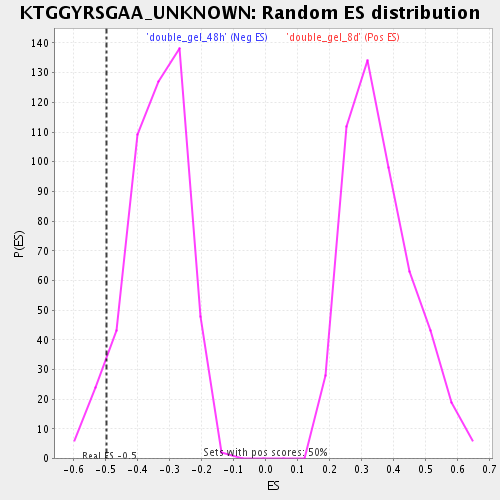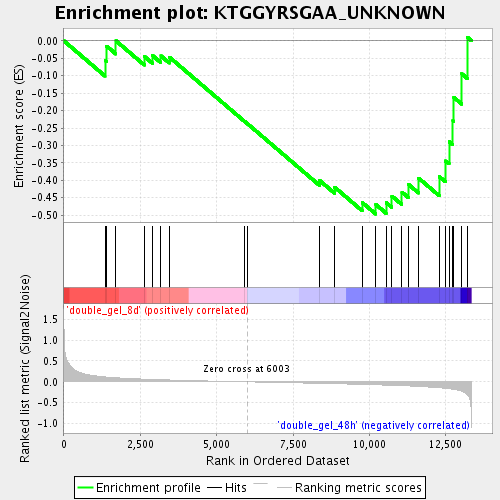
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | KTGGYRSGAA_UNKNOWN |
| Enrichment Score (ES) | -0.49700078 |
| Normalized Enrichment Score (NES) | -1.4584267 |
| Nominal p-value | 0.062374245 |
| FDR q-value | 0.28115305 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 1345 | 0.117 | -0.0555 | No |
| 2 | SH3KBP1 | SH3KBP1 Entrez, Source | SH3-domain kinase binding protein 1 | 1398 | 0.114 | -0.0150 | No |
| 3 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 1703 | 0.097 | 0.0002 | No |
| 4 | GDI1 | GDI1 Entrez, Source | GDP dissociation inhibitor 1 | 2631 | 0.064 | -0.0444 | No |
| 5 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 2897 | 0.057 | -0.0419 | No |
| 6 | CNTFR | CNTFR Entrez, Source | ciliary neurotrophic factor receptor | 3176 | 0.050 | -0.0432 | No |
| 7 | DLGAP4 | DLGAP4 Entrez, Source | discs, large (Drosophila) homolog-associated protein 4 | 3449 | 0.044 | -0.0464 | No |
| 8 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 5905 | 0.001 | -0.2302 | No |
| 9 | ORC1L | ORC1L Entrez, Source | origin recognition complex, subunit 1-like (yeast) | 5994 | 0.000 | -0.2367 | No |
| 10 | KCND2 | KCND2 Entrez, Source | potassium voltage-gated channel, Shal-related subfamily, member 2 | 8361 | -0.037 | -0.4000 | No |
| 11 | SPRED1 | SPRED1 Entrez, Source | sprouty-related, EVH1 domain containing 1 | 8858 | -0.045 | -0.4198 | No |
| 12 | SEC61A1 | SEC61A1 Entrez, Source | Sec61 alpha 1 subunit (S. cerevisiae) | 9764 | -0.061 | -0.4638 | No |
| 13 | TUBA4 | TUBA4 Entrez, Source | tubulin, alpha 4 | 10207 | -0.071 | -0.4694 | Yes |
| 14 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 10547 | -0.079 | -0.4640 | Yes |
| 15 | HTF9C | HTF9C Entrez, Source | - | 10732 | -0.084 | -0.4452 | Yes |
| 16 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 11062 | -0.092 | -0.4339 | Yes |
| 17 | SFRS7 | SFRS7 Entrez, Source | splicing factor, arginine/serine-rich 7, 35kDa | 11272 | -0.098 | -0.4115 | Yes |
| 18 | YARS | YARS Entrez, Source | tyrosyl-tRNA synthetase | 11607 | -0.110 | -0.3937 | Yes |
| 19 | PRIM1 | PRIM1 Entrez, Source | primase, polypeptide 1, 49kDa | 12284 | -0.141 | -0.3893 | Yes |
| 20 | LIG1 | LIG1 Entrez, Source | ligase I, DNA, ATP-dependent | 12490 | -0.154 | -0.3445 | Yes |
| 21 | PIM1 | PIM1 Entrez, Source | pim-1 oncogene | 12606 | -0.164 | -0.2892 | Yes |
| 22 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 12730 | -0.176 | -0.2297 | Yes |
| 23 | CDCA7 | CDCA7 Entrez, Source | cell division cycle associated 7 | 12759 | -0.178 | -0.1620 | Yes |
| 24 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13021 | -0.223 | -0.0945 | Yes |
| 25 | MCM6 | MCM6 Entrez, Source | MCM6 minichromosome maintenance deficient 6 (MIS5 homolog, S. pombe) (S. cerevisiae) | 13204 | -0.304 | 0.0104 | Yes |