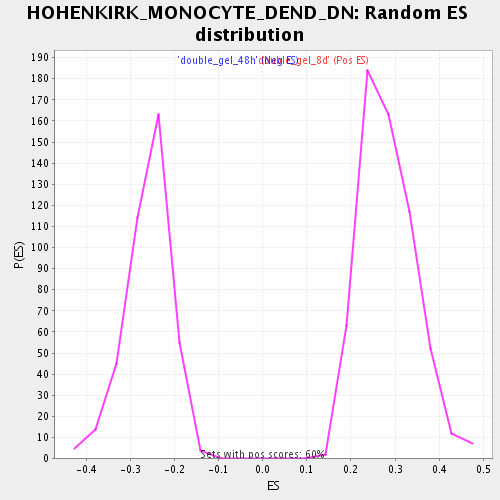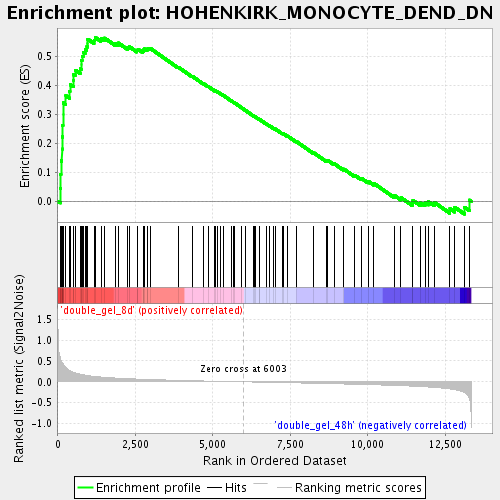
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | HOHENKIRK_MONOCYTE_DEND_DN |
| Enrichment Score (ES) | 0.5666313 |
| Normalized Enrichment Score (NES) | 2.0039809 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0043760356 |
| FWER p-Value | 0.191 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CORO1A | CORO1A Entrez, Source | coronin, actin binding protein, 1A | 77 | 0.560 | 0.0458 | Yes |
| 2 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 89 | 0.531 | 0.0940 | Yes |
| 3 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 101 | 0.510 | 0.1401 | Yes |
| 4 | LGALS2 | LGALS2 Entrez, Source | lectin, galactoside-binding, soluble, 2 (galectin 2) | 132 | 0.466 | 0.1808 | Yes |
| 5 | CXCL2 | CXCL2 Entrez, Source | chemokine (C-X-C motif) ligand 2 | 138 | 0.458 | 0.2227 | Yes |
| 6 | LYZ | LYZ Entrez, Source | lysozyme (renal amyloidosis) | 153 | 0.442 | 0.2624 | Yes |
| 7 | MARCKSL1 | MARCKSL1 Entrez, Source | MARCKS-like 1 | 166 | 0.428 | 0.3009 | Yes |
| 8 | AMPD3 | AMPD3 Entrez, Source | adenosine monophosphate deaminase (isoform E) | 167 | 0.426 | 0.3402 | Yes |
| 9 | PTK2B | PTK2B Entrez, Source | PTK2B protein tyrosine kinase 2 beta | 256 | 0.337 | 0.3646 | Yes |
| 10 | PLSCR1 | PLSCR1 Entrez, Source | phospholipid scramblase 1 | 376 | 0.270 | 0.3805 | Yes |
| 11 | IER3 | IER3 Entrez, Source | immediate early response 3 | 392 | 0.264 | 0.4037 | Yes |
| 12 | ZFP36L2 | ZFP36L2 Entrez, Source | zinc finger protein 36, C3H type-like 2 | 498 | 0.228 | 0.4168 | Yes |
| 13 | PF4 | PF4 Entrez, Source | platelet factor 4 (chemokine (C-X-C motif) ligand 4) | 506 | 0.226 | 0.4371 | Yes |
| 14 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 558 | 0.215 | 0.4530 | Yes |
| 15 | FBP1 | FBP1 Entrez, Source | fructose-1,6-bisphosphatase 1 | 720 | 0.182 | 0.4577 | Yes |
| 16 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 754 | 0.176 | 0.4714 | Yes |
| 17 | EMP3 | EMP3 Entrez, Source | epithelial membrane protein 3 | 755 | 0.176 | 0.4877 | Yes |
| 18 | HADH2 | HADH2 Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase, type II | 782 | 0.173 | 0.5016 | Yes |
| 19 | ITPR1 | ITPR1 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 1 | 816 | 0.167 | 0.5146 | Yes |
| 20 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 880 | 0.159 | 0.5244 | Yes |
| 21 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 934 | 0.153 | 0.5345 | Yes |
| 22 | S100A8 | S100A8 Entrez, Source | S100 calcium binding protein A8 | 960 | 0.151 | 0.5465 | Yes |
| 23 | S100A4 | S100A4 Entrez, Source | S100 calcium binding protein A4 | 962 | 0.151 | 0.5603 | Yes |
| 24 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 1166 | 0.131 | 0.5571 | Yes |
| 25 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 1197 | 0.128 | 0.5666 | Yes |
| 26 | ATP2A3 | ATP2A3 Entrez, Source | ATPase, Ca++ transporting, ubiquitous | 1390 | 0.114 | 0.5627 | No |
| 27 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 1504 | 0.108 | 0.5641 | No |
| 28 | LGMN | LGMN Entrez, Source | legumain | 1864 | 0.090 | 0.5453 | No |
| 29 | CTBP1 | CTBP1 Entrez, Source | C-terminal binding protein 1 | 1948 | 0.087 | 0.5471 | No |
| 30 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 2256 | 0.075 | 0.5308 | No |
| 31 | SOS2 | SOS2 Entrez, Source | son of sevenless homolog 2 (Drosophila) | 2302 | 0.074 | 0.5343 | No |
| 32 | CSF1R | CSF1R Entrez, Source | colony stimulating factor 1 receptor, formerly McDonough feline sarcoma viral (v-fms) oncogene homolog | 2552 | 0.066 | 0.5216 | No |
| 33 | IL2RG | IL2RG Entrez, Source | interleukin 2 receptor, gamma (severe combined immunodeficiency) | 2582 | 0.065 | 0.5254 | No |
| 34 | C3AR1 | C3AR1 Entrez, Source | complement component 3a receptor 1 | 2746 | 0.061 | 0.5188 | No |
| 35 | ITGAL | ITGAL Entrez, Source | integrin, alpha L (antigen CD11A (p180), lymphocyte function-associated antigen 1; alpha polypeptide) | 2766 | 0.061 | 0.5230 | No |
| 36 | CCR2 | CCR2 Entrez, Source | chemokine (C-C motif) receptor 2 | 2793 | 0.060 | 0.5266 | No |
| 37 | HTRA1 | HTRA1 Entrez, Source | HtrA serine peptidase 1 | 2880 | 0.058 | 0.5254 | No |
| 38 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 2902 | 0.057 | 0.5291 | No |
| 39 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 2983 | 0.055 | 0.5282 | No |
| 40 | DOK1 | DOK1 Entrez, Source | docking protein 1, 62kDa (downstream of tyrosine kinase 1) | 3881 | 0.035 | 0.4637 | No |
| 41 | PPIF | PPIF Entrez, Source | peptidylprolyl isomerase F (cyclophilin F) | 4337 | 0.027 | 0.4318 | No |
| 42 | SERPINF1 | SERPINF1 Entrez, Source | serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 | 4697 | 0.020 | 0.4066 | No |
| 43 | SPINK1 | SPINK1 Entrez, Source | serine peptidase inhibitor, Kazal type 1 | 4848 | 0.018 | 0.3969 | No |
| 44 | RASA3 | RASA3 Entrez, Source | RAS p21 protein activator 3 | 5066 | 0.014 | 0.3818 | No |
| 45 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 5086 | 0.014 | 0.3817 | No |
| 46 | LST1 | LST1 Entrez, Source | leukocyte specific transcript 1 | 5138 | 0.013 | 0.3790 | No |
| 47 | PPIB | PPIB Entrez, Source | peptidylprolyl isomerase B (cyclophilin B) | 5253 | 0.011 | 0.3714 | No |
| 48 | C3 | C3 Entrez, Source | complement component 3 | 5333 | 0.010 | 0.3663 | No |
| 49 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 5599 | 0.006 | 0.3468 | No |
| 50 | DNAJC8 | DNAJC8 Entrez, Source | DnaJ (Hsp40) homolog, subfamily C, member 8 | 5671 | 0.005 | 0.3419 | No |
| 51 | C5AR1 | C5AR1 Entrez, Source | complement component 5a receptor 1 | 5691 | 0.004 | 0.3409 | No |
| 52 | TNF | TNF Entrez, Source | tumor necrosis factor (TNF superfamily, member 2) | 5914 | 0.001 | 0.3243 | No |
| 53 | CD37 | CD37 Entrez, Source | CD37 molecule | 6038 | -0.001 | 0.3150 | No |
| 54 | SF3B4 | SF3B4 Entrez, Source | splicing factor 3b, subunit 4, 49kDa | 6322 | -0.005 | 0.2942 | No |
| 55 | NCF1 | NCF1 Entrez, Source | neutrophil cytosolic factor 1, (chronic granulomatous disease, autosomal 1) | 6355 | -0.006 | 0.2923 | No |
| 56 | ADAM10 | ADAM10 Entrez, Source | ADAM metallopeptidase domain 10 | 6375 | -0.006 | 0.2914 | No |
| 57 | SERPINA1 | SERPINA1 Entrez, Source | serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 1 | 6493 | -0.008 | 0.2832 | No |
| 58 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 6500 | -0.008 | 0.2835 | No |
| 59 | PDLIM7 | PDLIM7 Entrez, Source | PDZ and LIM domain 7 (enigma) | 6732 | -0.011 | 0.2671 | No |
| 60 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 6833 | -0.013 | 0.2607 | No |
| 61 | ANPEP | ANPEP Entrez, Source | alanyl (membrane) aminopeptidase (aminopeptidase N, aminopeptidase M, microsomal aminopeptidase, CD13, p150) | 6961 | -0.014 | 0.2525 | No |
| 62 | NAP1L4 | NAP1L4 Entrez, Source | nucleosome assembly protein 1-like 4 | 7012 | -0.015 | 0.2501 | No |
| 63 | BST1 | BST1 Entrez, Source | bone marrow stromal cell antigen 1 | 7247 | -0.019 | 0.2342 | No |
| 64 | PPBP | PPBP Entrez, Source | pro-platelet basic protein (chemokine (C-X-C motif) ligand 7) | 7274 | -0.019 | 0.2340 | No |
| 65 | NFYC | NFYC Entrez, Source | nuclear transcription factor Y, gamma | 7410 | -0.021 | 0.2258 | No |
| 66 | IRF7 | IRF7 Entrez, Source | interferon regulatory factor 7 | 7693 | -0.026 | 0.2069 | No |
| 67 | CYBB | CYBB Entrez, Source | cytochrome b-245, beta polypeptide (chronic granulomatous disease) | 8263 | -0.035 | 0.1672 | No |
| 68 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 8667 | -0.041 | 0.1406 | No |
| 69 | CD14 | CD14 Entrez, Source | CD14 molecule | 8706 | -0.042 | 0.1416 | No |
| 70 | SDC2 | SDC2 Entrez, Source | syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) | 8914 | -0.046 | 0.1302 | No |
| 71 | PPP2R5D | PPP2R5D Entrez, Source | protein phosphatase 2, regulatory subunit B (B56), delta isoform | 9222 | -0.051 | 0.1118 | No |
| 72 | DHX8 | DHX8 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 8 | 9578 | -0.057 | 0.0903 | No |
| 73 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 9785 | -0.062 | 0.0804 | No |
| 74 | SIAHBP1 | SIAHBP1 Entrez, Source | - | 10014 | -0.067 | 0.0694 | No |
| 75 | SLC1A3 | SLC1A3 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 3 | 10194 | -0.070 | 0.0624 | No |
| 76 | PSEN1 | PSEN1 Entrez, Source | presenilin 1 (Alzheimer disease 3) | 10866 | -0.087 | 0.0198 | No |
| 77 | PBEF1 | PBEF1 Entrez, Source | pre-B-cell colony enhancing factor 1 | 11072 | -0.092 | 0.0128 | No |
| 78 | TSC22D1 | TSC22D1 Entrez, Source | TSC22 domain family, member 1 | 11438 | -0.104 | -0.0051 | No |
| 79 | NINJ1 | NINJ1 Entrez, Source | ninjurin 1 | 11455 | -0.104 | 0.0033 | No |
| 80 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 11710 | -0.114 | -0.0054 | No |
| 81 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 11850 | -0.119 | -0.0049 | No |
| 82 | BAT2 | BAT2 Entrez, Source | HLA-B associated transcript 2 | 11954 | -0.124 | -0.0012 | No |
| 83 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 12150 | -0.134 | -0.0036 | No |
| 84 | ALG3 | ALG3 Entrez, Source | asparagine-linked glycosylation 3 homolog (S. cerevisiae, alpha-1,3-mannosyltransferase) | 12651 | -0.168 | -0.0258 | No |
| 85 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 12802 | -0.185 | -0.0200 | No |
| 86 | GSR | GSR Entrez, Source | glutathione reductase | 13122 | -0.259 | -0.0203 | No |
| 87 | CES1 | CES1 Entrez, Source | carboxylesterase 1 (monocyte/macrophage serine esterase 1) | 13281 | -0.399 | 0.0046 | No |