signal sequence receptor, alpha (translocon-associated protein alpha) | 12408 | -0.149 | -0.3000 | Yes |
| 39 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 12523 | -0.156 | -0.2706 | Yes |
| 40 | TRIM23 | TRIM23 Entrez, Source | tripartite motif-containing 23 | 12662 | -0.169 | -0.2398 | Yes |
| 41 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 12915 | -0.202 | -0.2096 | Yes |
| 42 | CASP2 | CASP2 Entrez, Source | caspase 2, apoptosis-related cysteine peptidase (neural precursor cell expressed, developmentally down-regulated 2) | 12967 | -0.213 | -0.1616 | Yes |
| 43 | PPAT | PPAT Entrez, Source | phosphoribosyl pyrophosphate amidotransferase | 13102 | -0.253 | -0.1102 | Yes |
| 44 | FNBP4 | FNBP4 Entrez, Source | formin binding protein 4 | 13108 | -0.255 | -0.0484 | Yes |
| 45 | GCLC | GCLC Entrez, Source | glutamate-cysteine ligase, catalytic subunit | 13151 | -0.271 | 0.0144 | Yes |