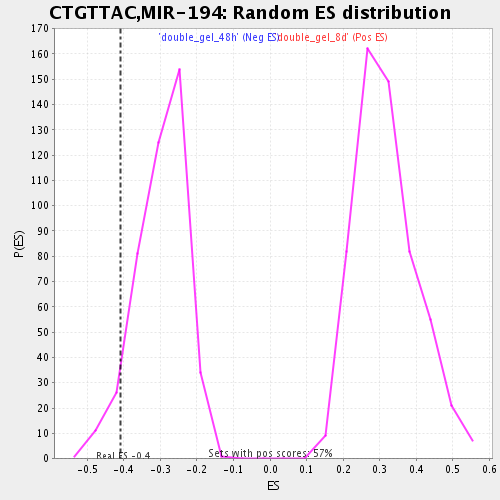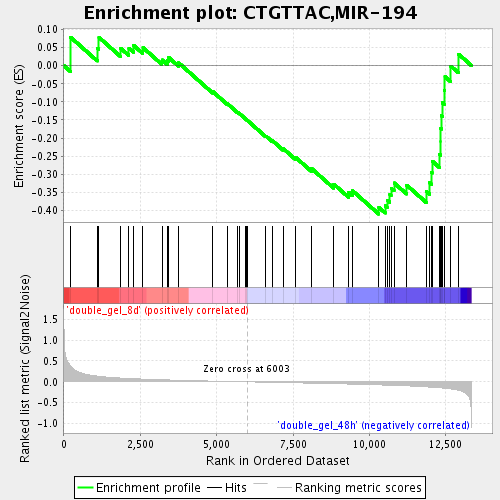
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | CTGTTAC,MIR-194 |
| Enrichment Score (ES) | -0.41024625 |
| Normalized Enrichment Score (NES) | -1.3779429 |
| Nominal p-value | 0.064665124 |
| FDR q-value | 0.36058083 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HS3ST2 | HS3ST2 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 2 | 219 | 0.370 | 0.0787 | No |
| 2 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 1094 | 0.137 | 0.0482 | No |
| 3 | PFKFB2 | PFKFB2 Entrez, Source | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 | 1146 | 0.132 | 0.0784 | No |
| 4 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 1849 | 0.091 | 0.0489 | No |
| 5 | TMEM43 | TMEM43 Entrez, Source | transmembrane protein 43 | 2127 | 0.080 | 0.0486 | No |
| 6 | SNX1 | SNX1 Entrez, Source | sorting nexin 1 | 2279 | 0.075 | 0.0564 | No |
| 7 | YTHDF1 | YTHDF1 Entrez, Source | YTH domain family, member 1 | 2587 | 0.065 | 0.0501 | No |
| 8 | FOXA1 | FOXA1 Entrez, Source | forkhead box A1 | 3210 | 0.049 | 0.0160 | No |
| 9 | SDC4 | SDC4 Entrez, Source | syndecan 4 (amphiglycan, ryudocan) | 3385 | 0.045 | 0.0146 | No |
| 10 | SOCS2 | SOCS2 Entrez, Source | suppressor of cytokine signaling 2 | 3428 | 0.044 | 0.0229 | No |
| 11 | PDHB | PDHB Entrez, Source | pyruvate dehydrogenase (lipoamide) beta | 3755 | 0.037 | 0.0080 | No |
| 12 | SUFU | SUFU Entrez, Source | suppressor of fused homolog (Drosophila) | 4870 | 0.017 | -0.0713 | No |
| 13 | TCF2 | TCF2 Entrez, Source | transcription factor 2, hepatic; LF-B3; variant hepatic nuclear factor | 5337 | 0.010 | -0.1039 | No |
| 14 | PTPN12 | PTPN12 Entrez, Source | protein tyrosine phosphatase, non-receptor type 12 | 5686 | 0.005 | -0.1290 | No |
| 15 | LIMK2 | LIMK2 Entrez, Source | LIM domain kinase 2 | 5743 | 0.004 | -0.1323 | No |
| 16 | CSNK1D | CSNK1D Entrez, Source | casein kinase 1, delta | 5941 | 0.001 | -0.1468 | No |
| 17 | TRIM9 | TRIM9 Entrez, Source | tripartite motif-containing 9 | 5979 | 0.000 | -0.1495 | No |
| 18 | GIPC1 | GIPC1 Entrez, Source | GIPC PDZ domain containing family, member 1 | 6006 | -0.000 | -0.1514 | No |
| 19 | GYG1 | GYG1 Entrez, Source | glycogenin 1 | 6609 | -0.009 | -0.1943 | No |
| 20 | SEPHS1 | SEPHS1 Entrez, Source | selenophosphate synthetase 1 | 6820 | -0.012 | -0.2069 | No |
| 21 | VAPA | VAPA Entrez, Source | VAMP (vesicle-associated membrane protein)-associated protein A, 33kDa | 7181 | -0.018 | -0.2294 | No |
| 22 | BCL2L11 | BCL2L11 Entrez, Source | BCL2-like 11 (apoptosis facilitator) | 7593 | -0.024 | -0.2541 | No |
| 23 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 8103 | -0.032 | -0.2841 | No |
| 24 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 8829 | -0.044 | -0.3272 | No |
| 25 | ST18 | ST18 Entrez, Source | suppression of tumorigenicity 18 (breast carcinoma) (zinc finger protein) | 9316 | -0.053 | -0.3502 | No |
| 26 | LGI1 | LGI1 Entrez, Source | leucine-rich, glioma inactivated 1 | 9443 | -0.055 | -0.3455 | No |
| 27 | MID1IP1 | MID1IP1 Entrez, Source | MID1 interacting protein 1 (gastrulation specific G12 homolog (zebrafish)) | 10305 | -0.073 | -0.3916 | Yes |
| 28 | SUMO2 | SUMO2 Entrez, Source | SMT3 suppressor of mif two 3 homolog 2 (S. cerevisiae) | 10516 | -0.078 | -0.3872 | Yes |
| 29 | DUSP9 | DUSP9 Entrez, Source | dual specificity phosphatase 9 | 10581 | -0.079 | -0.3716 | Yes |
| 30 | AP1G1 | AP1G1 Entrez, Source | adaptor-related protein complex 1, gamma 1 subunit | 10665 | -0.082 | -0.3569 | Yes |
| 31 | NUDC | NUDC Entrez, Source | nuclear distribution gene C homolog (A. nidulans) | 10710 | -0.083 | -0.3389 | Yes |
| 32 | GABRB3 | GABRB3 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 3 | 10807 | -0.085 | -0.3241 | Yes |
| 33 | ITSN1 | ITSN1 Entrez, Source | intersectin 1 (SH3 domain protein) | 11216 | -0.096 | -0.3301 | Yes |
| 34 | FMR1 | FMR1 Entrez, Source | fragile X mental retardation 1 | 11863 | -0.120 | -0.3478 | Yes |
| 35 | TRIP12 | TRIP12 Entrez, Source | thyroid hormone receptor interactor 12 | 11969 | -0.125 | -0.3236 | Yes |
| 36 | DNMT3A | DNMT3A Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 alpha | 12031 | -0.128 | -0.2953 | Yes |
| 37 | CHD4 | CHD4 Entrez, Source | chromodomain helicase DNA binding protein 4 | 12051 | -0.129 | -0.2636 | Yes |
| 38 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 12292 | -0.142 | -0.2452 | Yes |
| 39 | MECP2 | MECP2 Entrez, Source | methyl CpG binding protein 2 (Rett syndrome) | 12329 | -0.144 | -0.2108 | Yes |
| 40 | EXOC5 | EXOC5 Entrez, Source | exocyst complex component 5 | 12331 | -0.144 | -0.1737 | Yes |
| 41 | QKI | QKI Entrez, Source | quaking homolog, KH domain RNA binding (mouse) | 12354 | -0.146 | -0.1379 | Yes |
| 42 | GMFB | GMFB Entrez, Source | glia maturation factor, beta | 12387 | -0.148 | -0.1022 | Yes |
| 43 | HOOK3 | HOOK3 Entrez, Source | hook homolog 3 (Drosophila) | 12448 | -0.151 | -0.0679 | Yes |
| 44 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 12470 | -0.153 | -0.0301 | Yes |
| 45 | TRIM23 | TRIM23 Entrez, Source | tripartite motif-containing 23 | 12662 | -0.169 | -0.0010 | Yes |
| 46 | NCL | NCL Entrez, Source | nucleolin | 12916 | -0.202 | 0.0320 | Yes |