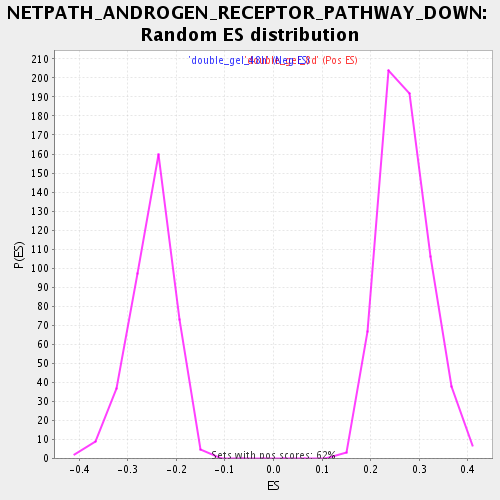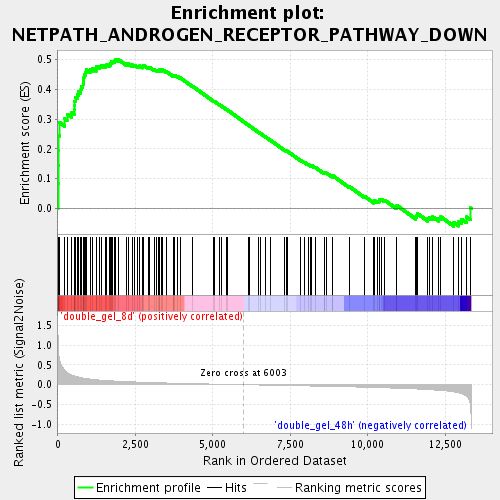
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | NETPATH_ANDROGEN_RECEPTOR_PATHWAY_DOWN |
| Enrichment Score (ES) | 0.5010938 |
| Normalized Enrichment Score (NES) | 1.8569158 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.017889446 |
| FWER p-Value | 0.822 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CYP17A1 | CYP17A1 Entrez, Source | cytochrome P450, family 17, subfamily A, polypeptide 1 | 3 | 1.233 | 0.0843 | Yes |
| 2 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 10 | 0.885 | 0.1445 | Yes |
| 3 | TIMP2 | TIMP2 Entrez, Source | TIMP metallopeptidase inhibitor 2 | 28 | 0.756 | 0.1950 | Yes |
| 4 | COL5A2 | COL5A2 Entrez, Source | collagen, type V, alpha 2 | 34 | 0.714 | 0.2436 | Yes |
| 5 | LPL | LPL Entrez, Source | lipoprotein lipase | 45 | 0.671 | 0.2888 | Yes |
| 6 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 222 | 0.366 | 0.3006 | Yes |
| 7 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 297 | 0.311 | 0.3164 | Yes |
| 8 | PKIB | PKIB Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor beta | 446 | 0.246 | 0.3220 | Yes |
| 9 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 525 | 0.222 | 0.3313 | Yes |
| 10 | CUTL1 | CUTL1 Entrez, Source | cut-like 1, CCAAT displacement protein (Drosophila) | 531 | 0.219 | 0.3460 | Yes |
| 11 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 538 | 0.219 | 0.3606 | Yes |
| 12 | IGFBP3 | IGFBP3 Entrez, Source | insulin-like growth factor binding protein 3 | 571 | 0.212 | 0.3727 | Yes |
| 13 | CRAT | CRAT Entrez, Source | carnitine acetyltransferase | 625 | 0.202 | 0.3825 | Yes |
| 14 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 659 | 0.195 | 0.3934 | Yes |
| 15 | NUCB2 | NUCB2 Entrez, Source | nucleobindin 2 | 743 | 0.177 | 0.3993 | Yes |
| 16 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 756 | 0.176 | 0.4104 | Yes |
| 17 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 809 | 0.168 | 0.4180 | Yes |
| 18 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 821 | 0.166 | 0.4286 | Yes |
| 19 | CX3CL1 | CX3CL1 Entrez, Source | chemokine (C-X3-C motif) ligand 1 | 837 | 0.164 | 0.4387 | Yes |
| 20 | PDK2 | PDK2 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 2 | 846 | 0.163 | 0.4493 | Yes |
| 21 | BTG2 | BTG2 Entrez, Source | BTG family, member 2 | 880 | 0.159 | 0.4577 | Yes |
| 22 | ERBB2 | ERBB2 Entrez, Source | v-erb-b2 erythroblastic leukemia viral oncogene homolog 2, neuro/glioblastoma derived oncogene homolog (avian) | 916 | 0.154 | 0.4656 | Yes |
| 23 | HRG | HRG Entrez, Source | histidine-rich glycoprotein | 1039 | 0.142 | 0.4661 | Yes |
| 24 | AGC1 | AGC1 Entrez, Source | aggrecan 1 (chondroitin sulfate proteoglycan 1, large aggregating proteoglycan, antigen identified by monoclonal antibody A0122) | 1118 | 0.135 | 0.4695 | Yes |
| 25 | SMPD1 | SMPD1 Entrez, Source | sphingomyelin phosphodiesterase 1, acid lysosomal (acid sphingomyelinase) | 1232 | 0.125 | 0.4695 | Yes |
| 26 | PLEKHB1 | PLEKHB1 Entrez, Source | pleckstrin homology domain containing, family B (evectins) member 1 | 1247 | 0.124 | 0.4770 | Yes |
| 27 | BARD1 | BARD1 Entrez, Source | BRCA1 associated RING domain 1 | 1344 | 0.117 | 0.4777 | Yes |
| 28 | RALB | RALB Entrez, Source | v-ral simian leukemia viral oncogene homolog B (ras related; GTP binding protein) | 1401 | 0.113 | 0.4812 | Yes |
| 29 | RNASE4 | RNASE4 Entrez, Source | ribonuclease, RNase A family, 4 | 1525 | 0.107 | 0.4793 | Yes |
| 30 | GCHFR | GCHFR Entrez, Source | GTP cyclohydrolase I feedback regulator | 1580 | 0.104 | 0.4823 | Yes |
| 31 | P2RY2 | P2RY2 Entrez, Source | purinergic receptor P2Y, G-protein coupled, 2 | 1654 | 0.100 | 0.4836 | Yes |
| 32 | CAMK2N2 | CAMK2N2 Entrez, Source | calcium/calmodulin-dependent protein kinase II inhibitor 2 | 1689 | 0.098 | 0.4877 | Yes |
| 33 | PAMCI | PAMCI Entrez, Source | peptidylglycine alpha-amidating monooxygenase COOH-terminal interactor | 1713 | 0.097 | 0.4926 | Yes |
| 34 | SPP1 | SPP1 Entrez, Source | secreted phosphoprotein 1 (osteopontin, bone sialoprotein I, early T-lymphocyte activation 1) | 1776 | 0.094 | 0.4944 | Yes |
| 35 | STXBP1 | STXBP1 Entrez, Source | syntaxin binding protein 1 | 1838 | 0.091 | 0.4960 | Yes |
| 36 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 1854 | 0.090 | 0.5011 | Yes |
| 37 | CTBP1 | CTBP1 Entrez, Source | C-terminal binding protein 1 | 1948 | 0.087 | 0.5000 | No |
| 38 | IGF1R | IGF1R Entrez, Source | insulin-like growth factor 1 receptor | 2201 | 0.077 | 0.4863 | No |
| 39 | SNX1 | SNX1 Entrez, Source | sorting nexin 1 | 2279 | 0.075 | 0.4856 | No |
| 40 | FLOT2 | FLOT2 Entrez, Source | flotillin 2 | 2393 | 0.071 | 0.4819 | No |
| 41 | PAM | PAM Entrez, Source | peptidylglycine alpha-amidating monooxygenase | 2485 | 0.069 | 0.4797 | No |
| 42 | GRN | GRN Entrez, Source | granulin | 2581 | 0.065 | 0.4770 | No |
| 43 | SERPINI1 | SERPINI1 Entrez, Source | serpin peptidase inhibitor, clade I (neuroserpin), member 1 | 2621 | 0.064 | 0.4785 | No |
| 44 | FN1 | FN1 Entrez, Source | fibronectin 1 | 2715 | 0.062 | 0.4757 | No |
| 45 | HMGCL | HMGCL Entrez, Source | 3-hydroxymethyl-3-methylglutaryl-Coenzyme A lyase (hydroxymethylglutaricaciduria) | 2739 | 0.062 | 0.4782 | No |
| 46 | BSG | BSG Entrez, Source | basigin (Ok blood group) | 2772 | 0.061 | 0.4800 | No |
| 47 | MINPP1 | MINPP1 Entrez, Source | multiple inositol polyphosphate histidine phosphatase, 1 | 2918 | 0.057 | 0.4729 | No |
| 48 | P4HA1 | P4HA1 Entrez, Source | procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha polypeptide I | 2948 | 0.056 | 0.4745 | No |
| 49 | JUN | JUN Entrez, Source | jun oncogene | 3104 | 0.052 | 0.4664 | No |
| 50 | CNTFR | CNTFR Entrez, Source | ciliary neurotrophic factor receptor | 3176 | 0.050 | 0.4644 | No |
| 51 | ALDH2 | ALDH2 Entrez, Source | aldehyde dehydrogenase 2 family (mitochondrial) | 3245 | 0.048 | 0.4626 | No |
| 52 | OXT | OXT Entrez, Source | oxytocin, prepro- (neurophysin I) | 3271 | 0.048 | 0.4640 | No |
| 53 | PCTK1 | PCTK1 Entrez, Source | PCTAIRE protein kinase 1 | 3282 | 0.048 | 0.4665 | No |
| 54 | DDC | DDC Entrez, Source | dopa decarboxylase (aromatic L-amino acid decarboxylase) | 3357 | 0.046 | 0.4641 | No |
| 55 | SDC4 | SDC4 Entrez, Source | syndecan 4 (amphiglycan, ryudocan) | 3385 | 0.045 | 0.4652 | No |
| 56 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 3506 | 0.043 | 0.4590 | No |
| 57 | CLU | CLU Entrez, Source | clusterin | 3739 | 0.038 | 0.4441 | No |
| 58 | NFKB1 | NFKB1 Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (p105) | 3740 | 0.038 | 0.4467 | No |
| 59 | CACNB3 | CACNB3 Entrez, Source | calcium channel, voltage-dependent, beta 3 subunit | 3774 | 0.037 | 0.4467 | No |
| 60 | IL6 | IL6 Entrez, Source | interleukin 6 (interferon, beta 2) | 3854 | 0.036 | 0.4432 | No |
| 61 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 3945 | 0.034 | 0.4387 | No |
| 62 | SULT2A1 | SULT2A1 Entrez, Source | sulfotransferase family, cytosolic, 2A, dehydroepiandrosterone (DHEA)-preferring, member 1 | 4352 | 0.026 | 0.4098 | No |
| 63 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 5025 | 0.015 | 0.3601 | No |
| 64 | TLE3 | TLE3 Entrez, Source | transducin-like enhancer of split 3 (E(sp1) homolog, Drosophila) | 5051 | 0.014 | 0.3591 | No |
| 65 | PLA2G2A | PLA2G2A Entrez, Source | phospholipase A2, group IIA (platelets, synovial fluid) | 5199 | 0.012 | 0.3488 | No |
| 66 | KLK3 | KLK3 Entrez, Source | kallikrein 3, (prostate specific antigen) | 5271 | 0.011 | 0.3442 | No |
| 67 | ACP6 | ACP6 Entrez, Source | acid phosphatase 6, lysophosphatidic | 5425 | 0.008 | 0.3332 | No |
| 68 | ST7 | ST7 Entrez, Source | suppression of tumorigenicity 7 | 5485 | 0.007 | 0.3293 | No |
| 69 | ACSL5 | ACSL5 Entrez, Source | acyl-CoA synthetase long-chain family member 5 | 6137 | -0.002 | 0.2802 | No |
| 70 | SQSTM1 | SQSTM1 Entrez, Source | sequestosome 1 | 6184 | -0.003 | 0.2769 | No |
| 71 | ENO2 | ENO2 Entrez, Source | enolase 2 (gamma, neuronal) | 6473 | -0.007 | 0.2557 | No |
| 72 | ACPP | ACPP Entrez, Source | acid phosphatase, prostate | 6543 | -0.008 | 0.2510 | No |
| 73 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 6690 | -0.011 | 0.2407 | No |
| 74 | LRRN1 | LRRN1 Entrez, Source | leucine rich repeat neuronal 1 | 6849 | -0.013 | 0.2296 | No |
| 75 | GATA2 | GATA2 Entrez, Source | GATA binding protein 2 | 7313 | -0.020 | 0.1960 | No |
| 76 | SFRP4 | SFRP4 Entrez, Source | secreted frizzled-related protein 4 | 7370 | -0.021 | 0.1932 | No |
| 77 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 7396 | -0.021 | 0.1927 | No |
| 78 | NOTCH1 | NOTCH1 Entrez, Source | Notch homolog 1, translocation-associated (Drosophila) | 7842 | -0.028 | 0.1610 | No |
| 79 | SMARCD3 | SMARCD3 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 | 7948 | -0.030 | 0.1551 | No |
| 80 | STAR | STAR Entrez, Source | steroidogenic acute regulator | 8077 | -0.032 | 0.1476 | No |
| 81 | PIK3R3 | PIK3R3 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 3 (p55, gamma) | 8161 | -0.033 | 0.1436 | No |
| 82 | KIF5B | KIF5B Entrez, Source | kinesin family member 5B | 8195 | -0.034 | 0.1435 | No |
| 83 | CASP9 | CASP9 Entrez, Source | caspase 9, apoptosis-related cysteine peptidase | 8297 | -0.036 | 0.1383 | No |
| 84 | CGA | CGA Entrez, Source | glycoprotein hormones, alpha polypeptide | 8595 | -0.040 | 0.1186 | No |
| 85 | ASNS | ASNS Entrez, Source | asparagine synthetase | 8596 | -0.040 | 0.1214 | No |
| 86 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 8667 | -0.041 | 0.1189 | No |
| 87 | PDYN | PDYN Entrez, Source | prodynorphin | 8860 | -0.045 | 0.1075 | No |
| 88 | MBTPS1 | MBTPS1 Entrez, Source | membrane-bound transcription factor peptidase, site 1 | 8862 | -0.045 | 0.1105 | No |
| 89 | SLC44A1 | SLC44A1 Entrez, Source | solute carrier family 44, member 1 | 9410 | -0.054 | 0.0729 | No |
| 90 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 9886 | -0.064 | 0.0413 | No |
| 91 | SCNN1A | SCNN1A Entrez, Source | sodium channel, nonvoltage-gated 1 alpha | 10199 | -0.071 | 0.0226 | No |
| 92 | COLEC12 | COLEC12 Entrez, Source | collectin sub-family member 12 | 10217 | -0.071 | 0.0262 | No |
| 93 | ATP2B1 | ATP2B1 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 1 | 10314 | -0.073 | 0.0239 | No |
| 94 | CAMK2N1 | CAMK2N1 Entrez, Source | calcium/calmodulin-dependent protein kinase II inhibitor 1 | 10364 | -0.074 | 0.0253 | No |
| 95 | SNRK | SNRK Entrez, Source | SNF related kinase | 10380 | -0.074 | 0.0292 | No |
| 96 | GLI3 | GLI3 Entrez, Source | GLI-Kruppel family member GLI3 (Greig cephalopolysyndactyly syndrome) | 10439 | -0.076 | 0.0300 | No |
| 97 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 10547 | -0.079 | 0.0274 | No |
| 98 | MDK | MDK Entrez, Source | midkine (neurite growth-promoting factor 2) | 10921 | -0.089 | 0.0052 | No |
| 99 | FZD3 | FZD3 Entrez, Source | frizzled homolog 3 (Drosophila) | 10939 | -0.089 | 0.0101 | No |
| 100 | SLC12A2 | SLC12A2 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 2 | 11551 | -0.108 | -0.0287 | No |
| 101 | NFIX | NFIX Entrez, Source | nuclear factor I/X (CCAAT-binding transcription factor) | 11585 | -0.109 | -0.0238 | No |
| 102 | PRKD1 | PRKD1 Entrez, Source | protein kinase D1 | 11594 | -0.109 | -0.0169 | No |
| 103 | SLC20A2 | SLC20A2 Entrez, Source | solute carrier family 20 (phosphate transporter), member 2 | 11939 | -0.123 | -0.0345 | No |
| 104 | MMP16 | MMP16 Entrez, Source | matrix metallopeptidase 16 (membrane-inserted) | 11993 | -0.126 | -0.0298 | No |
| 105 | EFNA1 | EFNA1 Entrez, Source | ephrin-A1 | 12095 | -0.131 | -0.0285 | No |
| 106 | BCHE | BCHE Entrez, Source | butyrylcholinesterase | 12286 | -0.141 | -0.0331 | No |
| 107 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 12360 | -0.146 | -0.0286 | No |
| 108 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 12777 | -0.181 | -0.0476 | No |
| 109 | SLC29A1 | SLC29A1 Entrez, Source | solute carrier family 29 (nucleoside transporters), member 1 | 12934 | -0.205 | -0.0453 | No |
| 110 | CEBPA | CEBPA Entrez, Source | CCAAT/enhancer binding protein (C/EBP), alpha | 13029 | -0.225 | -0.0370 | No |
| 111 | FOXQ1 | FOXQ1 Entrez, Source | forkhead box Q1 | 13187 | -0.293 | -0.0288 | No |
| 112 | GSTA2 | GSTA2 Entrez, Source | glutathione S-transferase A2 | 13321 | -0.590 | 0.0016 | No |