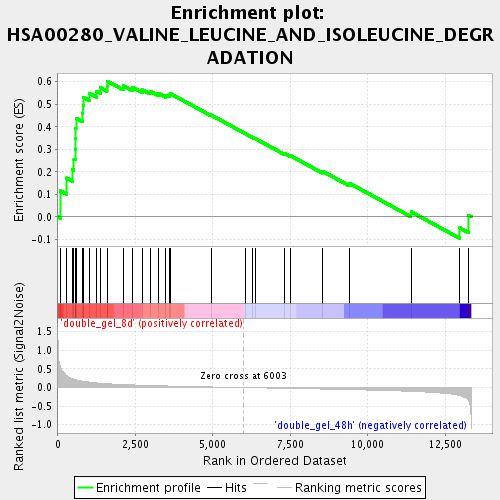
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | HSA00280_VALINE_LEUCINE_AND_ISOLEUCINE_DEGRADATION |
| Enrichment Score (ES) | 0.6029134 |
| Normalized Enrichment Score (NES) | 1.8171209 |
| Nominal p-value | 0.0036429872 |
| FDR q-value | 0.022726113 |
| FWER p-Value | 0.935 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | HMGCS2 | HMGCS2 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 2 (mitochondrial) | 83 | 0.552 | 0.1153 | Yes |
| 2 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 272 | 0.326 | 0.1729 | Yes |
| 3 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 470 | 0.238 | 0.2104 | Yes |
| 4 | ALDH3A1 | ALDH3A1 Entrez, Source | aldehyde dehydrogenase 3 family, memberA1 | 517 | 0.223 | 0.2561 | Yes |
| 5 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 558 | 0.215 | 0.3004 | Yes |
| 6 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 566 | 0.213 | 0.3467 | Yes |
| 7 | DBT | DBT Entrez, Source | dihydrolipoamide branched chain transacylase E2 | 577 | 0.210 | 0.3922 | Yes |
| 8 | ECHS1 | ECHS1 Entrez, Source | enoyl Coenzyme A hydratase, short chain, 1, mitochondrial | 585 | 0.209 | 0.4376 | Yes |
| 9 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 798 | 0.170 | 0.4591 | Yes |
| 10 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 809 | 0.168 | 0.4954 | Yes |
| 11 | EHHADH | EHHADH Entrez, Source | enoyl-Coenzyme A, hydratase/3-hydroxyacyl Coenzyme A dehydrogenase | 826 | 0.166 | 0.5306 | Yes |
| 12 | ACADS | ACADS Entrez, Source | acyl-Coenzyme A dehydrogenase, C-2 to C-3 short chain | 1025 | 0.145 | 0.5476 | Yes |
| 13 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 1252 | 0.124 | 0.5578 | Yes |
| 14 | HSD17B4 | HSD17B4 Entrez, Source | hydroxysteroid (17-beta) dehydrogenase 4 | 1381 | 0.115 | 0.5734 | Yes |
| 15 | ALDH6A1 | ALDH6A1 Entrez, Source | aldehyde dehydrogenase 6 family, member A1 | 1585 | 0.103 | 0.5808 | Yes |
| 16 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 1594 | 0.103 | 0.6029 | Yes |
| 17 | MCCC2 | MCCC2 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 2 (beta) | 2111 | 0.080 | 0.5818 | No |
| 18 | IVD | IVD Entrez, Source | isovaleryl Coenzyme A dehydrogenase | 2412 | 0.070 | 0.5748 | No |
| 19 | HMGCL | HMGCL Entrez, Source | 3-hydroxymethyl-3-methylglutaryl-Coenzyme A lyase (hydroxymethylglutaricaciduria) | 2739 | 0.062 | 0.5639 | No |
| 20 | PCCB | PCCB Entrez, Source | propionyl Coenzyme A carboxylase, beta polypeptide | 2977 | 0.055 | 0.5582 | No |
| 21 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 3245 | 0.048 | 0.5488 | No |
| 22 | ALDH1B1 | ALDH1B1 Entrez, Source | aldehyde dehydrogenase 1 family, member B1 | 3480 | 0.043 | 0.5407 | No |
| 23 | ALDH7A1 | ALDH7A1 Entrez, Source | aldehyde dehydrogenase 7 family, member A1 | 3592 | 0.041 | 0.5414 | No |
| 24 | HIBADH | HIBADH Entrez, Source | 3-hydroxyisobutyrate dehydrogenase | 3631 | 0.040 | 0.5474 | No |
| 25 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4941 | 0.016 | 0.4526 | No |
| 26 | HIBCH | HIBCH Entrez, Source | 3-hydroxyisobutyryl-Coenzyme A hydrolase | 6051 | -0.001 | 0.3694 | No |
| 27 | MCCC1 | MCCC1 Entrez, Source | methylcrotonoyl-Coenzyme A carboxylase 1 (alpha) | 6280 | -0.004 | 0.3532 | No |
| 28 | HADHB | HADHB Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), beta subunit | 6362 | -0.006 | 0.3484 | No |
| 29 | ABAT | ABAT Entrez, Source | 4-aminobutyrate aminotransferase | 7326 | -0.020 | 0.2804 | No |
| 30 | BCAT2 | BCAT2 Entrez, Source | branched chain aminotransferase 2, mitochondrial | 7503 | -0.022 | 0.2721 | No |
| 31 | ALDH9A1 | ALDH9A1 Entrez, Source | aldehyde dehydrogenase 9 family, member A1 | 8536 | -0.039 | 0.2033 | No |
| 32 | DLD | DLD Entrez, Source | dihydrolipoamide dehydrogenase | 9409 | -0.054 | 0.1497 | No |
| 33 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 11395 | -0.102 | 0.0230 | No |
| 34 | ACAT2 | ACAT2 Entrez, Source | acetyl-Coenzyme A acetyltransferase 2 (acetoacetyl Coenzyme A thiolase) | 12971 | -0.213 | -0.0485 | No |
| 35 | AOX1 | AOX1 Entrez, Source | aldehyde oxidase 1 | 13251 | -0.347 | 0.0068 | No |

