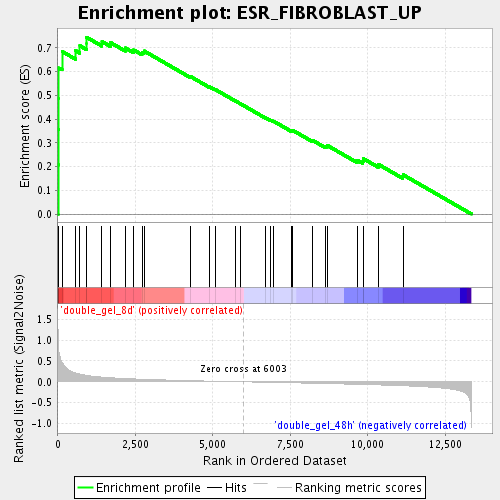
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | ESR_FIBROBLAST_UP |
| Enrichment Score (ES) | 0.7452171 |
| Normalized Enrichment Score (NES) | 2.1696372 |
| Nominal p-value | 0.0 |
| FDR q-value | 6.1850547E-4 |
| FWER p-Value | 0.0080 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VIM | VIM Entrez, Source | vimentin | 4 | 1.225 | 0.2070 | Yes |
| 2 | ANXA1 | ANXA1 Entrez, Source | annexin A1 | 11 | 0.884 | 0.3561 | Yes |
| 3 | COL3A1 | COL3A1 Entrez, Source | collagen, type III, alpha 1 (Ehlers-Danlos syndrome type IV, autosomal dominant) | 21 | 0.785 | 0.4882 | Yes |
| 4 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 27 | 0.757 | 0.6158 | Yes |
| 5 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 138 | 0.458 | 0.6851 | Yes |
| 6 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 571 | 0.212 | 0.6884 | Yes |
| 7 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 709 | 0.184 | 0.7093 | Yes |
| 8 | BNIP3L | BNIP3L Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3-like | 921 | 0.154 | 0.7194 | Yes |
| 9 | ARPC1B | ARPC1B Entrez, Source | actin related protein 2/3 complex, subunit 1B, 41kDa | 924 | 0.153 | 0.7452 | Yes |
| 10 | NGFRAP1 | NGFRAP1 Entrez, Source | nerve growth factor receptor (TNFRSF16) associated protein 1 | 1421 | 0.112 | 0.7269 | No |
| 11 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 1703 | 0.097 | 0.7222 | No |
| 12 | EFNB1 | EFNB1 Entrez, Source | ephrin-B1 | 2174 | 0.078 | 0.7002 | No |
| 13 | ZFP36L1 | ZFP36L1 Entrez, Source | zinc finger protein 36, C3H type-like 1 | 2445 | 0.070 | 0.6916 | No |
| 14 | FN1 | FN1 Entrez, Source | fibronectin 1 | 2715 | 0.062 | 0.6820 | No |
| 15 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 2779 | 0.061 | 0.6875 | No |
| 16 | AKAP12 | AKAP12 Entrez, Source | A kinase (PRKA) anchor protein (gravin) 12 | 4283 | 0.028 | 0.5792 | No |
| 17 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 4888 | 0.017 | 0.5368 | No |
| 18 | MARK3 | MARK3 Entrez, Source | MAP/microtubule affinity-regulating kinase 3 | 5075 | 0.014 | 0.5251 | No |
| 19 | TBCA | TBCA Entrez, Source | tubulin-specific chaperone a | 5734 | 0.004 | 0.4763 | No |
| 20 | FLOT1 | FLOT1 Entrez, Source | flotillin 1 | 5897 | 0.002 | 0.4644 | No |
| 21 | ATP5I | ATP5I Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit E | 6710 | -0.011 | 0.4052 | No |
| 22 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 6862 | -0.013 | 0.3961 | No |
| 23 | EIF4E | EIF4E Entrez, Source | eukaryotic translation initiation factor 4E | 6949 | -0.014 | 0.3921 | No |
| 24 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7545 | -0.023 | 0.3513 | No |
| 25 | TXNRD2 | TXNRD2 Entrez, Source | thioredoxin reductase 2 | 7578 | -0.024 | 0.3529 | No |
| 26 | OAZ1 | OAZ1 Entrez, Source | ornithine decarboxylase antizyme 1 | 8231 | -0.034 | 0.3098 | No |
| 27 | PRDX4 | PRDX4 Entrez, Source | peroxiredoxin 4 | 8646 | -0.041 | 0.2856 | No |
| 28 | GOLPH3 | GOLPH3 Entrez, Source | golgi phosphoprotein 3 (coat-protein) | 8690 | -0.042 | 0.2895 | No |
| 29 | TPR | TPR Entrez, Source | translocated promoter region (to activated MET oncogene) | 9660 | -0.059 | 0.2266 | No |
| 30 | FGFR4 | FGFR4 Entrez, Source | fibroblast growth factor receptor 4 | 9846 | -0.063 | 0.2234 | No |
| 31 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 9857 | -0.063 | 0.2333 | No |
| 32 | EIF3S6IP | EIF3S6IP Entrez, Source | eukaryotic translation initiation factor 3, subunit 6 interacting protein | 10336 | -0.073 | 0.2098 | No |
| 33 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 11137 | -0.094 | 0.1657 | No |

