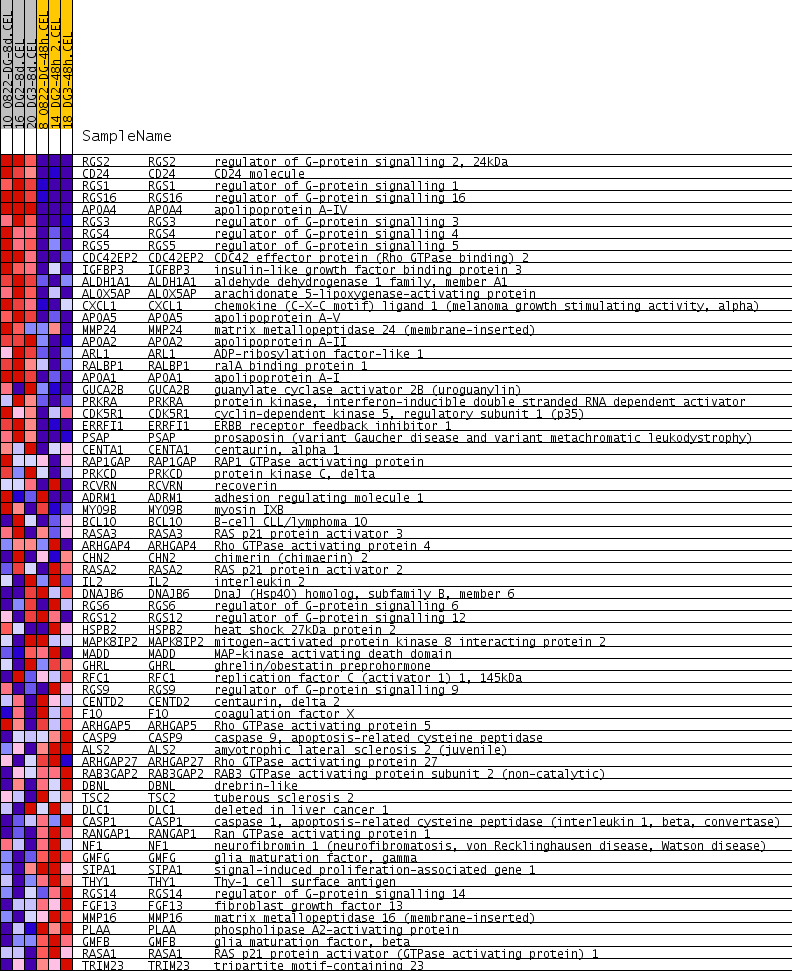
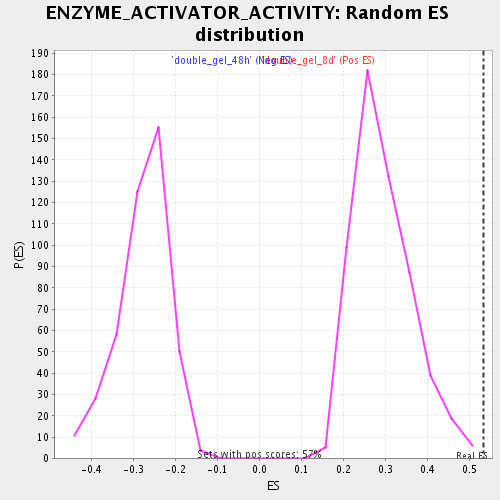
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | ENZYME_ACTIVATOR_ACTIVITY |
| Enrichment Score (ES) | 0.5346719 |
| Normalized Enrichment Score (NES) | 1.8164032 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.022690069 |
| FWER p-Value | 0.935 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 25 | 0.771 | 0.0959 | Yes |
| 2 | CD24 | CD24 Entrez, Source | CD24 molecule | 26 | 0.771 | 0.1937 | Yes |
| 3 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 75 | 0.565 | 0.2617 | Yes |
| 4 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 103 | 0.505 | 0.3238 | Yes |
| 5 | APOA4 | APOA4 Entrez, Source | apolipoprotein A-IV | 253 | 0.340 | 0.3556 | Yes |
| 6 | RGS3 | RGS3 Entrez, Source | regulator of G-protein signalling 3 | 338 | 0.287 | 0.3857 | Yes |
| 7 | RGS4 | RGS4 Entrez, Source | regulator of G-protein signalling 4 | 454 | 0.244 | 0.4080 | Yes |
| 8 | RGS5 | RGS5 Entrez, Source | regulator of G-protein signalling 5 | 513 | 0.224 | 0.4320 | Yes |
| 9 | CDC42EP2 | CDC42EP2 Entrez, Source | CDC42 effector protein (Rho GTPase binding) 2 | 522 | 0.222 | 0.4595 | Yes |
| 10 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 571 | 0.212 | 0.4828 | Yes |
| 11 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 739 | 0.178 | 0.4928 | Yes |
| 12 | ALOX5AP | ALOX5AP Entrez, Source | arachidonate 5-lipoxygenase-activating protein | 834 | 0.165 | 0.5066 | Yes |
| 13 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 1041 | 0.142 | 0.5091 | Yes |
| 14 | APOA5 | APOA5 Entrez, Source | apolipoprotein A-V | 1126 | 0.134 | 0.5198 | Yes |
| 15 | MMP24 | MMP24 Entrez, Source | matrix metallopeptidase 24 (membrane-inserted) | 1476 | 0.110 | 0.5074 | Yes |
| 16 | APOA2 | APOA2 Entrez, Source | apolipoprotein A-II | 1596 | 0.103 | 0.5115 | Yes |
| 17 | ARL1 | ARL1 Entrez, Source | ADP-ribosylation factor-like 1 | 1693 | 0.098 | 0.5166 | Yes |
| 18 | RALBP1 | RALBP1 Entrez, Source | ralA binding protein 1 | 1792 | 0.093 | 0.5211 | Yes |
| 19 | APOA1 | APOA1 Entrez, Source | apolipoprotein A-I | 1886 | 0.089 | 0.5254 | Yes |
| 20 | GUCA2B | GUCA2B Entrez, Source | guanylate cyclase activator 2B (uroguanylin) | 1912 | 0.088 | 0.5347 | Yes |
| 21 | PRKRA | PRKRA Entrez, Source | protein kinase, interferon-inducible double stranded RNA dependent activator | 2319 | 0.073 | 0.5134 | No |
| 22 | CDK5R1 | CDK5R1 Entrez, Source | cyclin-dependent kinase 5, regulatory subunit 1 (p35) | 2373 | 0.072 | 0.5185 | No |
| 23 | ERRFI1 | ERRFI1 Entrez, Source | ERBB receptor feedback inhibitor 1 | 2488 | 0.068 | 0.5186 | No |
| 24 | PSAP | PSAP Entrez, Source | prosaposin (variant Gaucher disease and variant metachromatic leukodystrophy) | 2642 | 0.064 | 0.5152 | No |
| 25 | CENTA1 | CENTA1 Entrez, Source | centaurin, alpha 1 | 3441 | 0.044 | 0.4606 | No |
| 26 | RAP1GAP | RAP1GAP Entrez, Source | RAP1 GTPase activating protein | 3624 | 0.040 | 0.4521 | No |
| 27 | PRKCD | PRKCD Entrez, Source | protein kinase C, delta | 3824 | 0.036 | 0.4416 | No |
| 28 | RCVRN | RCVRN Entrez, Source | recoverin | 4182 | 0.029 | 0.4185 | No |
| 29 | ADRM1 | ADRM1 Entrez, Source | adhesion regulating molecule 1 | 4443 | 0.025 | 0.4020 | No |
| 30 | MYO9B | MYO9B Entrez, Source | myosin IXB | 4486 | 0.024 | 0.4018 | No |
| 31 | BCL10 | BCL10 Entrez, Source | B-cell CLL/lymphoma 10 | 4947 | 0.016 | 0.3692 | No |
| 32 | RASA3 | RASA3 Entrez, Source | RAS p21 protein activator 3 | 5066 | 0.014 | 0.3621 | No |
| 33 | ARHGAP4 | ARHGAP4 Entrez, Source | Rho GTPase activating protein 4 | 5575 | 0.006 | 0.3246 | No |
| 34 | CHN2 | CHN2 Entrez, Source | chimerin (chimaerin) 2 | 5842 | 0.002 | 0.3049 | No |
| 35 | RASA2 | RASA2 Entrez, Source | RAS p21 protein activator 2 | 6217 | -0.003 | 0.2771 | No |
| 36 | IL2 | IL2 Entrez, Source | interleukin 2 | 6369 | -0.006 | 0.2665 | No |
| 37 | DNAJB6 | DNAJB6 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 6 | 6729 | -0.011 | 0.2409 | No |
| 38 | RGS6 | RGS6 Entrez, Source | regulator of G-protein signalling 6 | 6832 | -0.013 | 0.2348 | No |
| 39 | RGS12 | RGS12 Entrez, Source | regulator of G-protein signalling 12 | 6916 | -0.014 | 0.2303 | No |
| 40 | HSPB2 | HSPB2 Entrez, Source | heat shock 27kDa protein 2 | 6942 | -0.014 | 0.2302 | No |
| 41 | MAPK8IP2 | MAPK8IP2 Entrez, Source | mitogen-activated protein kinase 8 interacting protein 2 | 6981 | -0.015 | 0.2292 | No |
| 42 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 7149 | -0.017 | 0.2188 | No |
| 43 | GHRL | GHRL Entrez, Source | ghrelin/obestatin preprohormone | 7162 | -0.018 | 0.2202 | No |
| 44 | RFC1 | RFC1 Entrez, Source | replication factor C (activator 1) 1, 145kDa | 7389 | -0.021 | 0.2058 | No |
| 45 | RGS9 | RGS9 Entrez, Source | regulator of G-protein signalling 9 | 7456 | -0.022 | 0.2036 | No |
| 46 | CENTD2 | CENTD2 Entrez, Source | centaurin, delta 2 | 7725 | -0.026 | 0.1867 | No |
| 47 | F10 | F10 Entrez, Source | coagulation factor X | 7895 | -0.029 | 0.1777 | No |
| 48 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 8103 | -0.032 | 0.1662 | No |
| 49 | CASP9 | CASP9 Entrez, Source | caspase 9, apoptosis-related cysteine peptidase | 8297 | -0.036 | 0.1562 | No |
| 50 | ALS2 | ALS2 Entrez, Source | amyotrophic lateral sclerosis 2 (juvenile) | 8397 | -0.037 | 0.1534 | No |
| 51 | ARHGAP27 | ARHGAP27 Entrez, Source | Rho GTPase activating protein 27 | 8729 | -0.043 | 0.1339 | No |
| 52 | RAB3GAP2 | RAB3GAP2 Entrez, Source | RAB3 GTPase activating protein subunit 2 (non-catalytic) | 8768 | -0.043 | 0.1366 | No |
| 53 | DBNL | DBNL Entrez, Source | drebrin-like | 8828 | -0.044 | 0.1377 | No |
| 54 | TSC2 | TSC2 Entrez, Source | tuberous sclerosis 2 | 8912 | -0.046 | 0.1373 | No |
| 55 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 9141 | -0.050 | 0.1264 | No |
| 56 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 10182 | -0.070 | 0.0569 | No |
| 57 | RANGAP1 | RANGAP1 Entrez, Source | Ran GTPase activating protein 1 | 10603 | -0.080 | 0.0354 | No |
| 58 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 10739 | -0.084 | 0.0359 | No |
| 59 | GMFG | GMFG Entrez, Source | glia maturation factor, gamma | 11174 | -0.095 | 0.0153 | No |
| 60 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 11380 | -0.101 | 0.0127 | No |
| 61 | THY1 | THY1 Entrez, Source | Thy-1 cell surface antigen | 11400 | -0.102 | 0.0242 | No |
| 62 | RGS14 | RGS14 Entrez, Source | regulator of G-protein signalling 14 | 11575 | -0.108 | 0.0248 | No |
| 63 | FGF13 | FGF13 Entrez, Source | fibroblast growth factor 13 | 11831 | -0.118 | 0.0206 | No |
| 64 | MMP16 | MMP16 Entrez, Source | matrix metallopeptidase 16 (membrane-inserted) | 11993 | -0.126 | 0.0245 | No |
| 65 | PLAA | PLAA Entrez, Source | phospholipase A2-activating protein | 12215 | -0.137 | 0.0252 | No |
| 66 | GMFB | GMFB Entrez, Source | glia maturation factor, beta | 12387 | -0.148 | 0.0311 | No |
| 67 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 12457 | -0.152 | 0.0451 | No |
| 68 | TRIM23 | TRIM23 Entrez, Source | tripartite motif-containing 23 | 12662 | -0.169 | 0.0512 | No |