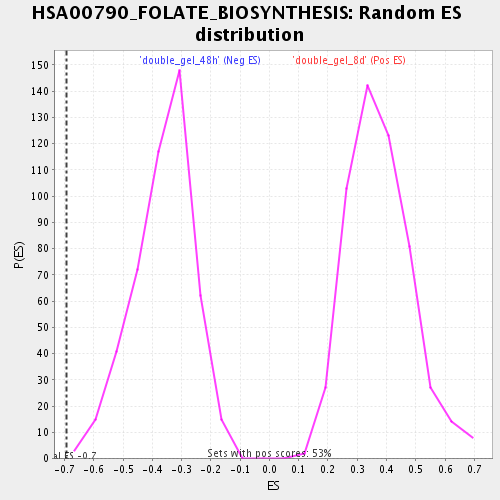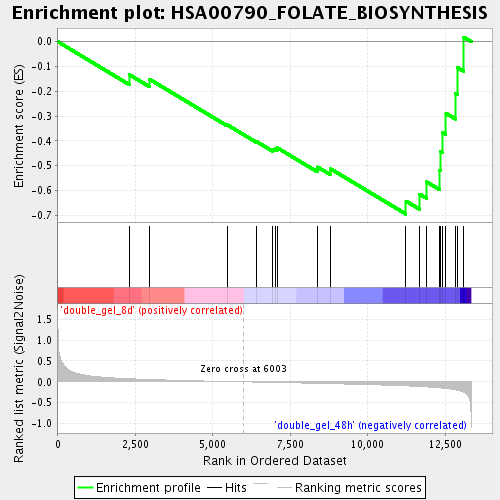
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | HSA00790_FOLATE_BIOSYNTHESIS |
| Enrichment Score (ES) | -0.69502765 |
| Normalized Enrichment Score (NES) | -1.9193428 |
| Nominal p-value | 0.0021141649 |
| FDR q-value | 0.01908465 |
| FWER p-Value | 0.385 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 2300 | 0.074 | -0.1323 | No |
| 2 | ALPL | ALPL Entrez, Source | alkaline phosphatase, liver/bone/kidney | 2954 | 0.056 | -0.1509 | No |
| 3 | ALPI | ALPI Entrez, Source | alkaline phosphatase, intestinal | 5468 | 0.008 | -0.3353 | No |
| 4 | DDX50 | DDX50 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 50 | 6409 | -0.006 | -0.4024 | No |
| 5 | DHFR | DHFR Entrez, Source | dihydrofolate reductase | 6938 | -0.014 | -0.4344 | No |
| 6 | PTS | PTS Entrez, Source | 6-pyruvoyltetrahydropterin synthase | 7010 | -0.015 | -0.4314 | No |
| 7 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 7085 | -0.016 | -0.4280 | No |
| 8 | SPR | SPR Entrez, Source | sepiapterin reductase (7,8-dihydrobiopterin:NADP+ oxidoreductase) | 8382 | -0.037 | -0.5051 | No |
| 9 | DDX47 | DDX47 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 | 8805 | -0.044 | -0.5129 | No |
| 10 | DDX52 | DDX52 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 52 | 11233 | -0.097 | -0.6425 | Yes |
| 11 | QDPR | QDPR Entrez, Source | quinoid dihydropteridine reductase | 11673 | -0.112 | -0.6144 | Yes |
| 12 | DDX18 | DDX18 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 18 | 11900 | -0.122 | -0.5651 | Yes |
| 13 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 12316 | -0.144 | -0.5180 | Yes |
| 14 | DDX56 | DDX56 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 56 | 12337 | -0.145 | -0.4408 | Yes |
| 15 | NUDT5 | NUDT5 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 5 | 12409 | -0.149 | -0.3650 | Yes |
| 16 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 12523 | -0.156 | -0.2885 | Yes |
| 17 | EP400 | EP400 Entrez, Source | E1A binding protein p400 | 12834 | -0.190 | -0.2083 | Yes |
| 18 | GGH | GGH Entrez, Source | gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) | 12902 | -0.200 | -0.1046 | Yes |
| 19 | ASCC3L1 | ASCC3L1 Entrez, Source | activating signal cointegrator 1 complex subunit 3-like 1 | 13101 | -0.253 | 0.0181 | Yes |