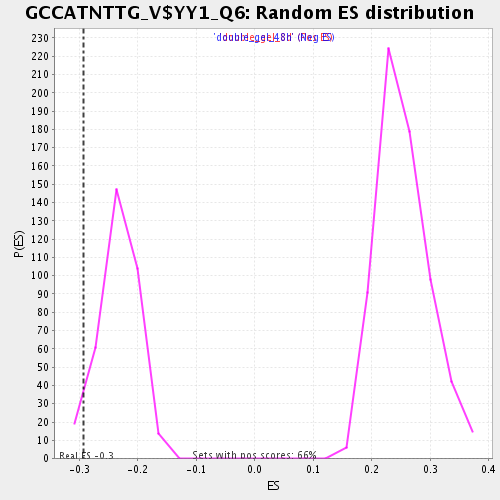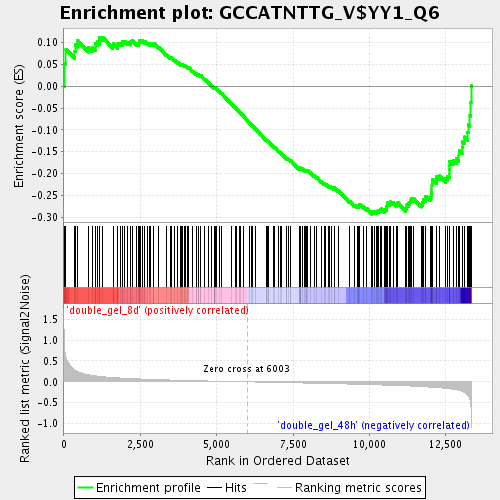
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | GCCATNTTG_V$YY1_Q6 |
| Enrichment Score (ES) | -0.2929433 |
| Normalized Enrichment Score (NES) | -1.2538767 |
| Nominal p-value | 0.052173913 |
| FDR q-value | 0.52804744 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 10 | 0.885 | 0.0532 | No |
| 2 | RAPGEF4 | RAPGEF4 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 4 | 66 | 0.600 | 0.0855 | No |
| 3 | NCDN | NCDN Entrez, Source | neurochondrin | 358 | 0.277 | 0.0803 | No |
| 4 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 375 | 0.270 | 0.0956 | No |
| 5 | WSB1 | WSB1 Entrez, Source | WD repeat and SOCS box-containing 1 | 450 | 0.245 | 0.1049 | No |
| 6 | POU2F1 | POU2F1 Entrez, Source | POU domain, class 2, transcription factor 1 | 797 | 0.170 | 0.0889 | No |
| 7 | GNAI2 | GNAI2 Entrez, Source | guanine nucleotide binding protein (G protein), alpha inhibiting activity polypeptide 2 | 931 | 0.153 | 0.0881 | No |
| 8 | SCRN1 | SCRN1 Entrez, Source | secernin 1 | 1023 | 0.145 | 0.0901 | No |
| 9 | LPHN1 | LPHN1 Entrez, Source | latrophilin 1 | 1024 | 0.145 | 0.0989 | No |
| 10 | TGIF | TGIF Entrez, Source | TGFB-induced factor (TALE family homeobox) | 1094 | 0.137 | 0.1020 | No |
| 11 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 1166 | 0.131 | 0.1046 | No |
| 12 | ABHD1 | ABHD1 Entrez, Source | abhydrolase domain containing 1 | 1175 | 0.130 | 0.1119 | No |
| 13 | SNX17 | SNX17 Entrez, Source | sorting nexin 17 | 1266 | 0.122 | 0.1125 | No |
| 14 | APBB1 | APBB1 Entrez, Source | amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) | 1606 | 0.102 | 0.0930 | No |
| 15 | PPP2CA | PPP2CA Entrez, Source | protein phosphatase 2 (formerly 2A), catalytic subunit, alpha isoform | 1627 | 0.101 | 0.0977 | No |
| 16 | CACNB2 | CACNB2 Entrez, Source | calcium channel, voltage-dependent, beta 2 subunit | 1764 | 0.095 | 0.0931 | No |
| 17 | DUSP7 | DUSP7 Entrez, Source | dual specificity phosphatase 7 | 1768 | 0.094 | 0.0986 | No |
| 18 | SFXN1 | SFXN1 Entrez, Source | sideroflexin 1 | 1842 | 0.091 | 0.0986 | No |
| 19 | IDH3G | IDH3G Entrez, Source | isocitrate dehydrogenase 3 (NAD+) gamma | 1907 | 0.088 | 0.0991 | No |
| 20 | FKRP | FKRP Entrez, Source | fukutin related protein | 1920 | 0.088 | 0.1036 | No |
| 21 | WASL | WASL Entrez, Source | Wiskott-Aldrich syndrome-like | 1998 | 0.085 | 0.1029 | No |
| 22 | DSCAM | DSCAM Entrez, Source | Down syndrome cell adhesion molecule | 2085 | 0.081 | 0.1013 | No |
| 23 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 2173 | 0.078 | 0.0995 | No |
| 24 | COX8A | COX8A Entrez, Source | cytochrome c oxidase subunit 8A (ubiquitous) | 2190 | 0.078 | 0.1030 | No |
| 25 | DZIP1 | DZIP1 Entrez, Source | DAZ interacting protein 1 | 2229 | 0.076 | 0.1048 | No |
| 26 | UQCRH | UQCRH Entrez, Source | ubiquinol-cytochrome c reductase hinge protein | 2381 | 0.071 | 0.0976 | No |
| 27 | PITX1 | PITX1 Entrez, Source | paired-like homeodomain transcription factor 1 | 2446 | 0.069 | 0.0970 | No |
| 28 | ING3 | ING3 Entrez, Source | inhibitor of growth family, member 3 | 2458 | 0.069 | 0.1004 | No |
| 29 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 2459 | 0.069 | 0.1046 | No |
| 30 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 2499 | 0.068 | 0.1058 | No |
| 31 | SLC6A1 | SLC6A1 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, GABA), member 1 | 2567 | 0.066 | 0.1047 | No |
| 32 | HSF1 | HSF1 Entrez, Source | heat shock transcription factor 1 | 2641 | 0.064 | 0.1030 | No |
| 33 | COX7B | COX7B Entrez, Source | cytochrome c oxidase subunit VIIb | 2748 | 0.061 | 0.0987 | No |
| 34 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 2813 | 0.060 | 0.0975 | No |
| 35 | MPDU1 | MPDU1 Entrez, Source | mannose-P-dolichol utilization defect 1 | 2839 | 0.059 | 0.0992 | No |
| 36 | CD4 | CD4 Entrez, Source | CD4 molecule | 2921 | 0.057 | 0.0965 | No |
| 37 | AIP | AIP Entrez, Source | aryl hydrocarbon receptor interacting protein | 2931 | 0.057 | 0.0993 | No |
| 38 | NUDT3 | NUDT3 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 3 | 3099 | 0.052 | 0.0897 | No |
| 39 | MLL5 | MLL5 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia 5 (trithorax homolog, Drosophila) | 3353 | 0.046 | 0.0733 | No |
| 40 | UBAP1 | UBAP1 Entrez, Source | ubiquitin associated protein 1 | 3485 | 0.043 | 0.0660 | No |
| 41 | NCOA6 | NCOA6 Entrez, Source | nuclear receptor coactivator 6 | 3527 | 0.042 | 0.0655 | No |
| 42 | TCF4 | TCF4 Entrez, Source | transcription factor 4 | 3628 | 0.040 | 0.0603 | No |
| 43 | HOXA2 | HOXA2 Entrez, Source | homeobox A2 | 3729 | 0.038 | 0.0550 | No |
| 44 | THRAP4 | THRAP4 Entrez, Source | thyroid hormone receptor associated protein 4 | 3822 | 0.036 | 0.0503 | No |
| 45 | ATF4 | ATF4 Entrez, Source | activating transcription factor 4 (tax-responsive enhancer element B67) | 3861 | 0.035 | 0.0495 | No |
| 46 | ZFP37 | ZFP37 Entrez, Source | zinc finger protein 37 homolog (mouse) | 3891 | 0.035 | 0.0494 | No |
| 47 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 3945 | 0.034 | 0.0475 | No |
| 48 | DGCR2 | DGCR2 Entrez, Source | DiGeorge syndrome critical region gene 2 | 3982 | 0.033 | 0.0468 | No |
| 49 | TIA1 | TIA1 Entrez, Source | TIA1 cytotoxic granule-associated RNA binding protein | 4046 | 0.032 | 0.0439 | No |
| 50 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 4083 | 0.031 | 0.0431 | No |
| 51 | RAB5A | RAB5A Entrez, Source | RAB5A, member RAS oncogene family | 4218 | 0.029 | 0.0347 | No |
| 52 | RBMS1 | RBMS1 Entrez, Source | RNA binding motif, single stranded interacting protein 1 | 4322 | 0.027 | 0.0285 | No |
| 53 | SHANK2 | SHANK2 Entrez, Source | SH3 and multiple ankyrin repeat domains 2 | 4334 | 0.027 | 0.0293 | No |
| 54 | MAEA | MAEA Entrez, Source | macrophage erythroblast attacher | 4393 | 0.026 | 0.0264 | No |
| 55 | BTRC | BTRC Entrez, Source | beta-transducin repeat containing | 4396 | 0.025 | 0.0278 | No |
| 56 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 4477 | 0.024 | 0.0232 | No |
| 57 | RAB4B | RAB4B Entrez, Source | RAB4B, member RAS oncogene family | 4479 | 0.024 | 0.0246 | No |
| 58 | RAB1A | RAB1A Entrez, Source | RAB1A, member RAS oncogene family | 4611 | 0.021 | 0.0159 | No |
| 59 | SFRS5 | SFRS5 Entrez, Source | splicing factor, arginine/serine-rich 5 | 4725 | 0.020 | 0.0085 | No |
| 60 | NFYA | NFYA Entrez, Source | nuclear transcription factor Y, alpha | 4833 | 0.018 | 0.0015 | No |
| 61 | RUFY1 | RUFY1 Entrez, Source | RUN and FYVE domain containing 1 | 4914 | 0.017 | -0.0036 | No |
| 62 | NR2C2 | NR2C2 Entrez, Source | nuclear receptor subfamily 2, group C, member 2 | 4923 | 0.016 | -0.0032 | No |
| 63 | PIAS3 | PIAS3 Entrez, Source | protein inhibitor of activated STAT, 3 | 4943 | 0.016 | -0.0037 | No |
| 64 | UBC | UBC Entrez, Source | ubiquitin C | 4954 | 0.016 | -0.0035 | No |
| 65 | USF1 | USF1 Entrez, Source | upstream transcription factor 1 | 4980 | 0.015 | -0.0044 | No |
| 66 | MARK3 | MARK3 Entrez, Source | MAP/microtubule affinity-regulating kinase 3 | 5075 | 0.014 | -0.0107 | No |
| 67 | ARF1 | ARF1 Entrez, Source | ADP-ribosylation factor 1 | 5149 | 0.012 | -0.0155 | No |
| 68 | PFN2 | PFN2 Entrez, Source | profilin 2 | 5491 | 0.007 | -0.0410 | No |
| 69 | ADK | ADK Entrez, Source | adenosine kinase | 5620 | 0.005 | -0.0504 | No |
| 70 | ARCN1 | ARCN1 Entrez, Source | archain 1 | 5640 | 0.005 | -0.0515 | No |
| 71 | ENSA | ENSA Entrez, Source | endosulfine alpha | 5749 | 0.003 | -0.0595 | No |
| 72 | MTMR3 | MTMR3 Entrez, Source | myotubularin related protein 3 | 5782 | 0.003 | -0.0618 | No |
| 73 | MAP2K5 | MAP2K5 Entrez, Source | mitogen-activated protein kinase kinase 5 | 5887 | 0.002 | -0.0695 | No |
| 74 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 6060 | -0.001 | -0.0826 | No |
| 75 | NAPE-PLD | NAPE-PLD Entrez, Source | - | 6144 | -0.002 | -0.0887 | No |
| 76 | CD164 | CD164 Entrez, Source | CD164 molecule, sialomucin | 6173 | -0.003 | -0.0907 | No |
| 77 | LYPLA2 | LYPLA2 Entrez, Source | lysophospholipase II | 6266 | -0.004 | -0.0974 | No |
| 78 | HSPA8 | HSPA8 Entrez, Source | heat shock 70kDa protein 8 | 6613 | -0.009 | -0.1232 | No |
| 79 | SP1 | SP1 Entrez, Source | Sp1 transcription factor | 6622 | -0.009 | -0.1232 | No |
| 80 | L3MBTL2 | L3MBTL2 Entrez, Source | l(3)mbt-like 2 (Drosophila) | 6673 | -0.010 | -0.1264 | No |
| 81 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 6690 | -0.011 | -0.1270 | No |
| 82 | RAB10 | RAB10 Entrez, Source | RAB10, member RAS oncogene family | 6696 | -0.011 | -0.1267 | No |
| 83 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 6862 | -0.013 | -0.1384 | No |
| 84 | U2AF2 | U2AF2 Entrez, Source | U2 small nuclear RNA auxiliary factor 2 | 6904 | -0.014 | -0.1407 | No |
| 85 | RAB28 | RAB28 Entrez, Source | RAB28, member RAS oncogene family | 7029 | -0.016 | -0.1492 | No |
| 86 | TTC17 | TTC17 Entrez, Source | tetratricopeptide repeat domain 17 | 7081 | -0.016 | -0.1521 | No |
| 87 | ATP5F1 | ATP5F1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit B1 | 7135 | -0.017 | -0.1550 | No |
| 88 | CUGBP1 | CUGBP1 Entrez, Source | CUG triplet repeat, RNA binding protein 1 | 7294 | -0.019 | -0.1659 | No |
| 89 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 7295 | -0.019 | -0.1647 | No |
| 90 | CLK4 | CLK4 Entrez, Source | CDC-like kinase 4 | 7341 | -0.020 | -0.1669 | No |
| 91 | NFYC | NFYC Entrez, Source | nuclear transcription factor Y, gamma | 7410 | -0.021 | -0.1708 | No |
| 92 | USP14 | USP14 Entrez, Source | ubiquitin specific peptidase 14 (tRNA-guanine transglycosylase) | 7427 | -0.021 | -0.1707 | No |
| 93 | PPP1CC | PPP1CC Entrez, Source | protein phosphatase 1, catalytic subunit, gamma isoform | 7701 | -0.026 | -0.1898 | No |
| 94 | GIT2 | GIT2 Entrez, Source | G protein-coupled receptor kinase interactor 2 | 7717 | -0.026 | -0.1894 | No |
| 95 | TAF6 | TAF6 Entrez, Source | TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa | 7744 | -0.027 | -0.1897 | No |
| 96 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 7751 | -0.027 | -0.1886 | No |
| 97 | SYT11 | SYT11 Entrez, Source | synaptotagmin XI | 7754 | -0.027 | -0.1871 | No |
| 98 | EIF2B4 | EIF2B4 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 4 delta, 67kDa | 7792 | -0.027 | -0.1883 | No |
| 99 | ATP5A1 | ATP5A1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F1 complex, alpha subunit 1, cardiac muscle | 7872 | -0.028 | -0.1925 | No |
| 100 | GNAO1 | GNAO1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O | 7919 | -0.029 | -0.1942 | No |
| 101 | RXRB | RXRB Entrez, Source | retinoid X receptor, beta | 7930 | -0.030 | -0.1932 | No |
| 102 | PSMC5 | PSMC5 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 5 | 7944 | -0.030 | -0.1924 | No |
| 103 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 7982 | -0.030 | -0.1933 | No |
| 104 | NSF | NSF Entrez, Source | N-ethylmaleimide-sensitive factor | 8063 | -0.032 | -0.1974 | No |
| 105 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 8193 | -0.034 | -0.2052 | No |
| 106 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 8274 | -0.035 | -0.2091 | No |
| 107 | ATP5G1 | ATP5G1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C1 (subunit 9) | 8437 | -0.038 | -0.2191 | No |
| 108 | PAPOLB | PAPOLB Entrez, Source | poly(A) polymerase beta (testis specific) | 8516 | -0.039 | -0.2227 | No |
| 109 | AP3D1 | AP3D1 Entrez, Source | adaptor-related protein complex 3, delta 1 subunit | 8568 | -0.040 | -0.2241 | No |
| 110 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 8660 | -0.041 | -0.2285 | No |
| 111 | AKAP8L | AKAP8L Entrez, Source | A kinase (PRKA) anchor protein 8-like | 8707 | -0.042 | -0.2294 | No |
| 112 | RAGE | RAGE Entrez, Source | renal tumor antigen | 8769 | -0.043 | -0.2314 | No |
| 113 | YWHAE | YWHAE Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon polypeptide | 8841 | -0.044 | -0.2341 | No |
| 114 | ANAPC4 | ANAPC4 Entrez, Source | anaphase promoting complex subunit 4 | 8855 | -0.045 | -0.2324 | No |
| 115 | RBM5 | RBM5 Entrez, Source | RNA binding motif protein 5 | 8977 | -0.047 | -0.2387 | No |
| 116 | RPL10 | RPL10 Entrez, Source | ribosomal protein L10 | 9343 | -0.053 | -0.2632 | No |
| 117 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 9515 | -0.056 | -0.2728 | No |
| 118 | SSR4 | SSR4 Entrez, Source | signal sequence receptor, delta (translocon-associated protein delta) | 9596 | -0.058 | -0.2753 | No |
| 119 | MYST2 | MYST2 Entrez, Source | MYST histone acetyltransferase 2 | 9642 | -0.059 | -0.2752 | No |
| 120 | EIF5 | EIF5 Entrez, Source | eukaryotic translation initiation factor 5 | 9653 | -0.059 | -0.2723 | No |
| 121 | CAB39L | CAB39L Entrez, Source | calcium binding protein 39-like | 9676 | -0.059 | -0.2704 | No |
| 122 | EIF4G2 | EIF4G2 Entrez, Source | eukaryotic translation initiation factor 4 gamma, 2 | 9796 | -0.062 | -0.2757 | No |
| 123 | DDX3X | DDX3X Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked | 9904 | -0.064 | -0.2799 | No |
| 124 | KCNMA1 | KCNMA1 Entrez, Source | potassium large conductance calcium-activated channel, subfamily M, alpha member 1 | 10077 | -0.068 | -0.2888 | Yes |
| 125 | PTBP1 | PTBP1 Entrez, Source | polypyrimidine tract binding protein 1 | 10104 | -0.068 | -0.2866 | Yes |
| 126 | RNF12 | RNF12 Entrez, Source | ring finger protein 12 | 10162 | -0.069 | -0.2867 | Yes |
| 127 | PTDSR | PTDSR Entrez, Source | phosphatidylserine receptor | 10242 | -0.071 | -0.2884 | Yes |
| 128 | MRPL11 | MRPL11 Entrez, Source | mitochondrial ribosomal protein L11 | 10267 | -0.072 | -0.2858 | Yes |
| 129 | RBM16 | RBM16 Entrez, Source | RNA binding motif protein 16 | 10310 | -0.073 | -0.2846 | Yes |
| 130 | CELSR3 | CELSR3 Entrez, Source | cadherin, EGF LAG seven-pass G-type receptor 3 (flamingo homolog, Drosophila) | 10349 | -0.074 | -0.2830 | Yes |
| 131 | TBC1D14 | TBC1D14 Entrez, Source | TBC1 domain family, member 14 | 10383 | -0.074 | -0.2809 | Yes |
| 132 | ATF1 | ATF1 Entrez, Source | activating transcription factor 1 | 10486 | -0.077 | -0.2840 | Yes |
| 133 | SUMO2 | SUMO2 Entrez, Source | SMT3 suppressor of mif two 3 homolog 2 (S. cerevisiae) | 10516 | -0.078 | -0.2814 | Yes |
| 134 | DAP3 | DAP3 Entrez, Source | death associated protein 3 | 10562 | -0.079 | -0.2800 | Yes |
| 135 | BTF3 | BTF3 Entrez, Source | basic transcription factor 3 | 10564 | -0.079 | -0.2753 | Yes |
| 136 | AKAP8 | AKAP8 Entrez, Source | A kinase (PRKA) anchor protein 8 | 10573 | -0.079 | -0.2710 | Yes |
| 137 | MAP4K3 | MAP4K3 Entrez, Source | mitogen-activated protein kinase kinase kinase kinase 3 | 10588 | -0.080 | -0.2673 | Yes |
| 138 | AP1G1 | AP1G1 Entrez, Source | adaptor-related protein complex 1, gamma 1 subunit | 10665 | -0.082 | -0.2681 | Yes |
| 139 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 10691 | -0.082 | -0.2650 | Yes |
| 140 | RPS8 | RPS8 Entrez, Source | ribosomal protein S8 | 10780 | -0.085 | -0.2665 | Yes |
| 141 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 10891 | -0.088 | -0.2695 | Yes |
| 142 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 10930 | -0.089 | -0.2670 | Yes |
| 143 | ING4 | ING4 Entrez, Source | inhibitor of growth family, member 4 | 11185 | -0.095 | -0.2804 | Yes |
| 144 | UBA52 | UBA52 Entrez, Source | ubiquitin A-52 residue ribosomal protein fusion product 1 | 11223 | -0.096 | -0.2774 | Yes |
| 145 | ATP5G2 | ATP5G2 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit C2 (subunit 9) | 11226 | -0.096 | -0.2717 | Yes |
| 146 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 11272 | -0.098 | -0.2691 | Yes |
| 147 | RPN1 | RPN1 Entrez, Source | ribophorin I | 11310 | -0.099 | -0.2659 | Yes |
| 148 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 11330 | -0.100 | -0.2613 | Yes |
| 149 | WHSC2 | WHSC2 Entrez, Source | Wolf-Hirschhorn syndrome candidate 2 | 11360 | -0.101 | -0.2573 | Yes |
| 150 | METTL3 | METTL3 Entrez, Source | methyltransferase like 3 | 11454 | -0.104 | -0.2580 | Yes |
| 151 | LRRC4 | LRRC4 Entrez, Source | leucine rich repeat containing 4 | 11701 | -0.113 | -0.2698 | Yes |
| 152 | HNRPC | HNRPC Entrez, Source | heterogeneous nuclear ribonucleoprotein C (C1/C2) | 11740 | -0.115 | -0.2658 | Yes |
| 153 | BOP1 | BOP1 Entrez, Source | block of proliferation 1 | 11761 | -0.115 | -0.2602 | Yes |
| 154 | CASC3 | CASC3 Entrez, Source | cancer susceptibility candidate 3 | 11825 | -0.118 | -0.2578 | Yes |
| 155 | EIF4A1 | EIF4A1 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 1 | 11843 | -0.119 | -0.2519 | Yes |
| 156 | SAFB | SAFB Entrez, Source | scaffold attachment factor B | 11983 | -0.126 | -0.2548 | Yes |
| 157 | RBM10 | RBM10 Entrez, Source | RNA binding motif protein 10 | 12015 | -0.127 | -0.2494 | Yes |
| 158 | FTSJ3 | FTSJ3 Entrez, Source | FtsJ homolog 3 (E. coli) | 12034 | -0.128 | -0.2430 | Yes |
| 159 | RALA | RALA Entrez, Source | v-ral simian leukemia viral oncogene homolog A (ras related) | 12043 | -0.128 | -0.2357 | Yes |
| 160 | RPL35A | RPL35A Entrez, Source | ribosomal protein L35a | 12045 | -0.129 | -0.2280 | Yes |
| 161 | GFER | GFER Entrez, Source | growth factor, augmenter of liver regeneration (ERV1 homolog, S. cerevisiae) | 12048 | -0.129 | -0.2203 | Yes |
| 162 | CSNK1A1 | CSNK1A1 Entrez, Source | casein kinase 1, alpha 1 | 12054 | -0.129 | -0.2128 | Yes |
| 163 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 12183 | -0.136 | -0.2142 | Yes |
| 164 | UGCGL1 | UGCGL1 Entrez, Source | UDP-glucose ceramide glucosyltransferase-like 1 | 12203 | -0.137 | -0.2073 | Yes |
| 165 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 12292 | -0.142 | -0.2054 | Yes |
| 166 | PTMA | PTMA Entrez, Source | prothymosin, alpha (gene sequence 28) | 12489 | -0.154 | -0.2109 | Yes |
| 167 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 12549 | -0.158 | -0.2057 | Yes |
| 168 | ADNP | ADNP Entrez, Source | activity-dependent neuroprotector | 12610 | -0.164 | -0.2003 | Yes |
| 169 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 12612 | -0.164 | -0.1904 | Yes |
| 170 | RBM12 | RBM12 Entrez, Source | RNA binding motif protein 12 | 12622 | -0.165 | -0.1810 | Yes |
| 171 | CRKL | CRKL Entrez, Source | v-crk sarcoma virus CT10 oncogene homolog (avian)-like | 12628 | -0.166 | -0.1713 | Yes |
| 172 | EXOC7 | EXOC7 Entrez, Source | exocyst complex component 7 | 12748 | -0.178 | -0.1695 | Yes |
| 173 | H1F0 | H1F0 Entrez, Source | H1 histone family, member 0 | 12841 | -0.191 | -0.1649 | Yes |
| 174 | PRPSAP2 | PRPSAP2 Entrez, Source | phosphoribosyl pyrophosphate synthetase-associated protein 2 | 12928 | -0.204 | -0.1590 | Yes |
| 175 | RBM3 | RBM3 Entrez, Source | RNA binding motif (RNP1, RRM) protein 3 | 12930 | -0.204 | -0.1466 | Yes |
| 176 | NASP | NASP Entrez, Source | nuclear autoantigenic sperm protein (histone-binding) | 13031 | -0.226 | -0.1404 | Yes |
| 177 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 13043 | -0.231 | -0.1272 | Yes |
| 178 | FUS | FUS Entrez, Source | fusion (involved in t(12;16) in malignant liposarcoma) | 13100 | -0.252 | -0.1161 | Yes |
| 179 | SFPQ | SFPQ Entrez, Source | splicing factor proline/glutamine-rich (polypyrimidine tract binding protein associated) | 13213 | -0.310 | -0.1057 | Yes |
| 180 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 13231 | -0.325 | -0.0871 | Yes |
| 181 | HNRPA1 | HNRPA1 Entrez, Source | heterogeneous nuclear ribonucleoprotein A1 | 13288 | -0.411 | -0.0663 | Yes |
| 182 | PSPC1 | PSPC1 Entrez, Source | paraspeckle component 1 | 13317 | -0.524 | -0.0365 | Yes |
| 183 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 13326 | -0.630 | 0.0012 | Yes |