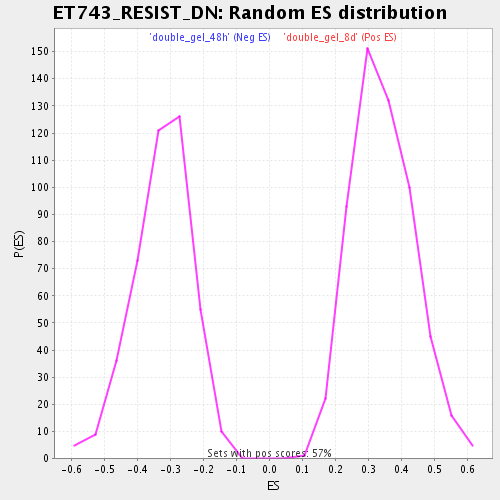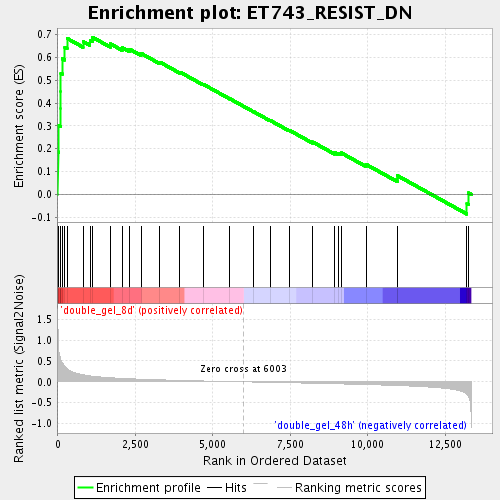
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | ET743_RESIST_DN |
| Enrichment Score (ES) | 0.68864214 |
| Normalized Enrichment Score (NES) | 2.0017374 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.004528611 |
| FWER p-Value | 0.197 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGFBP7 | IGFBP7 Entrez, Source | insulin-like growth factor binding protein 7 | 2 | 1.254 | 0.1859 | Yes |
| 2 | COL3A1 | COL3A1 Entrez, Source | collagen, type III, alpha 1 (Ehlers-Danlos syndrome type IV, autosomal dominant) | 21 | 0.785 | 0.3010 | Yes |
| 3 | FSTL1 | FSTL1 Entrez, Source | follistatin-like 1 | 91 | 0.531 | 0.3746 | Yes |
| 4 | ANKRD1 | ANKRD1 Entrez, Source | ankyrin repeat domain 1 (cardiac muscle) | 93 | 0.529 | 0.4531 | Yes |
| 5 | ADM | ADM Entrez, Source | adrenomedullin | 96 | 0.523 | 0.5305 | Yes |
| 6 | PRNP | PRNP Entrez, Source | prion protein (p27-30) (Creutzfeldt-Jakob disease, Gerstmann-Strausler-Scheinker syndrome, fatal familial insomnia) | 141 | 0.455 | 0.5947 | Yes |
| 7 | CDH13 | CDH13 Entrez, Source | cadherin 13, H-cadherin (heart) | 223 | 0.366 | 0.6430 | Yes |
| 8 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 297 | 0.311 | 0.6836 | Yes |
| 9 | PTTG1IP | PTTG1IP Entrez, Source | pituitary tumor-transforming 1 interacting protein | 811 | 0.168 | 0.6700 | Yes |
| 10 | LOXL1 | LOXL1 Entrez, Source | lysyl oxidase-like 1 | 1035 | 0.143 | 0.6745 | Yes |
| 11 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 1114 | 0.135 | 0.6886 | Yes |
| 12 | SLC16A3 | SLC16A3 Entrez, Source | solute carrier family 16, member 3 (monocarboxylic acid transporter 4) | 1690 | 0.098 | 0.6599 | No |
| 13 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 2074 | 0.082 | 0.6434 | No |
| 14 | BCAM | BCAM Entrez, Source | basal cell adhesion molecule (Lutheran blood group) | 2316 | 0.073 | 0.6361 | No |
| 15 | CBLB | CBLB Entrez, Source | Cas-Br-M (murine) ecotropic retroviral transforming sequence b | 2684 | 0.063 | 0.6179 | No |
| 16 | GPI | GPI Entrez, Source | glucose phosphate isomerase | 3287 | 0.048 | 0.5798 | No |
| 17 | RND3 | RND3 Entrez, Source | Rho family GTPase 3 | 3937 | 0.034 | 0.5360 | No |
| 18 | NOTCH3 | NOTCH3 Entrez, Source | Notch homolog 3 (Drosophila) | 4689 | 0.020 | 0.4826 | No |
| 19 | NID1 | NID1 Entrez, Source | nidogen 1 | 5553 | 0.007 | 0.4188 | No |
| 20 | PKN1 | PKN1 Entrez, Source | protein kinase N1 | 6325 | -0.005 | 0.3616 | No |
| 21 | CNTN2 | CNTN2 Entrez, Source | contactin 2 (axonal) | 6852 | -0.013 | 0.3240 | No |
| 22 | SIRPA | SIRPA Entrez, Source | signal-regulatory protein alpha | 7460 | -0.022 | 0.2817 | No |
| 23 | GABRA5 | GABRA5 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 5 | 8219 | -0.034 | 0.2298 | No |
| 24 | CSNK2A1 | CSNK2A1 Entrez, Source | casein kinase 2, alpha 1 polypeptide | 8931 | -0.046 | 0.1832 | No |
| 25 | CALR | CALR Entrez, Source | calreticulin | 9052 | -0.048 | 0.1814 | No |
| 26 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 9141 | -0.050 | 0.1821 | No |
| 27 | PLXNB2 | PLXNB2 Entrez, Source | plexin B2 | 9951 | -0.066 | 0.1311 | No |
| 28 | HSPA5 | HSPA5 Entrez, Source | heat shock 70kDa protein 5 (glucose-regulated protein, 78kDa) | 10956 | -0.089 | 0.0689 | No |
| 29 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 10960 | -0.089 | 0.0819 | No |
| 30 | SLC7A5 | SLC7A5 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 5 | 13201 | -0.300 | -0.0418 | No |
| 31 | SLC3A2 | SLC3A2 Entrez, Source | solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2 | 13255 | -0.353 | 0.0065 | No |