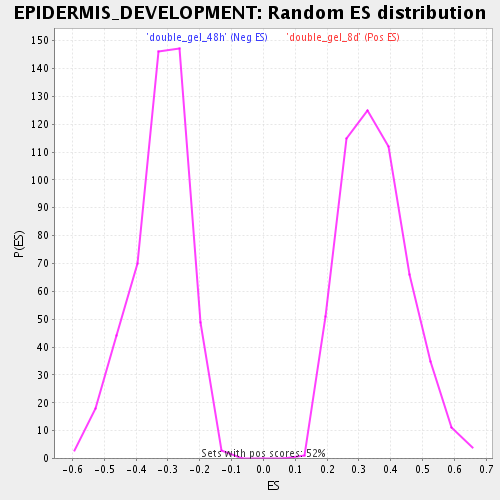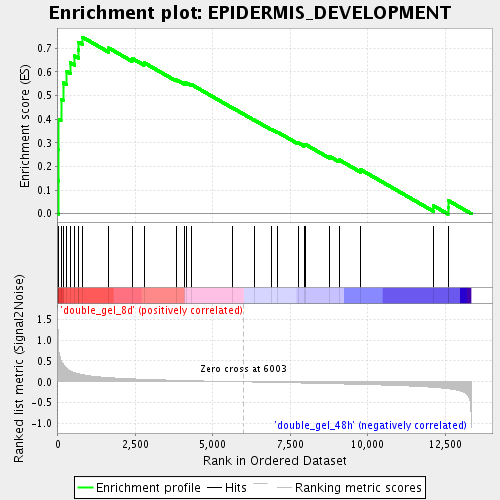
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | EPIDERMIS_DEVELOPMENT |
| Enrichment Score (ES) | 0.74659425 |
| Normalized Enrichment Score (NES) | 2.1214786 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0010795584 |
| FWER p-Value | 0.023 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LAMC2 | LAMC2 Entrez, Source | laminin, gamma 2 | 19 | 0.791 | 0.1389 | Yes |
| 2 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 27 | 0.757 | 0.2725 | Yes |
| 3 | COL5A2 | COL5A2 Entrez, Source | collagen, type V, alpha 2 | 34 | 0.714 | 0.3986 | Yes |
| 4 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 101 | 0.510 | 0.4840 | Yes |
| 5 | FST | FST Entrez, Source | follistatin | 169 | 0.423 | 0.5541 | Yes |
| 6 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 288 | 0.316 | 0.6013 | Yes |
| 7 | BTD | BTD Entrez, Source | biotinidase | 401 | 0.259 | 0.6387 | Yes |
| 8 | GJB5 | GJB5 Entrez, Source | gap junction protein, beta 5 (connexin 31.1) | 534 | 0.219 | 0.6677 | Yes |
| 9 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 659 | 0.195 | 0.6930 | Yes |
| 10 | COL1A1 | COL1A1 Entrez, Source | collagen, type I, alpha 1 | 673 | 0.190 | 0.7258 | Yes |
| 11 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 798 | 0.170 | 0.7466 | Yes |
| 12 | PLOD1 | PLOD1 Entrez, Source | procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1 | 1639 | 0.101 | 0.7013 | No |
| 13 | FLOT2 | FLOT2 Entrez, Source | flotillin 2 | 2393 | 0.071 | 0.6573 | No |
| 14 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 2779 | 0.061 | 0.6392 | No |
| 15 | RBP2 | RBP2 Entrez, Source | retinol binding protein 2, cellular | 3814 | 0.036 | 0.5679 | No |
| 16 | DSP | DSP Entrez, Source | desmoplakin | 4079 | 0.031 | 0.5536 | No |
| 17 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 4156 | 0.030 | 0.5533 | No |
| 18 | LAMA3 | LAMA3 Entrez, Source | laminin, alpha 3 | 4300 | 0.027 | 0.5474 | No |
| 19 | ATP2C1 | ATP2C1 Entrez, Source | ATPase, Ca++ transporting, type 2C, member 1 | 5638 | 0.005 | 0.4478 | No |
| 20 | GLI1 | GLI1 Entrez, Source | glioma-associated oncogene homolog 1 (zinc finger protein) | 6332 | -0.005 | 0.3967 | No |
| 21 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 6879 | -0.013 | 0.3580 | No |
| 22 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 7085 | -0.016 | 0.3456 | No |
| 23 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 7755 | -0.027 | 0.3000 | No |
| 24 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 7943 | -0.030 | 0.2913 | No |
| 25 | POU2F3 | POU2F3 Entrez, Source | POU domain, class 2, transcription factor 3 | 7981 | -0.030 | 0.2939 | No |
| 26 | PTHLH | PTHLH Entrez, Source | parathyroid hormone-like hormone | 8761 | -0.043 | 0.2430 | No |
| 27 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 9075 | -0.048 | 0.2281 | No |
| 28 | NME2 | NME2 Entrez, Source | non-metastatic cells 2, protein (NM23B) expressed in | 9773 | -0.061 | 0.1866 | No |
| 29 | FABP5 | FABP5 Entrez, Source | fatty acid binding protein 5 (psoriasis-associated) | 12121 | -0.133 | 0.0339 | No |
| 30 | KLK7 | KLK7 Entrez, Source | kallikrein 7 (chymotryptic, stratum corneum) | 12591 | -0.162 | 0.0274 | No |
| 31 | LAMB3 | LAMB3 Entrez, Source | laminin, beta 3 | 12600 | -0.163 | 0.0557 | No |