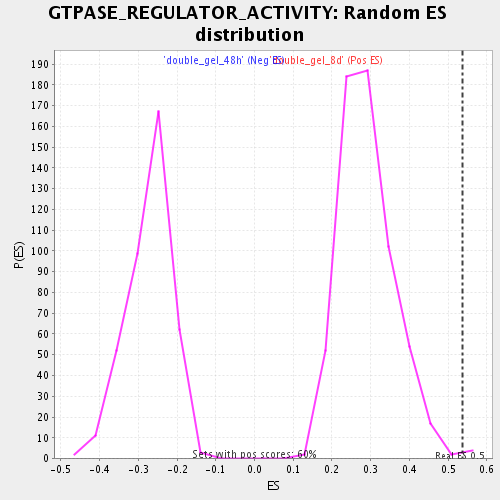
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | GTPASE_REGULATOR_ACTIVITY |
| Enrichment Score (ES) | 0.5371773 |
| Normalized Enrichment Score (NES) | 1.8520932 |
| Nominal p-value | 0.0066225166 |
| FDR q-value | 0.018254172 |
| FWER p-Value | 0.838 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 25 | 0.771 | 0.0953 | Yes |
| 2 | RAPGEF4 | RAPGEF4 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 4 | 66 | 0.600 | 0.1679 | Yes |
| 3 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 75 | 0.565 | 0.2384 | Yes |
| 4 | RGS16 | RGS16 Entrez, Source | regulator of G-protein signalling 16 | 103 | 0.505 | 0.3001 | Yes |
| 5 | PLCE1 | PLCE1 Entrez, Source | phospholipase C, epsilon 1 | 203 | 0.384 | 0.3409 | Yes |
| 6 | RGS3 | RGS3 Entrez, Source | regulator of G-protein signalling 3 | 338 | 0.287 | 0.3670 | Yes |
| 7 | RAPGEF5 | RAPGEF5 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 5 | 384 | 0.267 | 0.3972 | Yes |
| 8 | RGS4 | RGS4 Entrez, Source | regulator of G-protein signalling 4 | 454 | 0.244 | 0.4227 | Yes |
| 9 | RGS5 | RGS5 Entrez, Source | regulator of G-protein signalling 5 | 513 | 0.224 | 0.4465 | Yes |
| 10 | CDC42EP2 | CDC42EP2 Entrez, Source | CDC42 effector protein (Rho GTPase binding) 2 | 522 | 0.222 | 0.4739 | Yes |
| 11 | TNK2 | TNK2 Entrez, Source | tyrosine kinase, non-receptor, 2 | 629 | 0.201 | 0.4913 | Yes |
| 12 | ALDH1A1 | ALDH1A1 Entrez, Source | aldehyde dehydrogenase 1 family, member A1 | 739 | 0.178 | 0.5055 | Yes |
| 13 | EIF2B2 | EIF2B2 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 2 beta, 39kDa | 975 | 0.149 | 0.5066 | Yes |
| 14 | PSCD1 | PSCD1 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 1(cytohesin 1) | 1237 | 0.125 | 0.5026 | Yes |
| 15 | RTKN | RTKN Entrez, Source | rhotekin | 1496 | 0.108 | 0.4968 | Yes |
| 16 | PSCD2 | PSCD2 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 2 (cytohesin-2) | 1498 | 0.108 | 0.5104 | Yes |
| 17 | CDC42SE1 | CDC42SE1 Entrez, Source | CDC42 small effector 1 | 1617 | 0.102 | 0.5143 | Yes |
| 18 | RALGDS | RALGDS Entrez, Source | ral guanine nucleotide dissociation stimulator | 1715 | 0.097 | 0.5192 | Yes |
| 19 | ARHGDIB | ARHGDIB Entrez, Source | Rho GDP dissociation inhibitor (GDI) beta | 1774 | 0.094 | 0.5267 | Yes |
| 20 | RALBP1 | RALBP1 Entrez, Source | ralA binding protein 1 | 1792 | 0.093 | 0.5372 | Yes |
| 21 | WASL | WASL Entrez, Source | Wiskott-Aldrich syndrome-like | 1998 | 0.085 | 0.5324 | No |
| 22 | ERRFI1 | ERRFI1 Entrez, Source | ERBB receptor feedback inhibitor 1 | 2488 | 0.068 | 0.5042 | No |
| 23 | PSCD3 | PSCD3 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 3 | 2590 | 0.065 | 0.5048 | No |
| 24 | RAPGEF3 | RAPGEF3 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 3 | 2950 | 0.056 | 0.4848 | No |
| 25 | CENTA1 | CENTA1 Entrez, Source | centaurin, alpha 1 | 3441 | 0.044 | 0.4535 | No |
| 26 | RAP1GAP | RAP1GAP Entrez, Source | RAP1 GTPase activating protein | 3624 | 0.040 | 0.4449 | No |
| 27 | ARHGEF1 | ARHGEF1 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 1 | 3821 | 0.036 | 0.4346 | No |
| 28 | EIF2B1 | EIF2B1 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 1 alpha, 26kDa | 4466 | 0.024 | 0.3892 | No |
| 29 | MYO9B | MYO9B Entrez, Source | myosin IXB | 4486 | 0.024 | 0.3907 | No |
| 30 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 5025 | 0.015 | 0.3521 | No |
| 31 | RASA3 | RASA3 Entrez, Source | RAS p21 protein activator 3 | 5066 | 0.014 | 0.3508 | No |
| 32 | RASGRP4 | RASGRP4 Entrez, Source | RAS guanyl releasing protein 4 | 5408 | 0.009 | 0.3262 | No |
| 33 | RIMS1 | RIMS1 Entrez, Source | regulating synaptic membrane exocytosis 1 | 5444 | 0.008 | 0.3246 | No |
| 34 | ARHGAP4 | ARHGAP4 Entrez, Source | Rho GTPase activating protein 4 | 5575 | 0.006 | 0.3156 | No |
| 35 | FGD1 | FGD1 Entrez, Source | FYVE, RhoGEF and PH domain containing 1 (faciogenital dysplasia) | 5692 | 0.004 | 0.3074 | No |
| 36 | CHN2 | CHN2 Entrez, Source | chimerin (chimaerin) 2 | 5842 | 0.002 | 0.2964 | No |
| 37 | RASA2 | RASA2 Entrez, Source | RAS p21 protein activator 2 | 6217 | -0.003 | 0.2687 | No |
| 38 | ARHGEF11 | ARHGEF11 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 11 | 6256 | -0.004 | 0.2663 | No |
| 39 | GPS1 | GPS1 Entrez, Source | G protein pathway suppressor 1 | 6668 | -0.010 | 0.2366 | No |
| 40 | RGS6 | RGS6 Entrez, Source | regulator of G-protein signalling 6 | 6832 | -0.013 | 0.2259 | No |
| 41 | RGS12 | RGS12 Entrez, Source | regulator of G-protein signalling 12 | 6916 | -0.014 | 0.2214 | No |
| 42 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 7149 | -0.017 | 0.2061 | No |
| 43 | KALRN | KALRN Entrez, Source | kalirin, RhoGEF kinase | 7276 | -0.019 | 0.1991 | No |
| 44 | RGS9 | RGS9 Entrez, Source | regulator of G-protein signalling 9 | 7456 | -0.022 | 0.1884 | No |
| 45 | CENTD2 | CENTD2 Entrez, Source | centaurin, delta 2 | 7725 | -0.026 | 0.1715 | No |
| 46 | GDI2 | GDI2 Entrez, Source | GDP dissociation inhibitor 2 | 7776 | -0.027 | 0.1711 | No |
| 47 | EIF2B4 | EIF2B4 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 4 delta, 67kDa | 7792 | -0.027 | 0.1734 | No |
| 48 | ARHGAP5 | ARHGAP5 Entrez, Source | Rho GTPase activating protein 5 | 8103 | -0.032 | 0.1541 | No |
| 49 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 8130 | -0.033 | 0.1563 | No |
| 50 | ALS2 | ALS2 Entrez, Source | amyotrophic lateral sclerosis 2 (juvenile) | 8397 | -0.037 | 0.1410 | No |
| 51 | ARHGAP27 | ARHGAP27 Entrez, Source | Rho GTPase activating protein 27 | 8729 | -0.043 | 0.1214 | No |
| 52 | RAB3GAP2 | RAB3GAP2 Entrez, Source | RAB3 GTPase activating protein subunit 2 (non-catalytic) | 8768 | -0.043 | 0.1240 | No |
| 53 | TSC2 | TSC2 Entrez, Source | tuberous sclerosis 2 | 8912 | -0.046 | 0.1190 | No |
| 54 | TIAM1 | TIAM1 Entrez, Source | T-cell lymphoma invasion and metastasis 1 | 9112 | -0.049 | 0.1102 | No |
| 55 | DLC1 | DLC1 Entrez, Source | deleted in liver cancer 1 | 9141 | -0.050 | 0.1143 | No |
| 56 | ARL2BP | ARL2BP Entrez, Source | ADP-ribosylation factor-like 2 binding protein | 9323 | -0.053 | 0.1074 | No |
| 57 | FGD4 | FGD4 Entrez, Source | FYVE, RhoGEF and PH domain containing 4 | 10187 | -0.070 | 0.0512 | No |
| 58 | ARHGDIA | ARHGDIA Entrez, Source | Rho GDP dissociation inhibitor (GDI) alpha | 10450 | -0.076 | 0.0411 | No |
| 59 | RANGAP1 | RANGAP1 Entrez, Source | Ran GTPase activating protein 1 | 10603 | -0.080 | 0.0397 | No |
| 60 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 10739 | -0.084 | 0.0401 | No |
| 61 | EIF2B3 | EIF2B3 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 3 gamma, 58kDa | 11098 | -0.093 | 0.0249 | No |
| 62 | RCBTB2 | RCBTB2 Entrez, Source | regulator of chromosome condensation (RCC1) and BTB (POZ) domain containing protein 2 | 11303 | -0.099 | 0.0220 | No |
| 63 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 11380 | -0.101 | 0.0290 | No |
| 64 | THY1 | THY1 Entrez, Source | Thy-1 cell surface antigen | 11400 | -0.102 | 0.0404 | No |
| 65 | RGS14 | RGS14 Entrez, Source | regulator of G-protein signalling 14 | 11575 | -0.108 | 0.0410 | No |
| 66 | ARL2 | ARL2 Entrez, Source | ADP-ribosylation factor-like 2 | 12040 | -0.128 | 0.0222 | No |
| 67 | RASA1 | RASA1 Entrez, Source | RAS p21 protein activator (GTPase activating protein) 1 | 12457 | -0.152 | 0.0100 | No |
| 68 | EIF2B5 | EIF2B5 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 5 epsilon, 82kDa | 12669 | -0.170 | 0.0155 | No |
| 69 | TRIO | TRIO Entrez, Source | triple functional domain (PTPRF interacting) | 13166 | -0.279 | 0.0133 | No |