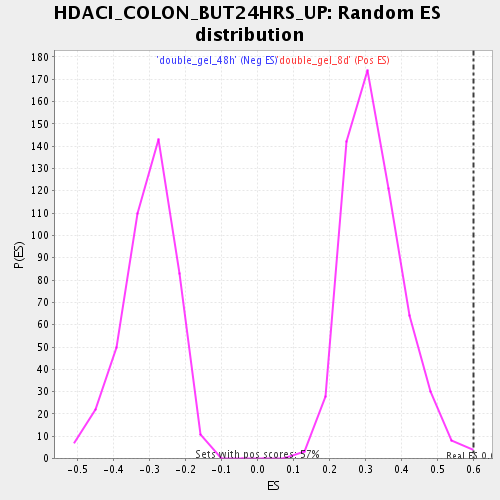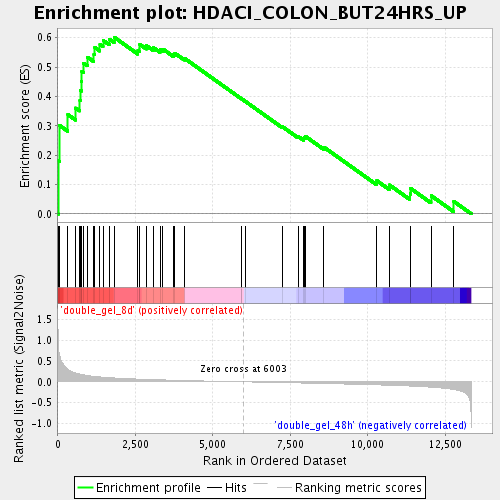
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | HDACI_COLON_BUT24HRS_UP |
| Enrichment Score (ES) | 0.6015331 |
| Normalized Enrichment Score (NES) | 1.8548237 |
| Nominal p-value | 0.0034843206 |
| FDR q-value | 0.018239953 |
| FWER p-Value | 0.827 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 9 | 0.936 | 0.1820 | Yes |
| 2 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 56 | 0.628 | 0.3009 | Yes |
| 3 | ZSWIM6 | ZSWIM6 Entrez, Source | zinc finger, SWIM-type containing 6 | 320 | 0.292 | 0.3382 | Yes |
| 4 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 571 | 0.212 | 0.3607 | Yes |
| 5 | RHOB | RHOB Entrez, Source | ras homolog gene family, member B | 709 | 0.184 | 0.3863 | Yes |
| 6 | GABBR1 | GABBR1 Entrez, Source | gamma-aminobutyric acid (GABA) B receptor, 1 | 722 | 0.182 | 0.4209 | Yes |
| 7 | TIMP1 | TIMP1 Entrez, Source | TIMP metallopeptidase inhibitor 1 | 759 | 0.176 | 0.4525 | Yes |
| 8 | CALCOCO1 | CALCOCO1 Entrez, Source | calcium binding and coiled-coil domain 1 | 771 | 0.174 | 0.4856 | Yes |
| 9 | NEU1 | NEU1 Entrez, Source | sialidase 1 (lysosomal sialidase) | 827 | 0.166 | 0.5138 | Yes |
| 10 | PLD3 | PLD3 Entrez, Source | phospholipase D family, member 3 | 959 | 0.151 | 0.5333 | Yes |
| 11 | PLCG2 | PLCG2 Entrez, Source | phospholipase C, gamma 2 (phosphatidylinositol-specific) | 1161 | 0.131 | 0.5439 | Yes |
| 12 | ANXA5 | ANXA5 Entrez, Source | annexin A5 | 1191 | 0.129 | 0.5668 | Yes |
| 13 | SLC27A1 | SLC27A1 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 1 | 1356 | 0.116 | 0.5771 | Yes |
| 14 | IL17RE | IL17RE Entrez, Source | interleukin 17 receptor E | 1457 | 0.110 | 0.5910 | Yes |
| 15 | BIRC4 | BIRC4 Entrez, Source | baculoviral IAP repeat-containing 4 | 1664 | 0.099 | 0.5949 | Yes |
| 16 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 1816 | 0.092 | 0.6015 | Yes |
| 17 | GRN | GRN Entrez, Source | granulin | 2581 | 0.065 | 0.5568 | No |
| 18 | HSF1 | HSF1 Entrez, Source | heat shock transcription factor 1 | 2641 | 0.064 | 0.5648 | No |
| 19 | PSAP | PSAP Entrez, Source | prosaposin (variant Gaucher disease and variant metachromatic leukodystrophy) | 2642 | 0.064 | 0.5773 | No |
| 20 | MAP1LC3B | MAP1LC3B Entrez, Source | microtubule-associated protein 1 light chain 3 beta | 2854 | 0.059 | 0.5729 | No |
| 21 | INPP5E | INPP5E Entrez, Source | inositol polyphosphate-5-phosphatase, 72 kDa | 3078 | 0.052 | 0.5663 | No |
| 22 | CDH1 | CDH1 Entrez, Source | cadherin 1, type 1, E-cadherin (epithelial) | 3296 | 0.047 | 0.5592 | No |
| 23 | ROCK1 | ROCK1 Entrez, Source | Rho-associated, coiled-coil containing protein kinase 1 | 3389 | 0.045 | 0.5612 | No |
| 24 | CLU | CLU Entrez, Source | clusterin | 3739 | 0.038 | 0.5423 | No |
| 25 | SPARCL1 | SPARCL1 Entrez, Source | SPARC-like 1 (mast9, hevin) | 3771 | 0.037 | 0.5473 | No |
| 26 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 4084 | 0.031 | 0.5299 | No |
| 27 | KCNA1 | KCNA1 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 1 (episodic ataxia with myokymia) | 5912 | 0.001 | 0.3928 | No |
| 28 | ATP6AP2 | ATP6AP2 Entrez, Source | ATPase, H+ transporting, lysosomal accessory protein 2 | 5915 | 0.001 | 0.3929 | No |
| 29 | CD81 | CD81 Entrez, Source | CD81 molecule | 6049 | -0.001 | 0.3830 | No |
| 30 | DNM3 | DNM3 Entrez, Source | dynamin 3 | 7241 | -0.019 | 0.2972 | No |
| 31 | HSP90AA1 | HSP90AA1 Entrez, Source | heat shock protein 90kDa alpha (cytosolic), class A member 1 | 7748 | -0.027 | 0.2643 | No |
| 32 | RXRB | RXRB Entrez, Source | retinoid X receptor, beta | 7930 | -0.030 | 0.2565 | No |
| 33 | SMARCD3 | SMARCD3 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 | 7948 | -0.030 | 0.2610 | No |
| 34 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 7989 | -0.031 | 0.2640 | No |
| 35 | FLNB | FLNB Entrez, Source | filamin B, beta (actin binding protein 278) | 8583 | -0.040 | 0.2273 | No |
| 36 | PRDX1 | PRDX1 Entrez, Source | peroxiredoxin 1 | 10293 | -0.072 | 0.1129 | No |
| 37 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 10699 | -0.083 | 0.0986 | No |
| 38 | TSNAX | TSNAX Entrez, Source | translin-associated factor X | 11363 | -0.101 | 0.0684 | No |
| 39 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 11380 | -0.101 | 0.0869 | No |
| 40 | SKAP2 | SKAP2 Entrez, Source | src kinase associated phosphoprotein 2 | 12042 | -0.128 | 0.0623 | No |
| 41 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 12777 | -0.181 | 0.0425 | No |