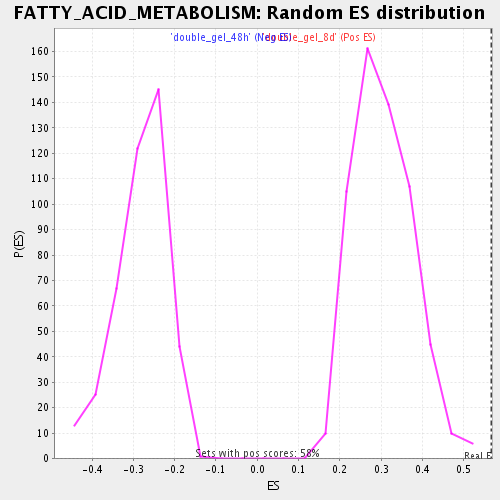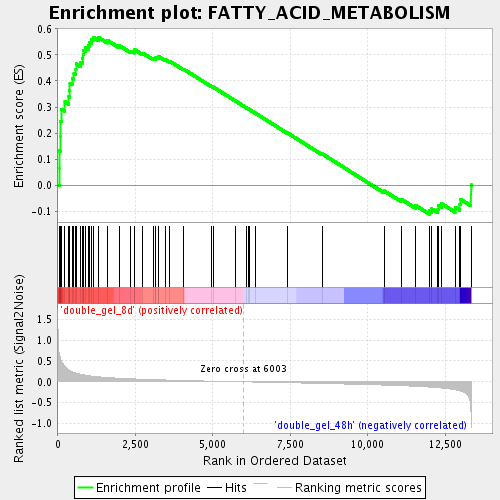
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | FATTY_ACID_METABOLISM |
| Enrichment Score (ES) | 0.5679345 |
| Normalized Enrichment Score (NES) | 1.8599346 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.018130446 |
| FWER p-Value | 0.806 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LPL | LPL Entrez, Source | lipoprotein lipase | 45 | 0.671 | 0.0649 | Yes |
| 2 | CPT1A | CPT1A Entrez, Source | carnitine palmitoyltransferase 1A (liver) | 47 | 0.660 | 0.1319 | Yes |
| 3 | CYP1A1 | CYP1A1 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 1 | 72 | 0.571 | 0.1882 | Yes |
| 4 | ADHFE1 | ADHFE1 Entrez, Source | alcohol dehydrogenase, iron containing, 1 | 74 | 0.567 | 0.2459 | Yes |
| 5 | FABP2 | FABP2 Entrez, Source | fatty acid binding protein 2, intestinal | 121 | 0.476 | 0.2909 | Yes |
| 6 | DCI | DCI Entrez, Source | dodecenoyl-Coenzyme A delta isomerase (3,2 trans-enoyl-Coenzyme A isomerase) | 226 | 0.365 | 0.3202 | Yes |
| 7 | CPT2 | CPT2 Entrez, Source | carnitine palmitoyltransferase II | 351 | 0.281 | 0.3394 | Yes |
| 8 | ACOX2 | ACOX2 Entrez, Source | acyl-Coenzyme A oxidase 2, branched chain | 379 | 0.269 | 0.3647 | Yes |
| 9 | CYP2E1 | CYP2E1 Entrez, Source | cytochrome P450, family 2, subfamily E, polypeptide 1 | 389 | 0.265 | 0.3910 | Yes |
| 10 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 470 | 0.238 | 0.4091 | Yes |
| 11 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 517 | 0.223 | 0.4284 | Yes |
| 12 | FABP1 | FABP1 Entrez, Source | fatty acid binding protein 1, liver | 580 | 0.210 | 0.4451 | Yes |
| 13 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 585 | 0.209 | 0.4660 | Yes |
| 14 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 739 | 0.178 | 0.4726 | Yes |
| 15 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 798 | 0.170 | 0.4855 | Yes |
| 16 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 809 | 0.168 | 0.5019 | Yes |
| 17 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 826 | 0.166 | 0.5175 | Yes |
| 18 | PECI | PECI Entrez, Source | peroxisomal D3,D2-enoyl-CoA isomerase | 894 | 0.157 | 0.5285 | Yes |
| 19 | ACADL | ACADL Entrez, Source | acyl-Coenzyme A dehydrogenase, long chain | 976 | 0.149 | 0.5375 | Yes |
| 20 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 1025 | 0.145 | 0.5486 | Yes |
| 21 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 1086 | 0.137 | 0.5581 | Yes |
| 22 | SDS | SDS Entrez, Source | serine dehydratase | 1136 | 0.133 | 0.5679 | Yes |
| 23 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 1310 | 0.119 | 0.5670 | No |
| 24 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 1594 | 0.103 | 0.5562 | No |
| 25 | CYP2C9 | CYP2C9 Entrez, Source | cytochrome P450, family 2, subfamily C, polypeptide 9 | 1984 | 0.085 | 0.5356 | No |
| 26 | ACSL1 | ACSL1 Entrez, Source | acyl-CoA synthetase long-chain family member 1 | 2355 | 0.072 | 0.5151 | No |
| 27 | CYP19A1 | CYP19A1 Entrez, Source | cytochrome P450, family 19, subfamily A, polypeptide 1 | 2460 | 0.069 | 0.5143 | No |
| 28 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 2484 | 0.069 | 0.5196 | No |
| 29 | ACOX1 | ACOX1 Entrez, Source | acyl-Coenzyme A oxidase 1, palmitoyl | 2738 | 0.062 | 0.5068 | No |
| 30 | ADH1A | ADH1A Entrez, Source | alcohol dehydrogenase 1A (class I), alpha polypeptide | 3072 | 0.052 | 0.4871 | No |
| 31 | CYP3A4 | CYP3A4 Entrez, Source | cytochrome P450, family 3, subfamily A, polypeptide 4 | 3149 | 0.051 | 0.4865 | No |
| 32 | ABCD2 | ABCD2 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 2 | 3159 | 0.051 | 0.4910 | No |
| 33 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 3245 | 0.048 | 0.4895 | No |
| 34 | ADH7 | ADH7 Entrez, Source | alcohol dehydrogenase 7 (class IV), mu or sigma polypeptide | 3254 | 0.048 | 0.4938 | No |
| 35 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 3480 | 0.043 | 0.4813 | No |
| 36 | LIPE | LIPE Entrez, Source | lipase, hormone-sensitive | 3607 | 0.041 | 0.4759 | No |
| 37 | PECR | PECR Entrez, Source | peroxisomal trans-2-enoyl-CoA reductase | 4050 | 0.032 | 0.4459 | No |
| 38 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4941 | 0.016 | 0.3805 | No |
| 39 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 5009 | 0.015 | 0.3770 | No |
| 40 | ADH6 | ADH6 Entrez, Source | alcohol dehydrogenase 6 (class V) | 5740 | 0.004 | 0.3224 | No |
| 41 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 6090 | -0.001 | 0.2963 | No |
| 42 | ACSL5 | ACSL5 Entrez, Source | acyl-CoA synthetase long-chain family member 5 | 6137 | -0.002 | 0.2931 | No |
| 43 | ADH4 | ADH4 Entrez, Source | alcohol dehydrogenase 4 (class II), pi polypeptide | 6171 | -0.003 | 0.2908 | No |
| 44 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 6362 | -0.006 | 0.2771 | No |
| 45 | ALDH1A2 | ALDH1A2 Entrez, Source | aldehyde dehydrogenase 1 family, member A2 | 7402 | -0.021 | 0.2010 | No |
| 46 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 8536 | -0.039 | 0.1198 | No |
| 47 | ACADSB | ACADSB Entrez, Source | acyl-Coenzyme A dehydrogenase, short/branched chain | 10528 | -0.078 | -0.0221 | No |
| 48 | ACOX3 | ACOX3 Entrez, Source | acyl-Coenzyme A oxidase 3, pristanoyl | 11075 | -0.092 | -0.0538 | No |
| 49 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 11541 | -0.107 | -0.0779 | No |
| 50 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 11986 | -0.126 | -0.0986 | No |
| 51 | FASN | FASN Entrez, Source | fatty acid synthase | 12064 | -0.130 | -0.0912 | No |
| 52 | CYP51A1 | CYP51A1 Entrez, Source | cytochrome P450, family 51, subfamily A, polypeptide 1 | 12263 | -0.140 | -0.0918 | No |
| 53 | PPARA | PPARA Entrez, Source | peroxisome proliferative activated receptor, alpha | 12287 | -0.141 | -0.0791 | No |
| 54 | CYP1A2 | CYP1A2 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 2 | 12372 | -0.147 | -0.0705 | No |
| 55 | CYP4B1 | CYP4B1 Entrez, Source | cytochrome P450, family 4, subfamily B, polypeptide 1 | 12818 | -0.188 | -0.0849 | No |
| 56 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 12971 | -0.213 | -0.0747 | No |
| 57 | GNPAT | GNPAT Entrez, Source | glyceronephosphate O-acyltransferase | 12982 | -0.215 | -0.0536 | No |
| 58 | FABP7 | FABP7 Entrez, Source | fatty acid binding protein 7, brain | 13332 | -0.793 | 0.0008 | No |