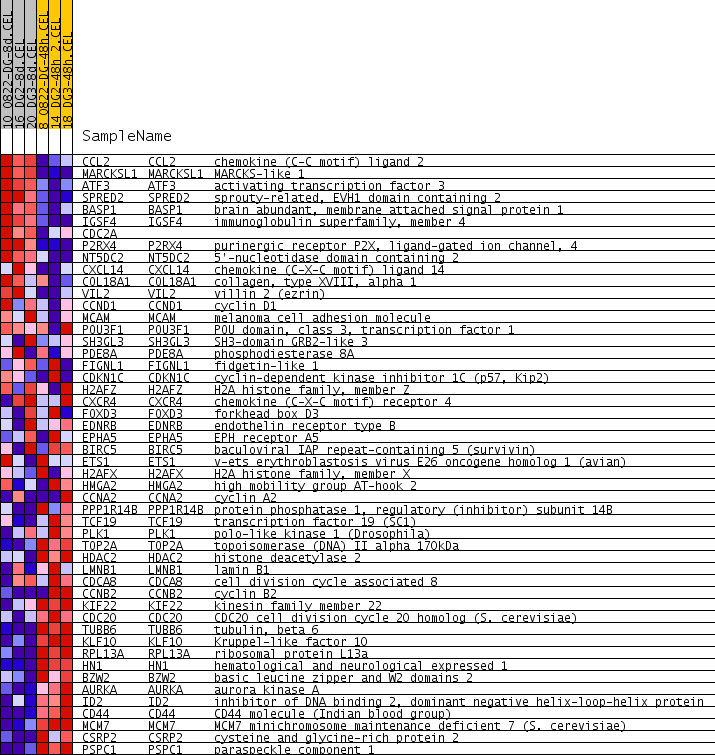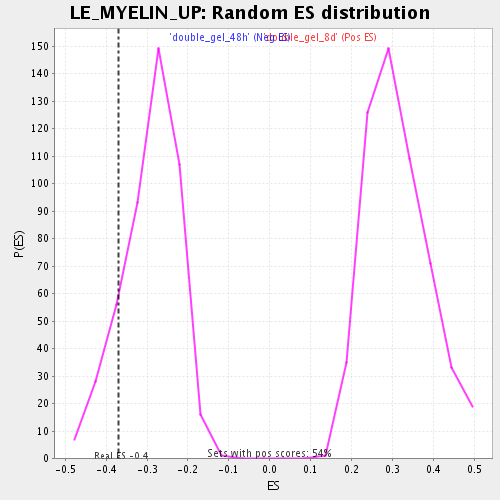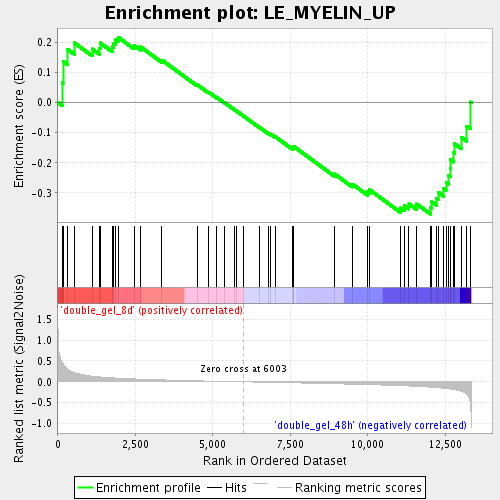
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | LE_MYELIN_UP |
| Enrichment Score (ES) | -0.3702065 |
| Normalized Enrichment Score (NES) | -1.2664925 |
| Nominal p-value | 0.14660831 |
| FDR q-value | 0.52797335 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 145 | 0.452 | 0.0657 | No |
| 2 | MARCKSL1 | MARCKSL1 Entrez, Source | MARCKS-like 1 | 166 | 0.428 | 0.1368 | No |
| 3 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 310 | 0.299 | 0.1769 | No |
| 4 | SPRED2 | SPRED2 Entrez, Source | sprouty-related, EVH1 domain containing 2 | 528 | 0.221 | 0.1981 | No |
| 5 | BASP1 | BASP1 Entrez, Source | brain abundant, membrane attached signal protein 1 | 1105 | 0.136 | 0.1778 | No |
| 6 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 1345 | 0.117 | 0.1796 | No |
| 7 | CDC2A | 1363 | 0.115 | 0.1979 | No | ||
| 8 | P2RX4 | P2RX4 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 4 | 1753 | 0.095 | 0.1847 | No |
| 9 | NT5DC2 | NT5DC2 Entrez, Source | 5'-nucleotidase domain containing 2 | 1797 | 0.093 | 0.1973 | No |
| 10 | CXCL14 | CXCL14 Entrez, Source | chemokine (C-X-C motif) ligand 14 | 1856 | 0.090 | 0.2082 | No |
| 11 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 1942 | 0.087 | 0.2166 | No |
| 12 | VIL2 | VIL2 Entrez, Source | villin 2 (ezrin) | 2459 | 0.069 | 0.1895 | No |
| 13 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 2668 | 0.063 | 0.1846 | No |
| 14 | MCAM | MCAM Entrez, Source | melanoma cell adhesion molecule | 3354 | 0.046 | 0.1409 | No |
| 15 | POU3F1 | POU3F1 Entrez, Source | POU domain, class 3, transcription factor 1 | 4491 | 0.024 | 0.0594 | No |
| 16 | SH3GL3 | SH3GL3 Entrez, Source | SH3-domain GRB2-like 3 | 4859 | 0.017 | 0.0348 | No |
| 17 | PDE8A | PDE8A Entrez, Source | phosphodiesterase 8A | 5133 | 0.013 | 0.0164 | No |
| 18 | FIGNL1 | FIGNL1 Entrez, Source | fidgetin-like 1 | 5379 | 0.009 | -0.0005 | No |
| 19 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 5704 | 0.004 | -0.0241 | No |
| 20 | H2AFZ | H2AFZ Entrez, Source | H2A histone family, member Z | 5766 | 0.003 | -0.0282 | No |
| 21 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 5981 | 0.000 | -0.0442 | No |
| 22 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 6499 | -0.008 | -0.0818 | No |
| 23 | EDNRB | EDNRB Entrez, Source | endothelin receptor type B | 6802 | -0.012 | -0.1025 | No |
| 24 | EPHA5 | EPHA5 Entrez, Source | EPH receptor A5 | 6875 | -0.013 | -0.1057 | No |
| 25 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 7015 | -0.015 | -0.1135 | No |
| 26 | ETS1 | ETS1 Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog 1 (avian) | 7566 | -0.024 | -0.1509 | No |
| 27 | H2AFX | H2AFX Entrez, Source | H2A histone family, member X | 7594 | -0.024 | -0.1488 | No |
| 28 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 7610 | -0.024 | -0.1458 | No |
| 29 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 8927 | -0.046 | -0.2370 | No |
| 30 | PPP1R14B | PPP1R14B Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 14B | 9504 | -0.056 | -0.2709 | No |
| 31 | TCF19 | TCF19 Entrez, Source | transcription factor 19 (SC1) | 9995 | -0.066 | -0.2965 | No |
| 32 | PLK1 | PLK1 Entrez, Source | polo-like kinase 1 (Drosophila) | 10056 | -0.067 | -0.2896 | No |
| 33 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 11066 | -0.092 | -0.3498 | Yes |
| 34 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 11182 | -0.095 | -0.3423 | Yes |
| 35 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 11330 | -0.100 | -0.3365 | Yes |
| 36 | CDCA8 | CDCA8 Entrez, Source | cell division cycle associated 8 | 11577 | -0.108 | -0.3366 | Yes |
| 37 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12025 | -0.128 | -0.3486 | Yes |
| 38 | KIF22 | KIF22 Entrez, Source | kinesin family member 22 | 12066 | -0.130 | -0.3295 | Yes |
| 39 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 12218 | -0.137 | -0.3176 | Yes |
| 40 | TUBB6 | TUBB6 Entrez, Source | tubulin, beta 6 | 12270 | -0.141 | -0.2976 | Yes |
| 41 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 12453 | -0.151 | -0.2855 | Yes |
| 42 | RPL13A | RPL13A Entrez, Source | ribosomal protein L13a | 12527 | -0.156 | -0.2645 | Yes |
| 43 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 12612 | -0.164 | -0.2430 | Yes |
| 44 | BZW2 | BZW2 Entrez, Source | basic leucine zipper and W2 domains 2 | 12664 | -0.170 | -0.2180 | Yes |
| 45 | AURKA | AURKA Entrez, Source | aurora kinase A | 12673 | -0.170 | -0.1897 | Yes |
| 46 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 12777 | -0.181 | -0.1667 | Yes |
| 47 | CD44 | CD44 Entrez, Source | CD44 molecule (Indian blood group) | 12802 | -0.185 | -0.1370 | Yes |
| 48 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13021 | -0.223 | -0.1156 | Yes |
| 49 | CSRP2 | CSRP2 Entrez, Source | cysteine and glycine-rich protein 2 | 13196 | -0.299 | -0.0780 | Yes |
| 50 | PSPC1 | PSPC1 Entrez, Source | paraspeckle component 1 | 13317 | -0.524 | 0.0019 | Yes |