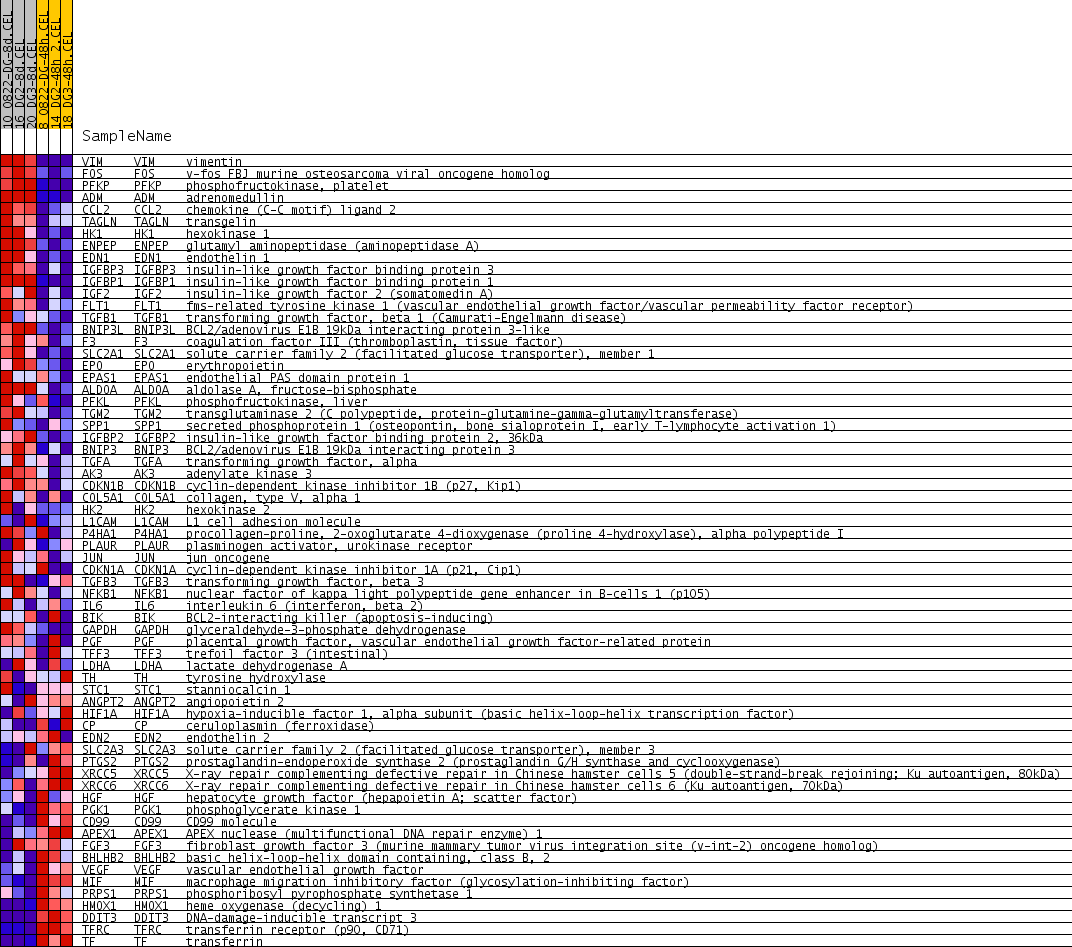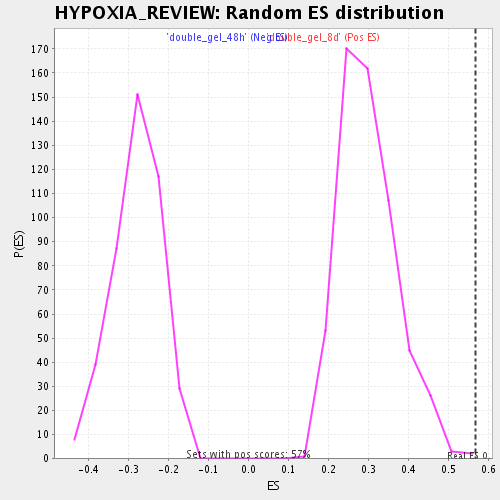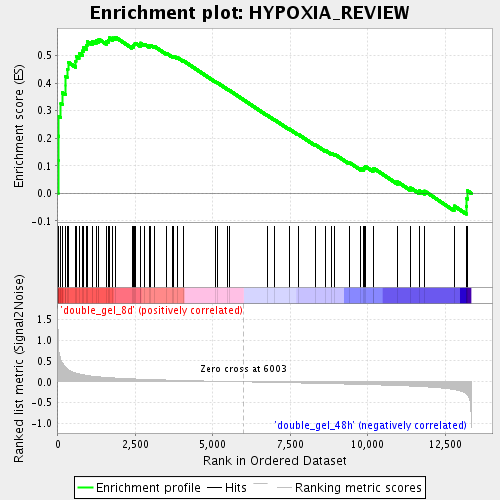
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | HYPOXIA_REVIEW |
| Enrichment Score (ES) | 0.5668321 |
| Normalized Enrichment Score (NES) | 1.891723 |
| Nominal p-value | 0.0017574693 |
| FDR q-value | 0.013440252 |
| FWER p-Value | 0.647 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VIM | VIM Entrez, Source | vimentin | 4 | 1.225 | 0.1181 | Yes |
| 2 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 9 | 0.936 | 0.2083 | Yes |
| 3 | PFKP | PFKP Entrez, Source | phosphofructokinase, platelet | 29 | 0.753 | 0.2796 | Yes |
| 4 | ADM | ADM Entrez, Source | adrenomedullin | 96 | 0.523 | 0.3252 | Yes |
| 5 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 145 | 0.452 | 0.3652 | Yes |
| 6 | TAGLN | TAGLN Entrez, Source | transgelin | 250 | 0.344 | 0.3906 | Yes |
| 7 | HK1 | HK1 Entrez, Source | hexokinase 1 | 251 | 0.342 | 0.4236 | Yes |
| 8 | ENPEP | ENPEP Entrez, Source | glutamyl aminopeptidase (aminopeptidase A) | 298 | 0.310 | 0.4501 | Yes |
| 9 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 326 | 0.291 | 0.4763 | Yes |
| 10 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 571 | 0.212 | 0.4784 | Yes |
| 11 | IGFBP1 | IGFBP1 Entrez, Source | insulin-like growth factor binding protein 1 | 584 | 0.209 | 0.4976 | Yes |
| 12 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 682 | 0.188 | 0.5085 | Yes |
| 13 | FLT1 | FLT1 Entrez, Source | fms-related tyrosine kinase 1 (vascular endothelial growth factor/vascular permeability factor receptor) | 789 | 0.172 | 0.5171 | Yes |
| 14 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 821 | 0.166 | 0.5309 | Yes |
| 15 | BNIP3L | BNIP3L Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3-like | 921 | 0.154 | 0.5383 | Yes |
| 16 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 943 | 0.152 | 0.5514 | Yes |
| 17 | SLC2A1 | SLC2A1 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 1 | 1110 | 0.135 | 0.5519 | Yes |
| 18 | EPO | EPO Entrez, Source | erythropoietin | 1235 | 0.125 | 0.5546 | Yes |
| 19 | EPAS1 | EPAS1 Entrez, Source | endothelial PAS domain protein 1 | 1321 | 0.118 | 0.5596 | Yes |
| 20 | ALDOA | ALDOA Entrez, Source | aldolase A, fructose-bisphosphate | 1564 | 0.104 | 0.5515 | Yes |
| 21 | PFKL | PFKL Entrez, Source | phosphofructokinase, liver | 1624 | 0.101 | 0.5568 | Yes |
| 22 | TGM2 | TGM2 Entrez, Source | transglutaminase 2 (C polypeptide, protein-glutamine-gamma-glutamyltransferase) | 1650 | 0.100 | 0.5646 | Yes |
| 23 | SPP1 | SPP1 Entrez, Source | secreted phosphoprotein 1 (osteopontin, bone sialoprotein I, early T-lymphocyte activation 1) | 1776 | 0.094 | 0.5643 | Yes |
| 24 | IGFBP2 | IGFBP2 Entrez, Source | insulin-like growth factor binding protein 2, 36kDa | 1859 | 0.090 | 0.5668 | Yes |
| 25 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 2391 | 0.071 | 0.5337 | No |
| 26 | TGFA | TGFA Entrez, Source | transforming growth factor, alpha | 2430 | 0.070 | 0.5376 | No |
| 27 | AK3 | AK3 Entrez, Source | adenylate kinase 3 | 2456 | 0.069 | 0.5424 | No |
| 28 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 2499 | 0.068 | 0.5458 | No |
| 29 | COL5A1 | COL5A1 Entrez, Source | collagen, type V, alpha 1 | 2660 | 0.063 | 0.5399 | No |
| 30 | HK2 | HK2 Entrez, Source | hexokinase 2 | 2661 | 0.063 | 0.5460 | No |
| 31 | L1CAM | L1CAM Entrez, Source | L1 cell adhesion molecule | 2799 | 0.060 | 0.5415 | No |
| 32 | P4HA1 | P4HA1 Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha polypeptide I | 2948 | 0.056 | 0.5357 | No |
| 33 | PLAUR | PLAUR Entrez, Source | plasminogen activator, urokinase receptor | 2983 | 0.055 | 0.5385 | No |
| 34 | JUN | JUN Entrez, Source | jun oncogene | 3104 | 0.052 | 0.5344 | No |
| 35 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 3506 | 0.043 | 0.5084 | No |
| 36 | TGFB3 | TGFB3 Entrez, Source | transforming growth factor, beta 3 | 3692 | 0.039 | 0.4982 | No |
| 37 | NFKB1 | NFKB1 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (p105) | 3740 | 0.038 | 0.4983 | No |
| 38 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 3854 | 0.036 | 0.4932 | No |
| 39 | BIK | BIK Entrez, Source | BCL2-interacting killer (apoptosis-inducing) | 4042 | 0.032 | 0.4822 | No |
| 40 | GAPDH | GAPDH Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase | 5074 | 0.014 | 0.4059 | No |
| 41 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 5143 | 0.012 | 0.4020 | No |
| 42 | TFF3 | TFF3 Entrez, Source | trefoil factor 3 (intestinal) | 5480 | 0.008 | 0.3774 | No |
| 43 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 5536 | 0.007 | 0.3739 | No |
| 44 | TH | TH Entrez, Source | tyrosine hydroxylase | 6761 | -0.012 | 0.2829 | No |
| 45 | STC1 | STC1 Entrez, Source | stanniocalcin 1 | 6986 | -0.015 | 0.2674 | No |
| 46 | ANGPT2 | ANGPT2 Entrez, Source | angiopoietin 2 | 7458 | -0.022 | 0.2341 | No |
| 47 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 7751 | -0.027 | 0.2147 | No |
| 48 | CP | CP Entrez, Source | ceruloplasmin (ferroxidase) | 8305 | -0.036 | 0.1765 | No |
| 49 | EDN2 | EDN2 Entrez, Source | endothelin 2 | 8626 | -0.041 | 0.1563 | No |
| 50 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 8829 | -0.044 | 0.1454 | No |
| 51 | PTGS2 | PTGS2 Entrez, Source | prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) | 8937 | -0.046 | 0.1418 | No |
| 52 | XRCC5 | XRCC5 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 5 (double-strand-break rejoining; Ku autoantigen, 80kDa) | 9405 | -0.054 | 0.1118 | No |
| 53 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 9780 | -0.062 | 0.0896 | No |
| 54 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 9857 | -0.063 | 0.0900 | No |
| 55 | PGK1 | PGK1 Entrez, Source | phosphoglycerate kinase 1 | 9880 | -0.064 | 0.0945 | No |
| 56 | CD99 | CD99 Entrez, Source | CD99 molecule | 9921 | -0.065 | 0.0978 | No |
| 57 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 10173 | -0.070 | 0.0856 | No |
| 58 | FGF3 | FGF3 Entrez, Source | fibroblast growth factor 3 (murine mammary tumor virus integration site (v-int-2) oncogene homolog) | 10191 | -0.070 | 0.0911 | No |
| 59 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 10960 | -0.089 | 0.0419 | No |
| 60 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 11386 | -0.102 | 0.0197 | No |
| 61 | MIF | MIF Entrez, Source | macrophage migration inhibitory factor (glycosylation-inhibiting factor) | 11668 | -0.112 | 0.0094 | No |
| 62 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 11832 | -0.118 | 0.0085 | No |
| 63 | HMOX1 | HMOX1 Entrez, Source | heme oxygenase (decycling) 1 | 12788 | -0.183 | -0.0458 | No |
| 64 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 13182 | -0.289 | -0.0475 | No |
| 65 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 13197 | -0.299 | -0.0196 | No |
| 66 | TF | TF Entrez, Source | transferrin | 13216 | -0.315 | 0.0095 | No |