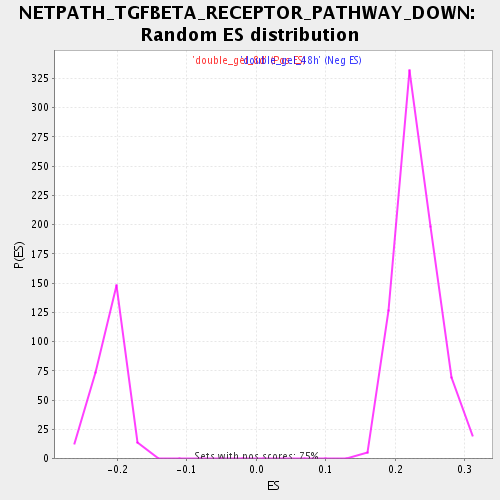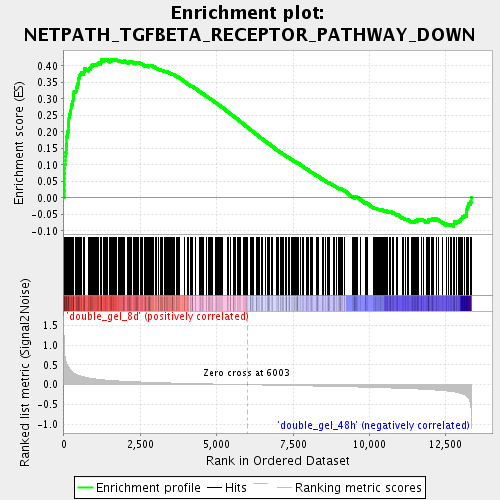
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | NETPATH_TGFBETA_RECEPTOR_PATHWAY_DOWN |
| Enrichment Score (ES) | 0.42029062 |
| Normalized Enrichment Score (NES) | 1.8186687 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.02313824 |
| FWER p-Value | 0.932 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | VIM | VIM Entrez, Source | vimentin | 4 | 1.225 | 0.0225 | Yes |
| 2 | ADAMTS1 | ADAMTS1 Entrez, Source | ADAM metallopeptidase with thrombospondin type 1 motif, 1 | 8 | 1.023 | 0.0413 | Yes |
| 3 | CCND2 | CCND2 Entrez, Source | cyclin D2 | 10 | 0.885 | 0.0576 | Yes |
| 4 | MAP3K8 | MAP3K8 Entrez, Source | mitogen-activated protein kinase kinase kinase 8 | 12 | 0.878 | 0.0739 | Yes |
| 5 | LAMC2 | LAMC2 Entrez, Source | laminin, gamma 2 | 19 | 0.791 | 0.0881 | Yes |
| 6 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 25 | 0.771 | 0.1021 | Yes |
| 7 | DBP | DBP Entrez, Source | D site of albumin promoter (albumin D-box) binding protein | 41 | 0.688 | 0.1137 | Yes |
| 8 | LPL | LPL Entrez, Source | lipoprotein lipase | 45 | 0.671 | 0.1260 | Yes |
| 9 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 56 | 0.628 | 0.1369 | Yes |
| 10 | CYP1A1 | CYP1A1 Entrez, Source | cytochrome P450, family 1, subfamily A, polypeptide 1 | 72 | 0.571 | 0.1463 | Yes |
| 11 | GPRC5A | GPRC5A Entrez, Source | G protein-coupled receptor, family C, group 5, member A | 78 | 0.559 | 0.1563 | Yes |
| 12 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 87 | 0.537 | 0.1657 | Yes |
| 13 | RNASE1 | RNASE1 Entrez, Source | ribonuclease, RNase A family, 1 (pancreatic) | 94 | 0.529 | 0.1751 | Yes |
| 14 | ADM | ADM Entrez, Source | adrenomedullin | 96 | 0.523 | 0.1847 | Yes |
| 15 | CRYAB | CRYAB Entrez, Source | crystallin, alpha B | 106 | 0.494 | 0.1932 | Yes |
| 16 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 115 | 0.484 | 0.2016 | Yes |
| 17 | F2R | F2R Entrez, Source | coagulation factor II (thrombin) receptor | 142 | 0.455 | 0.2080 | Yes |
| 18 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 145 | 0.452 | 0.2162 | Yes |
| 19 | APOC1 | APOC1 Entrez, Source | apolipoprotein C-I | 151 | 0.445 | 0.2241 | Yes |
| 20 | CLDN1 | CLDN1 Entrez, Source | claudin 1 | 158 | 0.436 | 0.2318 | Yes |
| 21 | BMP2 | BMP2 Entrez, Source | bone morphogenetic protein 2 | 164 | 0.430 | 0.2394 | Yes |
| 22 | FST | FST Entrez, Source | follistatin | 169 | 0.423 | 0.2470 | Yes |
| 23 | BGN | BGN Entrez, Source | biglycan | 175 | 0.408 | 0.2542 | Yes |
| 24 | NOL3 | NOL3 Entrez, Source | nucleolar protein 3 (apoptosis repressor with CARD domain) | 202 | 0.385 | 0.2593 | Yes |
| 25 | CDH13 | CDH13 Entrez, Source | cadherin 13, H-cadherin (heart) | 223 | 0.366 | 0.2646 | Yes |
| 26 | ST14 | ST14 Entrez, Source | suppression of tumorigenicity 14 (colon carcinoma) | 232 | 0.362 | 0.2707 | Yes |
| 27 | PLA1A | PLA1A Entrez, Source | phospholipase A1 member A | 236 | 0.355 | 0.2770 | Yes |
| 28 | LAMC1 | LAMC1 Entrez, Source | laminin, gamma 1 (formerly LAMB2) | 252 | 0.342 | 0.2822 | Yes |
| 29 | PLCL1 | PLCL1 Entrez, Source | phospholipase C-like 1 | 269 | 0.328 | 0.2871 | Yes |
| 30 | FMO3 | FMO3 Entrez, Source | flavin containing monooxygenase 3 | 281 | 0.322 | 0.2922 | Yes |
| 31 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 297 | 0.311 | 0.2968 | Yes |
| 32 | HSPA4 | HSPA4 Entrez, Source | heat shock 70kDa protein 4 | 305 | 0.303 | 0.3019 | Yes |
| 33 | ATF3 | ATF3 Entrez, Source | activating transcription factor 3 | 310 | 0.299 | 0.3072 | Yes |
| 34 | TFPI | TFPI Entrez, Source | tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) | 316 | 0.294 | 0.3122 | Yes |
| 35 | EDN1 | EDN1 Entrez, Source | endothelin 1 | 326 | 0.291 | 0.3169 | Yes |
| 36 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 329 | 0.291 | 0.3222 | Yes |
| 37 | GDF15 | GDF15 Entrez, Source | growth differentiation factor 15 | 385 | 0.267 | 0.3229 | Yes |
| 38 | BTD | BTD Entrez, Source | biotinidase | 401 | 0.259 | 0.3265 | Yes |
| 39 | PTGER4 | PTGER4 Entrez, Source | prostaglandin E receptor 4 (subtype EP4) | 402 | 0.258 | 0.3313 | Yes |
| 40 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 404 | 0.258 | 0.3360 | Yes |
| 41 | RGL2 | RGL2 Entrez, Source | ral guanine nucleotide dissociation stimulator-like 2 | 439 | 0.247 | 0.3380 | Yes |
| 42 | RGS4 | RGS4 Entrez, Source | regulator of G-protein signalling 4 | 454 | 0.244 | 0.3414 | Yes |
| 43 | RBP4 | RBP4 Entrez, Source | retinol binding protein 4, plasma | 458 | 0.242 | 0.3457 | Yes |
| 44 | LEPR | LEPR Entrez, Source | leptin receptor | 471 | 0.237 | 0.3492 | Yes |
| 45 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 472 | 0.237 | 0.3536 | Yes |
| 46 | THBD | THBD Entrez, Source | thrombomodulin | 474 | 0.236 | 0.3579 | Yes |
| 47 | SLC20A1 | SLC20A1 Entrez, Source | solute carrier family 20 (phosphate transporter), member 1 | 484 | 0.232 | 0.3615 | Yes |
| 48 | ZFP36L2 | ZFP36L2 Entrez, Source | zinc finger protein 36, C3H type-like 2 | 498 | 0.228 | 0.3647 | Yes |
| 49 | RGS5 | RGS5 Entrez, Source | regulator of G-protein signalling 5 | 513 | 0.224 | 0.3678 | Yes |
| 50 | VCAM1 | VCAM1 Entrez, Source | vascular cell adhesion molecule 1 | 516 | 0.223 | 0.3718 | Yes |
| 51 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 538 | 0.219 | 0.3743 | Yes |
| 52 | EHD3 | EHD3 Entrez, Source | EH-domain containing 3 | 559 | 0.215 | 0.3767 | Yes |
| 53 | CLDN4 | CLDN4 Entrez, Source | claudin 4 | 570 | 0.212 | 0.3799 | Yes |
| 54 | CCND3 | CCND3 Entrez, Source | cyclin D3 | 643 | 0.198 | 0.3779 | Yes |
| 55 | RASSF1 | RASSF1 Entrez, Source | Ras association (RalGDS/AF-6) domain family 1 | 652 | 0.196 | 0.3810 | Yes |
| 56 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 659 | 0.195 | 0.3841 | Yes |
| 57 | TMSB10 | TMSB10 Entrez, Source | thymosin, beta 10 | 670 | 0.192 | 0.3869 | Yes |
| 58 | COL1A1 | COL1A1 Entrez, Source | collagen, type I, alpha 1 | 673 | 0.190 | 0.3903 | Yes |
| 59 | IGF2 | IGF2 Entrez, Source | insulin-like growth factor 2 (somatomedin A) | 682 | 0.188 | 0.3932 | Yes |
| 60 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 798 | 0.170 | 0.3874 | Yes |
| 61 | CYB5A | CYB5A Entrez, Source | cytochrome b5 type A (microsomal) | 817 | 0.167 | 0.3891 | Yes |
| 62 | TGFB1 | TGFB1 Entrez, Source | transforming growth factor, beta 1 (Camurati-Engelmann disease) | 821 | 0.166 | 0.3920 | Yes |
| 63 | CCL7 | CCL7 Entrez, Source | chemokine (C-C motif) ligand 7 | 860 | 0.161 | 0.3920 | Yes |
| 64 | SLC40A1 | SLC40A1 Entrez, Source | solute carrier family 40 (iron-regulated transporter), member 1 | 870 | 0.160 | 0.3943 | Yes |
| 65 | GAS6 | GAS6 Entrez, Source | growth arrest-specific 6 | 883 | 0.158 | 0.3963 | Yes |
| 66 | GSTM3 | GSTM3 Entrez, Source | glutathione S-transferase M3 (brain) | 890 | 0.157 | 0.3988 | Yes |
| 67 | SULT1E1 | SULT1E1 Entrez, Source | sulfotransferase family 1E, estrogen-preferring, member 1 | 908 | 0.156 | 0.4003 | Yes |
| 68 | F3 | F3 Entrez, Source | coagulation factor III (thromboplastin, tissue factor) | 943 | 0.152 | 0.4005 | Yes |
| 69 | NFIA | NFIA Entrez, Source | nuclear factor I/A | 948 | 0.151 | 0.4030 | Yes |
| 70 | S100A4 | S100A4 Entrez, Source | S100 calcium binding protein A4 | 962 | 0.151 | 0.4048 | Yes |
| 71 | IFRD1 | IFRD1 Entrez, Source | interferon-related developmental regulator 1 | 999 | 0.147 | 0.4047 | Yes |
| 72 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 1041 | 0.142 | 0.4042 | Yes |
| 73 | GCNT1 | GCNT1 Entrez, Source | glucosaminyl (N-acetyl) transferase 1, core 2 (beta-1,6-N-acetylglucosaminyltransferase) | 1068 | 0.139 | 0.4048 | Yes |
| 74 | CAST | CAST Entrez, Source | calpastatin | 1090 | 0.137 | 0.4057 | Yes |
| 75 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 1114 | 0.135 | 0.4064 | Yes |
| 76 | CNTNAP1 | CNTNAP1 Entrez, Source | contactin associated protein 1 | 1143 | 0.133 | 0.4067 | Yes |
| 77 | MGAT1 | MGAT1 Entrez, Source | mannosyl (alpha-1,3-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase | 1145 | 0.133 | 0.4091 | Yes |
| 78 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 1197 | 0.128 | 0.4075 | Yes |
| 79 | LIF | LIF Entrez, Source | leukemia inhibitory factor (cholinergic differentiation factor) | 1203 | 0.127 | 0.4095 | Yes |
| 80 | PCOLCE | PCOLCE Entrez, Source | procollagen C-endopeptidase enhancer | 1214 | 0.126 | 0.4111 | Yes |
| 81 | PHLDA1 | PHLDA1 Entrez, Source | pleckstrin homology-like domain, family A, member 1 | 1221 | 0.126 | 0.4129 | Yes |
| 82 | MAPKAPK2 | MAPKAPK2 Entrez, Source | mitogen-activated protein kinase-activated protein kinase 2 | 1230 | 0.125 | 0.4147 | Yes |
| 83 | VTN | VTN Entrez, Source | vitronectin | 1239 | 0.124 | 0.4163 | Yes |
| 84 | TFPI2 | TFPI2 Entrez, Source | tissue factor pathway inhibitor 2 | 1241 | 0.124 | 0.4186 | Yes |
| 85 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 1310 | 0.119 | 0.4155 | Yes |
| 86 | EPAS1 | EPAS1 Entrez, Source | endothelial PAS domain protein 1 | 1321 | 0.118 | 0.4169 | Yes |
| 87 | NID2 | NID2 Entrez, Source | nidogen 2 (osteonidogen) | 1349 | 0.116 | 0.4170 | Yes |
| 88 | VAPB | VAPB Entrez, Source | VAMP (vesicle-associated membrane protein)-associated protein B and C | 1350 | 0.116 | 0.4192 | Yes |
| 89 | PIR | PIR Entrez, Source | pirin (iron-binding nuclear protein) | 1402 | 0.113 | 0.4173 | Yes |
| 90 | PTPN2 | PTPN2 Entrez, Source | protein tyrosine phosphatase, non-receptor type 2 | 1414 | 0.113 | 0.4185 | Yes |
| 91 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 1506 | 0.108 | 0.4135 | Yes |
| 92 | PLK2 | PLK2 Entrez, Source | polo-like kinase 2 (Drosophila) | 1523 | 0.107 | 0.4142 | Yes |
| 93 | OSBPL1A | OSBPL1A Entrez, Source | oxysterol binding protein-like 1A | 1527 | 0.107 | 0.4160 | Yes |
| 94 | SFRP1 | SFRP1 Entrez, Source | secreted frizzled-related protein 1 | 1546 | 0.105 | 0.4165 | Yes |
| 95 | GSTM1 | GSTM1 Entrez, Source | glutathione S-transferase M1 | 1557 | 0.105 | 0.4177 | Yes |
| 96 | KHDRBS3 | KHDRBS3 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 3 | 1559 | 0.105 | 0.4196 | Yes |
| 97 | NFIB | NFIB Entrez, Source | nuclear factor I/B | 1582 | 0.103 | 0.4198 | Yes |
| 98 | HRH2 | HRH2 Entrez, Source | histamine receptor H2 | 1623 | 0.101 | 0.4186 | Yes |
| 99 | ADFP | ADFP Entrez, Source | adipose differentiation-related protein | 1626 | 0.101 | 0.4203 | Yes |
| 100 | FGFR3 | FGFR3 Entrez, Source | fibroblast growth factor receptor 3 (achondroplasia, thanatophoric dwarfism) | 1667 | 0.099 | 0.4190 | No |
| 101 | DAXX | DAXX Entrez, Source | death-associated protein 6 | 1703 | 0.097 | 0.4181 | No |
| 102 | CSF1 | CSF1 Entrez, Source | colony stimulating factor 1 (macrophage) | 1723 | 0.096 | 0.4184 | No |
| 103 | RALBP1 | RALBP1 Entrez, Source | ralA binding protein 1 | 1792 | 0.093 | 0.4149 | No |
| 104 | PLAG1 | PLAG1 Entrez, Source | pleiomorphic adenoma gene 1 | 1816 | 0.092 | 0.4148 | No |
| 105 | CTSD | CTSD Entrez, Source | cathepsin D (lysosomal aspartyl peptidase) | 1839 | 0.091 | 0.4148 | No |
| 106 | GJB1 | GJB1 Entrez, Source | gap junction protein, beta 1, 32kDa (connexin 32, Charcot-Marie-Tooth neuropathy, X-linked) | 1871 | 0.090 | 0.4141 | No |
| 107 | AXIN2 | AXIN2 Entrez, Source | axin 2 (conductin, axil) | 1922 | 0.088 | 0.4118 | No |
| 108 | COL18A1 | COL18A1 Entrez, Source | collagen, type XVIII, alpha 1 | 1942 | 0.087 | 0.4119 | No |
| 109 | CTBP1 | CTBP1 Entrez, Source | C-terminal binding protein 1 | 1948 | 0.087 | 0.4132 | No |
| 110 | LCP2 | LCP2 Entrez, Source | lymphocyte cytosolic protein 2 (SH2 domain containing leukocyte protein of 76kDa) | 1958 | 0.087 | 0.4141 | No |
| 111 | CEBPD | CEBPD Entrez, Source | CCAAT/enhancer binding protein (C/EBP), delta | 1977 | 0.086 | 0.4143 | No |
| 112 | SEMA3A | SEMA3A Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A | 1982 | 0.086 | 0.4156 | No |
| 113 | ACTA2 | ACTA2 Entrez, Source | actin, alpha 2, smooth muscle, aorta | 2074 | 0.082 | 0.4100 | No |
| 114 | RABEP2 | RABEP2 Entrez, Source | rabaptin, RAB GTPase binding effector protein 2 | 2084 | 0.082 | 0.4108 | No |
| 115 | CD244 | CD244 Entrez, Source | CD244 molecule, natural killer cell receptor 2B4 | 2118 | 0.080 | 0.4098 | No |
| 116 | APOE | APOE Entrez, Source | apolipoprotein E | 2123 | 0.080 | 0.4109 | No |
| 117 | HEXB | HEXB Entrez, Source | hexosaminidase B (beta polypeptide) | 2133 | 0.080 | 0.4117 | No |
| 118 | HPN | HPN Entrez, Source | hepsin (transmembrane protease, serine 1) | 2141 | 0.079 | 0.4127 | No |
| 119 | MIPEP | MIPEP Entrez, Source | mitochondrial intermediate peptidase | 2154 | 0.079 | 0.4132 | No |
| 120 | EFNB1 | EFNB1 Entrez, Source | ephrin-B1 | 2174 | 0.078 | 0.4132 | No |
| 121 | MAPK10 | MAPK10 Entrez, Source | mitogen-activated protein kinase 10 | 2176 | 0.078 | 0.4145 | No |
| 122 | ITGAM | ITGAM Entrez, Source | integrin, alpha M (complement component 3 receptor 3 subunit) | 2219 | 0.077 | 0.4127 | No |
| 123 | WISP2 | WISP2 Entrez, Source | WNT1 inducible signaling pathway protein 2 | 2285 | 0.074 | 0.4090 | No |
| 124 | TPO | TPO Entrez, Source | thyroid peroxidase | 2311 | 0.074 | 0.4085 | No |
| 125 | BRE | BRE Entrez, Source | brain and reproductive organ-expressed (TNFRSF1A modulator) | 2350 | 0.072 | 0.4069 | No |
| 126 | MAP2K2 | MAP2K2 Entrez, Source | mitogen-activated protein kinase kinase 2 | 2358 | 0.072 | 0.4077 | No |
| 127 | LTA | LTA Entrez, Source | lymphotoxin alpha (TNF superfamily, member 1) | 2360 | 0.072 | 0.4089 | No |
| 128 | BDKRB2 | BDKRB2 Entrez, Source | bradykinin receptor B2 | 2364 | 0.072 | 0.4100 | No |
| 129 | CDK5R1 | CDK5R1 Entrez, Source | cyclin-dependent kinase 5, regulatory subunit 1 (p35) | 2373 | 0.072 | 0.4107 | No |
| 130 | ATP1B1 | ATP1B1 Entrez, Source | ATPase, Na+/K+ transporting, beta 1 polypeptide | 2395 | 0.071 | 0.4104 | No |
| 131 | SDCCAG1 | SDCCAG1 Entrez, Source | serologically defined colon cancer antigen 1 | 2404 | 0.071 | 0.4111 | No |
| 132 | TBC1D10A | TBC1D10A Entrez, Source | TBC1 domain family, member 10A | 2421 | 0.070 | 0.4112 | No |
| 133 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 2451 | 0.069 | 0.4102 | No |
| 134 | CDKN1B | CDKN1B Entrez, Source | cyclin-dependent kinase inhibitor 1B (p27, Kip1) | 2499 | 0.068 | 0.4078 | No |
| 135 | NUDT2 | NUDT2 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 2 | 2523 | 0.067 | 0.4073 | No |
| 136 | GRN | GRN Entrez, Source | granulin | 2581 | 0.065 | 0.4041 | No |
| 137 | PDGFRB | PDGFRB Entrez, Source | platelet-derived growth factor receptor, beta polypeptide | 2644 | 0.064 | 0.4005 | No |
| 138 | CDK5RAP2 | CDK5RAP2 Entrez, Source | CDK5 regulatory subunit associated protein 2 | 2664 | 0.063 | 0.4002 | No |
| 139 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 2668 | 0.063 | 0.4011 | No |
| 140 | CPD | CPD Entrez, Source | carboxypeptidase D | 2683 | 0.063 | 0.4012 | No |
| 141 | EDG2 | EDG2 Entrez, Source | endothelial differentiation, lysophosphatidic acid G-protein-coupled receptor, 2 | 2690 | 0.063 | 0.4019 | No |
| 142 | CTSH | CTSH Entrez, Source | cathepsin H | 2723 | 0.062 | 0.4005 | No |
| 143 | FKBP1B | FKBP1B Entrez, Source | FK506 binding protein 1B, 12.6 kDa | 2776 | 0.061 | 0.3976 | No |
| 144 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 2779 | 0.061 | 0.3986 | No |
| 145 | ANGPT1 | ANGPT1 Entrez, Source | angiopoietin 1 | 2782 | 0.061 | 0.3996 | No |
| 146 | INPP1 | INPP1 Entrez, Source | inositol polyphosphate-1-phosphatase | 2784 | 0.061 | 0.4006 | No |
| 147 | CCR2 | CCR2 Entrez, Source | chemokine (C-C motif) receptor 2 | 2793 | 0.060 | 0.4011 | No |
| 148 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 2806 | 0.060 | 0.4013 | No |
| 149 | AGPAT1 | AGPAT1 Entrez, Source | 1-acylglycerol-3-phosphate O-acyltransferase 1 (lysophosphatidic acid acyltransferase, alpha) | 2813 | 0.060 | 0.4020 | No |
| 150 | LITAF | LITAF Entrez, Source | lipopolysaccharide-induced TNF factor | 2826 | 0.059 | 0.4021 | No |
| 151 | IMPA2 | IMPA2 Entrez, Source | inositol(myo)-1(or 4)-monophosphatase 2 | 2844 | 0.059 | 0.4019 | No |
| 152 | CBFB | CBFB Entrez, Source | core-binding factor, beta subunit | 2867 | 0.058 | 0.4013 | No |
| 153 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 2902 | 0.057 | 0.3997 | No |
| 154 | ARL6IP5 | ARL6IP5 Entrez, Source | ADP-ribosylation-like factor 6 interacting protein 5 | 2947 | 0.056 | 0.3973 | No |
| 155 | S100A10 | S100A10 Entrez, Source | S100 calcium binding protein A10 | 2981 | 0.055 | 0.3958 | No |
| 156 | TGFBR3 | TGFBR3 Entrez, Source | transforming growth factor, beta receptor III (betaglycan, 300kDa) | 3034 | 0.054 | 0.3927 | No |
| 157 | ACO1 | ACO1 Entrez, Source | aconitase 1, soluble | 3097 | 0.052 | 0.3889 | No |
| 158 | HNMT | HNMT Entrez, Source | histamine N-methyltransferase | 3098 | 0.052 | 0.3899 | No |
| 159 | JUN | JUN Entrez, Source | jun oncogene | 3104 | 0.052 | 0.3904 | No |
| 160 | P2RXL1 | P2RXL1 Entrez, Source | purinergic receptor P2X-like 1, orphan receptor | 3146 | 0.051 | 0.3882 | No |
| 161 | BCL2L1 | BCL2L1 Entrez, Source | BCL2-like 1 | 3166 | 0.050 | 0.3876 | No |
| 162 | STOM | STOM Entrez, Source | stomatin | 3184 | 0.050 | 0.3873 | No |
| 163 | GPR30 | GPR30 Entrez, Source | G protein-coupled receptor 30 | 3198 | 0.050 | 0.3872 | No |
| 164 | SPINT2 | SPINT2 Entrez, Source | serine peptidase inhibitor, Kunitz type, 2 | 3215 | 0.049 | 0.3868 | No |
| 165 | NPC2 | NPC2 Entrez, Source | Niemann-Pick disease, type C2 | 3237 | 0.049 | 0.3861 | No |
| 166 | PSMA6 | PSMA6 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 6 | 3279 | 0.048 | 0.3838 | No |
| 167 | CDH1 | CDH1 Entrez, Source | cadherin 1, type 1, E-cadherin (epithelial) | 3296 | 0.047 | 0.3835 | No |
| 168 | ID4 | ID4 Entrez, Source | inhibitor of DNA binding 4, dominant negative helix-loop-helix protein | 3304 | 0.047 | 0.3838 | No |
| 169 | CCNA1 | CCNA1 Entrez, Source | cyclin A1 | 3333 | 0.046 | 0.3825 | No |
| 170 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 3345 | 0.046 | 0.3825 | No |
| 171 | JAG2 | JAG2 Entrez, Source | jagged 2 | 3370 | 0.046 | 0.3815 | No |
| 172 | LSR | LSR Entrez, Source | lipolysis stimulated lipoprotein receptor | 3374 | 0.046 | 0.3821 | No |
| 173 | SDC4 | SDC4 Entrez, Source | syndecan 4 (amphiglycan, ryudocan) | 3385 | 0.045 | 0.3822 | No |
| 174 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 3412 | 0.045 | 0.3810 | No |
| 175 | PDGFD | PDGFD Entrez, Source | platelet derived growth factor D | 3442 | 0.044 | 0.3795 | No |
| 176 | HMGB1 | HMGB1 Entrez, Source | high-mobility group box 1 | 3490 | 0.043 | 0.3767 | No |
| 177 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 3506 | 0.043 | 0.3763 | No |
| 178 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 3540 | 0.042 | 0.3745 | No |
| 179 | MPP3 | MPP3 Entrez, Source | membrane protein, palmitoylated 3 (MAGUK p55 subfamily member 3) | 3553 | 0.042 | 0.3744 | No |
| 180 | PDLIM3 | PDLIM3 Entrez, Source | PDZ and LIM domain 3 | 3560 | 0.042 | 0.3747 | No |
| 181 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 3563 | 0.042 | 0.3753 | No |
| 182 | MST1 | MST1 Entrez, Source | macrophage stimulating 1 (hepatocyte growth factor-like) | 3594 | 0.041 | 0.3737 | No |
| 183 | SEPP1 | SEPP1 Entrez, Source | selenoprotein P, plasma, 1 | 3625 | 0.040 | 0.3722 | No |
| 184 | TGFB3 | TGFB3 Entrez, Source | transforming growth factor, beta 3 | 3692 | 0.039 | 0.3677 | No |
| 185 | ADORA2B | ADORA2B Entrez, Source | adenosine A2b receptor | 3693 | 0.039 | 0.3685 | No |
| 186 | HOXA2 | HOXA2 Entrez, Source | homeobox A2 | 3729 | 0.038 | 0.3665 | No |
| 187 | AKAP2 | AKAP2 Entrez, Source | A kinase (PRKA) anchor protein 2 | 3738 | 0.038 | 0.3665 | No |
| 188 | SLC27A4 | SLC27A4 Entrez, Source | solute carrier family 27 (fatty acid transporter), member 4 | 3783 | 0.037 | 0.3638 | No |
| 189 | MGST1 | MGST1 Entrez, Source | microsomal glutathione S-transferase 1 | 3932 | 0.034 | 0.3529 | No |
| 190 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 3949 | 0.034 | 0.3523 | No |
| 191 | GPM6B | GPM6B Entrez, Source | glycoprotein M6B | 4058 | 0.032 | 0.3445 | No |
| 192 | POU6F1 | POU6F1 Entrez, Source | POU domain, class 6, transcription factor 1 | 4073 | 0.031 | 0.3440 | No |
| 193 | CTF1 | CTF1 Entrez, Source | cardiotrophin 1 | 4135 | 0.030 | 0.3399 | No |
| 194 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 4156 | 0.030 | 0.3389 | No |
| 195 | RAB13 | RAB13 Entrez, Source | RAB13, member RAS oncogene family | 4159 | 0.030 | 0.3393 | No |
| 196 | PAK3 | PAK3 Entrez, Source | p21 (CDKN1A)-activated kinase 3 | 4196 | 0.029 | 0.3370 | No |
| 197 | PTPRF | PTPRF Entrez, Source | protein tyrosine phosphatase, receptor type, F | 4202 | 0.029 | 0.3372 | No |
| 198 | DRD2 | DRD2 Entrez, Source | dopamine receptor D2 | 4207 | 0.029 | 0.3374 | No |
| 199 | SYNGR2 | SYNGR2 Entrez, Source | synaptogyrin 2 | 4223 | 0.029 | 0.3368 | No |
| 200 | EPHB6 | EPHB6 Entrez, Source | EPH receptor B6 | 4295 | 0.027 | 0.3318 | No |
| 201 | LAMA3 | LAMA3 Entrez, Source | laminin, alpha 3 | 4300 | 0.027 | 0.3320 | No |
| 202 | LPHN2 | LPHN2 Entrez, Source | latrophilin 2 | 4430 | 0.025 | 0.3224 | No |
| 203 | NAB2 | NAB2 Entrez, Source | NGFI-A binding protein 2 (EGR1 binding protein 2) | 4454 | 0.024 | 0.3211 | No |
| 204 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 4483 | 0.024 | 0.3193 | No |
| 205 | SDCBP | SDCBP Entrez, Source | syndecan binding protein (syntenin) | 4506 | 0.023 | 0.3181 | No |
| 206 | LUM | LUM Entrez, Source | lumican | 4533 | 0.023 | 0.3165 | No |
| 207 | CLCN7 | CLCN7 Entrez, Source | chloride channel 7 | 4535 | 0.023 | 0.3168 | No |
| 208 | STK17B | STK17B Entrez, Source | serine/threonine kinase 17b (apoptosis-inducing) | 4570 | 0.022 | 0.3146 | No |
| 209 | PTGIS | PTGIS Entrez, Source | prostaglandin I2 (prostacyclin) synthase | 4653 | 0.021 | 0.3086 | No |
| 210 | PSMC3 | PSMC3 Entrez, Source | proteasome (prosome, macropain) 26S subunit, ATPase, 3 | 4668 | 0.021 | 0.3079 | No |
| 211 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 4720 | 0.020 | 0.3043 | No |
| 212 | ADRB2 | ADRB2 Entrez, Source | adrenergic, beta-2-, receptor, surface | 4733 | 0.020 | 0.3037 | No |
| 213 | SOCS1 | SOCS1 Entrez, Source | suppressor of cytokine signaling 1 | 4762 | 0.019 | 0.3019 | No |
| 214 | FASTK | FASTK Entrez, Source | Fas-activated serine/threonine kinase | 4763 | 0.019 | 0.3023 | No |
| 215 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 4794 | 0.019 | 0.3003 | No |
| 216 | TNFRSF1A | TNFRSF1A Entrez, Source | tumor necrosis factor receptor superfamily, member 1A | 4825 | 0.018 | 0.2983 | No |
| 217 | SUFU | SUFU Entrez, Source | suppressor of fused homolog (Drosophila) | 4870 | 0.017 | 0.2952 | No |
| 218 | CCNG1 | CCNG1 Entrez, Source | cyclin G1 | 4877 | 0.017 | 0.2950 | No |
| 219 | CRYAA | CRYAA Entrez, Source | crystallin, alpha A | 4950 | 0.016 | 0.2897 | No |
| 220 | PPP1CA | PPP1CA Entrez, Source | protein phosphatase 1, catalytic subunit, alpha isoform | 5008 | 0.015 | 0.2856 | No |
| 221 | CROT | CROT Entrez, Source | carnitine O-octanoyltransferase | 5009 | 0.015 | 0.2859 | No |
| 222 | IFIT2 | IFIT2 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 2 | 5017 | 0.015 | 0.2856 | No |
| 223 | GAPDH | GAPDH Entrez, Source | glyceraldehyde-3-phosphate dehydrogenase | 5074 | 0.014 | 0.2815 | No |
| 224 | FGL2 | FGL2 Entrez, Source | fibrinogen-like 2 | 5086 | 0.014 | 0.2809 | No |
| 225 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 5124 | 0.013 | 0.2783 | No |
| 226 | MAPK7 | MAPK7 Entrez, Source | mitogen-activated protein kinase 7 | 5142 | 0.013 | 0.2772 | No |
| 227 | LCN2 | LCN2 Entrez, Source | lipocalin 2 (oncogene 24p3) | 5175 | 0.012 | 0.2749 | No |
| 228 | ARPC1A | ARPC1A Entrez, Source | actin related protein 2/3 complex, subunit 1A, 41kDa | 5358 | 0.009 | 0.2610 | No |
| 229 | CCNF | CCNF Entrez, Source | cyclin F | 5366 | 0.009 | 0.2606 | No |
| 230 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 5399 | 0.009 | 0.2583 | No |
| 231 | GALT | GALT Entrez, Source | galactose-1-phosphate uridylyltransferase | 5437 | 0.008 | 0.2555 | No |
| 232 | ST6GALNAC2 | ST6GALNAC2 Entrez, Source | ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 2 | 5440 | 0.008 | 0.2555 | No |
| 233 | CTSL | CTSL Entrez, Source | cathepsin L | 5451 | 0.008 | 0.2549 | No |
| 234 | CLDN8 | CLDN8 Entrez, Source | claudin 8 | 5544 | 0.007 | 0.2479 | No |
| 235 | CDC42 | CDC42 Entrez, Source | cell division cycle 42 (GTP binding protein, 25kDa) | 5548 | 0.007 | 0.2478 | No |
| 236 | NR2F2 | NR2F2 Entrez, Source | nuclear receptor subfamily 2, group F, member 2 | 5565 | 0.006 | 0.2467 | No |
| 237 | LDLR | LDLR Entrez, Source | low density lipoprotein receptor (familial hypercholesterolemia) | 5574 | 0.006 | 0.2461 | No |
| 238 | SCG2 | SCG2 Entrez, Source | secretogranin II (chromogranin C) | 5614 | 0.005 | 0.2432 | No |
| 239 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 5669 | 0.005 | 0.2391 | No |
| 240 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 5704 | 0.004 | 0.2366 | No |
| 241 | ENG | ENG Entrez, Source | endoglin (Osler-Rendu-Weber syndrome 1) | 5761 | 0.003 | 0.2323 | No |
| 242 | MMP2 | MMP2 Entrez, Source | matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) | 5780 | 0.003 | 0.2309 | No |
| 243 | ASNA1 | ASNA1 Entrez, Source | arsA arsenite transporter, ATP-binding, homolog 1 (bacterial) | 5871 | 0.002 | 0.2240 | No |
| 244 | WEE1 | WEE1 Entrez, Source | WEE1 homolog (S. pombe) | 5905 | 0.001 | 0.2214 | No |
| 245 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 5955 | 0.001 | 0.2176 | No |
| 246 | ACVR2A | ACVR2A Entrez, Source | activin A receptor, type IIA | 5960 | 0.001 | 0.2173 | No |
| 247 | RDH10 | RDH10 Entrez, Source | retinol dehydrogenase 10 (all-trans) | 5982 | 0.000 | 0.2157 | No |
| 248 | CDC25A | CDC25A Entrez, Source | cell division cycle 25A | 6002 | 0.000 | 0.2142 | No |
| 249 | ACSL4 | ACSL4 Entrez, Source | acyl-CoA synthetase long-chain family member 4 | 6090 | -0.001 | 0.2075 | No |
| 250 | CCNE1 | CCNE1 Entrez, Source | cyclin E1 | 6092 | -0.002 | 0.2075 | No |
| 251 | PDGFRA | PDGFRA Entrez, Source | platelet-derived growth factor receptor, alpha polypeptide | 6101 | -0.002 | 0.2069 | No |
| 252 | ARNT2 | ARNT2 Entrez, Source | aryl-hydrocarbon receptor nuclear translocator 2 | 6133 | -0.002 | 0.2045 | No |
| 253 | OXTR | OXTR Entrez, Source | oxytocin receptor | 6165 | -0.002 | 0.2021 | No |
| 254 | SQSTM1 | SQSTM1 Entrez, Source | sequestosome 1 | 6184 | -0.003 | 0.2008 | No |
| 255 | FMO2 | FMO2 Entrez, Source | flavin containing monooxygenase 2 (non-functional) | 6188 | -0.003 | 0.2006 | No |
| 256 | PSG4 | PSG4 Entrez, Source | pregnancy specific beta-1-glycoprotein 4 | 6287 | -0.004 | 0.1931 | No |
| 257 | PPIA | PPIA Entrez, Source | peptidylprolyl isomerase A (cyclophilin A) | 6293 | -0.004 | 0.1928 | No |
| 258 | COPS3 | COPS3 Entrez, Source | COP9 constitutive photomorphogenic homolog subunit 3 (Arabidopsis) | 6295 | -0.005 | 0.1928 | No |
| 259 | LMNA | LMNA Entrez, Source | lamin A/C | 6342 | -0.005 | 0.1893 | No |
| 260 | PIGC | PIGC Entrez, Source | phosphatidylinositol glycan anchor biosynthesis, class C | 6374 | -0.006 | 0.1870 | No |
| 261 | ACACB | ACACB Entrez, Source | acetyl-Coenzyme A carboxylase beta | 6400 | -0.006 | 0.1852 | No |
| 262 | RYR1 | RYR1 Entrez, Source | ryanodine receptor 1 (skeletal) | 6453 | -0.007 | 0.1813 | No |
| 263 | ABCA3 | ABCA3 Entrez, Source | ATP-binding cassette, sub-family A (ABC1), member 3 | 6472 | -0.007 | 0.1800 | No |
| 264 | SH3BP2 | SH3BP2 Entrez, Source | SH3-domain binding protein 2 | 6483 | -0.007 | 0.1794 | No |
| 265 | BRD2 | BRD2 Entrez, Source | bromodomain containing 2 | 6500 | -0.008 | 0.1783 | No |
| 266 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 6595 | -0.009 | 0.1711 | No |
| 267 | APLN | APLN Entrez, Source | apelin, AGTRL1 ligand | 6665 | -0.010 | 0.1660 | No |
| 268 | STK25 | STK25 Entrez, Source | serine/threonine kinase 25 (STE20 homolog, yeast) | 6672 | -0.010 | 0.1657 | No |
| 269 | GSTM5 | GSTM5 Entrez, Source | glutathione S-transferase M5 | 6680 | -0.010 | 0.1653 | No |
| 270 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 6682 | -0.010 | 0.1655 | No |
| 271 | SLC22A6 | SLC22A6 Entrez, Source | solute carrier family 22 (organic anion transporter), member 6 | 6692 | -0.011 | 0.1650 | No |
| 272 | HSF2 | HSF2 Entrez, Source | heat shock transcription factor 2 | 6714 | -0.011 | 0.1635 | No |
| 273 | TCF21 | TCF21 Entrez, Source | transcription factor 21 | 6717 | -0.011 | 0.1636 | No |
| 274 | BRCA1 | BRCA1 Entrez, Source | breast cancer 1, early onset | 6790 | -0.012 | 0.1582 | No |
| 275 | GRB7 | GRB7 Entrez, Source | growth factor receptor-bound protein 7 | 6828 | -0.013 | 0.1556 | No |
| 276 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 6833 | -0.013 | 0.1555 | No |
| 277 | SERPINF2 | SERPINF2 Entrez, Source | serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 2 | 6956 | -0.014 | 0.1463 | No |
| 278 | STC1 | STC1 Entrez, Source | stanniocalcin 1 | 6986 | -0.015 | 0.1443 | No |
| 279 | FAM3C | FAM3C Entrez, Source | family with sequence similarity 3, member C | 7017 | -0.015 | 0.1423 | No |
| 280 | CNKSR1 | CNKSR1 Entrez, Source | connector enhancer of kinase suppressor of Ras 1 | 7028 | -0.016 | 0.1418 | No |
| 281 | PPP2R1B | PPP2R1B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), beta isoform | 7092 | -0.017 | 0.1372 | No |
| 282 | CDH11 | CDH11 Entrez, Source | cadherin 11, type 2, OB-cadherin (osteoblast) | 7101 | -0.017 | 0.1369 | No |
| 283 | GSK3A | GSK3A Entrez, Source | glycogen synthase kinase 3 alpha | 7109 | -0.017 | 0.1367 | No |
| 284 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 7149 | -0.017 | 0.1339 | No |
| 285 | GHRL | GHRL Entrez, Source | ghrelin/obestatin preprohormone | 7162 | -0.018 | 0.1333 | No |
| 286 | CCNC | CCNC Entrez, Source | cyclin C | 7174 | -0.018 | 0.1328 | No |
| 287 | CRK | CRK Entrez, Source | v-crk sarcoma virus CT10 oncogene homolog (avian) | 7199 | -0.018 | 0.1313 | No |
| 288 | BST1 | BST1 Entrez, Source | bone marrow stromal cell antigen 1 | 7247 | -0.019 | 0.1280 | No |
| 289 | FGF6 | FGF6 Entrez, Source | fibroblast growth factor 6 | 7269 | -0.019 | 0.1267 | No |
| 290 | CDH2 | CDH2 Entrez, Source | cadherin 2, type 1, N-cadherin (neuronal) | 7290 | -0.019 | 0.1255 | No |
| 291 | SCAMP3 | SCAMP3 Entrez, Source | secretory carrier membrane protein 3 | 7292 | -0.019 | 0.1258 | No |
| 292 | ATRX | ATRX Entrez, Source | alpha thalassemia/mental retardation syndrome X-linked (RAD54 homolog, S. cerevisiae) | 7295 | -0.019 | 0.1260 | No |
| 293 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 7356 | -0.020 | 0.1217 | No |
| 294 | SFRP4 | SFRP4 Entrez, Source | secreted frizzled-related protein 4 | 7370 | -0.021 | 0.1211 | No |
| 295 | CDK2 | CDK2 Entrez, Source | cyclin-dependent kinase 2 | 7386 | -0.021 | 0.1203 | No |
| 296 | PPARG | PPARG Entrez, Source | peroxisome proliferative activated receptor, gamma | 7396 | -0.021 | 0.1200 | No |
| 297 | ANGPT2 | ANGPT2 Entrez, Source | angiopoietin 2 | 7458 | -0.022 | 0.1157 | No |
| 298 | ICOS | ICOS Entrez, Source | inducible T-cell co-stimulator | 7475 | -0.022 | 0.1149 | No |
| 299 | UNC119 | UNC119 Entrez, Source | unc-119 homolog (C. elegans) | 7505 | -0.023 | 0.1130 | No |
| 300 | SOD1 | SOD1 Entrez, Source | superoxide dismutase 1, soluble (amyotrophic lateral sclerosis 1 (adult)) | 7545 | -0.023 | 0.1104 | No |
| 301 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 7557 | -0.023 | 0.1100 | No |
| 302 | PDGFRL | PDGFRL Entrez, Source | platelet-derived growth factor receptor-like | 7568 | -0.024 | 0.1097 | No |
| 303 | MME | MME Entrez, Source | membrane metallo-endopeptidase (neutral endopeptidase, enkephalinase) | 7622 | -0.025 | 0.1060 | No |
| 304 | RHOA | RHOA Entrez, Source | ras homolog gene family, member A | 7625 | -0.025 | 0.1063 | No |
| 305 | PPP3CB | PPP3CB Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, beta isoform (calcineurin A beta) | 7629 | -0.025 | 0.1065 | No |
| 306 | PGRMC1 | PGRMC1 Entrez, Source | progesterone receptor membrane component 1 | 7638 | -0.025 | 0.1064 | No |
| 307 | PIK3R1 | PIK3R1 Entrez, Source | phosphoinositide-3-kinase, regulatory subunit 1 (p85 alpha) | 7649 | -0.025 | 0.1061 | No |
| 308 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 7656 | -0.025 | 0.1061 | No |
| 309 | EPS15 | EPS15 Entrez, Source | epidermal growth factor receptor pathway substrate 15 | 7677 | -0.026 | 0.1050 | No |
| 310 | EEF1A1 | EEF1A1 Entrez, Source | eukaryotic translation elongation factor 1 alpha 1 | 7752 | -0.027 | 0.0997 | No |
| 311 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 7811 | -0.028 | 0.0957 | No |
| 312 | C6 | C6 Entrez, Source | complement component 6 | 7841 | -0.028 | 0.0940 | No |
| 313 | ZFP161 | ZFP161 Entrez, Source | zinc finger protein 161 homolog (mouse) | 7931 | -0.030 | 0.0877 | No |
| 314 | CAMK2D | CAMK2D Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II delta | 7932 | -0.030 | 0.0882 | No |
| 315 | MVK | MVK Entrez, Source | mevalonate kinase (mevalonic aciduria) | 7967 | -0.030 | 0.0861 | No |
| 316 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 7989 | -0.031 | 0.0851 | No |
| 317 | AKR1B10 | AKR1B10 Entrez, Source | aldo-keto reductase family 1, member B10 (aldose reductase) | 8074 | -0.032 | 0.0791 | No |
| 318 | TG | TG Entrez, Source | thyroglobulin | 8094 | -0.032 | 0.0783 | No |
| 319 | CTSK | CTSK Entrez, Source | cathepsin K (pycnodysostosis) | 8117 | -0.032 | 0.0772 | No |
| 320 | EFEMP1 | EFEMP1 Entrez, Source | EGF-containing fibulin-like extracellular matrix protein 1 | 8124 | -0.033 | 0.0773 | No |
| 321 | DHPS | DHPS Entrez, Source | deoxyhypusine synthase | 8259 | -0.035 | 0.0675 | No |
| 322 | CSDE1 | CSDE1 Entrez, Source | cold shock domain containing E1, RNA-binding | 8262 | -0.035 | 0.0680 | No |
| 323 | PNOC | PNOC Entrez, Source | prepronociceptin | 8291 | -0.036 | 0.0665 | No |
| 324 | CDK4 | CDK4 Entrez, Source | cyclin-dependent kinase 4 | 8298 | -0.036 | 0.0667 | No |
| 325 | PCYT2 | PCYT2 Entrez, Source | phosphate cytidylyltransferase 2, ethanolamine | 8300 | -0.036 | 0.0673 | No |
| 326 | MAPK6 | MAPK6 Entrez, Source | mitogen-activated protein kinase 6 | 8336 | -0.036 | 0.0653 | No |
| 327 | WDR39 | WDR39 Entrez, Source | WD repeat domain 39 | 8447 | -0.038 | 0.0574 | No |
| 328 | PDE1A | PDE1A Entrez, Source | phosphodiesterase 1A, calmodulin-dependent | 8459 | -0.038 | 0.0573 | No |
| 329 | BFSP1 | BFSP1 Entrez, Source | beaded filament structural protein 1, filensin | 8509 | -0.039 | 0.0542 | No |
| 330 | SAP18 | SAP18 Entrez, Source | Sin3A-associated protein, 18kDa | 8573 | -0.040 | 0.0501 | No |
| 331 | COPA | COPA Entrez, Source | coatomer protein complex, subunit alpha | 8614 | -0.041 | 0.0477 | No |
| 332 | HNRPDL | HNRPDL Entrez, Source | heterogeneous nuclear ribonucleoprotein D-like | 8663 | -0.041 | 0.0447 | No |
| 333 | TXNIP | TXNIP Entrez, Source | thioredoxin interacting protein | 8667 | -0.041 | 0.0453 | No |
| 334 | CAMK2B | CAMK2B Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II beta | 8673 | -0.041 | 0.0457 | No |
| 335 | SLIT3 | SLIT3 Entrez, Source | slit homolog 3 (Drosophila) | 8691 | -0.042 | 0.0451 | No |
| 336 | PPP2R1A | PPP2R1A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit A (PR 65), alpha isoform | 8698 | -0.042 | 0.0454 | No |
| 337 | C1R | C1R Entrez, Source | complement component 1, r subcomponent | 8819 | -0.044 | 0.0369 | No |
| 338 | KCNMB1 | KCNMB1 Entrez, Source | potassium large conductance calcium-activated channel, subfamily M, beta member 1 | 8825 | -0.044 | 0.0374 | No |
| 339 | MBTPS1 | MBTPS1 Entrez, Source | membrane-bound transcription factor peptidase, site 1 | 8862 | -0.045 | 0.0354 | No |
| 340 | SST | SST Entrez, Source | somatostatin | 8868 | -0.045 | 0.0359 | No |
| 341 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 8927 | -0.046 | 0.0322 | No |
| 342 | DPP4 | DPP4 Entrez, Source | dipeptidyl-peptidase 4 (CD26, adenosine deaminase complexing protein 2) | 8984 | -0.047 | 0.0287 | No |
| 343 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 9017 | -0.048 | 0.0271 | No |
| 344 | SNX10 | SNX10 Entrez, Source | sorting nexin 10 | 9028 | -0.048 | 0.0272 | No |
| 345 | IL6ST | IL6ST Entrez, Source | interleukin 6 signal transducer (gp130, oncostatin M receptor) | 9035 | -0.048 | 0.0277 | No |
| 346 | UBE2D2 | UBE2D2 Entrez, Source | ubiquitin-conjugating enzyme E2D 2 (UBC4/5 homolog, yeast) | 9056 | -0.048 | 0.0270 | No |
| 347 | GALNT5 | GALNT5 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 5 (GalNAc-T5) | 9062 | -0.048 | 0.0275 | No |
| 348 | IGFBP4 | IGFBP4 Entrez, Source | insulin-like growth factor binding protein 4 | 9082 | -0.049 | 0.0269 | No |
| 349 | PCTK3 | PCTK3 Entrez, Source | PCTAIRE protein kinase 3 | 9103 | -0.049 | 0.0263 | No |
| 350 | PRODH | PRODH Entrez, Source | proline dehydrogenase (oxidase) 1 | 9179 | -0.050 | 0.0214 | No |
| 351 | GPNMB | GPNMB Entrez, Source | glycoprotein (transmembrane) nmb | 9192 | -0.050 | 0.0214 | No |
| 352 | CTSC | CTSC Entrez, Source | cathepsin C | 9429 | -0.055 | 0.0041 | No |
| 353 | SLC25A11 | SLC25A11 Entrez, Source | solute carrier family 25 (mitochondrial carrier; oxoglutarate carrier), member 11 | 9476 | -0.056 | 0.0016 | No |
| 354 | PAX8 | PAX8 Entrez, Source | paired box gene 8 | 9509 | -0.056 | 0.0001 | No |
| 355 | IFIT1 | IFIT1 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 1 | 9511 | -0.056 | 0.0011 | No |
| 356 | EPOR | EPOR Entrez, Source | erythropoietin receptor | 9512 | -0.056 | 0.0021 | No |
| 357 | ELF3 | ELF3 Entrez, Source | E74-like factor 3 (ets domain transcription factor, epithelial-specific ) | 9517 | -0.056 | 0.0029 | No |
| 358 | IFIT3 | IFIT3 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 3 | 9528 | -0.056 | 0.0031 | No |
| 359 | KCNE3 | KCNE3 Entrez, Source | potassium voltage-gated channel, Isk-related family, member 3 | 9543 | -0.057 | 0.0031 | No |
| 360 | RASSF5 | RASSF5 Entrez, Source | Ras association (RalGDS/AF-6) domain family 5 | 9569 | -0.057 | 0.0022 | No |
| 361 | RPL5 | RPL5 Entrez, Source | ribosomal protein L5 | 9572 | -0.057 | 0.0031 | No |
| 362 | SREBF1 | SREBF1 Entrez, Source | sterol regulatory element binding transcription factor 1 | 9603 | -0.058 | 0.0019 | No |
| 363 | AHR | AHR Entrez, Source | aryl hydrocarbon receptor | 9715 | -0.060 | -0.0056 | No |
| 364 | MFNG | MFNG Entrez, Source | manic fringe homolog (Drosophila) | 9855 | -0.063 | -0.0153 | No |
| 365 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 9857 | -0.063 | -0.0142 | No |
| 366 | CDC25B | CDC25B Entrez, Source | cell division cycle 25B | 9872 | -0.064 | -0.0141 | No |
| 367 | CTNNB1 | CTNNB1 Entrez, Source | catenin (cadherin-associated protein), beta 1, 88kDa | 9886 | -0.064 | -0.0139 | No |
| 368 | FGF18 | FGF18 Entrez, Source | fibroblast growth factor 18 | 9933 | -0.065 | -0.0162 | No |
| 369 | STK38 | STK38 Entrez, Source | serine/threonine kinase 38 | 10135 | -0.069 | -0.0306 | No |
| 370 | APEX1 | APEX1 Entrez, Source | APEX nuclease (multifunctional DNA repair enzyme) 1 | 10173 | -0.070 | -0.0322 | No |
| 371 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 10182 | -0.070 | -0.0315 | No |
| 372 | RAE1 | RAE1 Entrez, Source | RAE1 RNA export 1 homolog (S. pombe) | 10200 | -0.071 | -0.0315 | No |
| 373 | COLEC12 | COLEC12 Entrez, Source | collectin sub-family member 12 | 10217 | -0.071 | -0.0314 | No |
| 374 | S100A1 | S100A1 Entrez, Source | S100 calcium binding protein A1 | 10276 | -0.072 | -0.0346 | No |
| 375 | TOB1 | TOB1 Entrez, Source | transducer of ERBB2, 1 | 10294 | -0.072 | -0.0345 | No |
| 376 | SOD3 | SOD3 Entrez, Source | superoxide dismutase 3, extracellular | 10312 | -0.073 | -0.0345 | No |
| 377 | AK2 | AK2 Entrez, Source | adenylate kinase 2 | 10352 | -0.074 | -0.0362 | No |
| 378 | SRPX | SRPX Entrez, Source | sushi-repeat-containing protein, X-linked | 10360 | -0.074 | -0.0353 | No |
| 379 | IL1RN | IL1RN Entrez, Source | interleukin 1 receptor antagonist | 10398 | -0.075 | -0.0368 | No |
| 380 | AMACR | AMACR Entrez, Source | alpha-methylacyl-CoA racemase | 10410 | -0.075 | -0.0363 | No |
| 381 | OPRS1 | OPRS1 Entrez, Source | opioid receptor, sigma 1 | 10458 | -0.077 | -0.0385 | No |
| 382 | CLEC11A | CLEC11A Entrez, Source | C-type lectin domain family 11, member A | 10483 | -0.077 | -0.0389 | No |
| 383 | CDC16 | CDC16 Entrez, Source | CDC16 cell division cycle 16 homolog (S. cerevisiae) | 10523 | -0.078 | -0.0405 | No |
| 384 | HOXA1 | HOXA1 Entrez, Source | homeobox A1 | 10536 | -0.079 | -0.0400 | No |
| 385 | MYC | MYC Entrez, Source | v-myc myelocytomatosis viral oncogene homolog (avian) | 10547 | -0.079 | -0.0393 | No |
| 386 | TGFBR2 | TGFBR2 Entrez, Source | transforming growth factor, beta receptor II (70/80kDa) | 10602 | -0.080 | -0.0420 | No |
| 387 | RANGAP1 | RANGAP1 Entrez, Source | Ran GTPase activating protein 1 | 10603 | -0.080 | -0.0405 | No |
| 388 | STMN1 | STMN1 Entrez, Source | stathmin 1/oncoprotein 18 | 10649 | -0.081 | -0.0425 | No |
| 389 | ATP2B2 | ATP2B2 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 2 | 10683 | -0.082 | -0.0435 | No |
| 390 | GLMN | GLMN Entrez, Source | glomulin, FKBP associated protein | 10689 | -0.082 | -0.0424 | No |
| 391 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 10699 | -0.083 | -0.0416 | No |
| 392 | HAS2 | HAS2 Entrez, Source | hyaluronan synthase 2 | 10766 | -0.084 | -0.0451 | No |
| 393 | ICAM1 | ICAM1 Entrez, Source | intercellular adhesion molecule 1 (CD54), human rhinovirus receptor | 10770 | -0.085 | -0.0438 | No |
| 394 | CFL1 | CFL1 Entrez, Source | cofilin 1 (non-muscle) | 10892 | -0.088 | -0.0516 | No |
| 395 | PTN | PTN Entrez, Source | pleiotrophin (heparin binding growth factor 8, neurite growth-promoting factor 1) | 10905 | -0.088 | -0.0508 | No |
| 396 | CCNH | CCNH Entrez, Source | cyclin H | 10928 | -0.089 | -0.0509 | No |
| 397 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 11066 | -0.092 | -0.0598 | No |
| 398 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 11118 | -0.094 | -0.0620 | No |
| 399 | INSR | INSR Entrez, Source | insulin receptor | 11168 | -0.095 | -0.0641 | No |
| 400 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 11192 | -0.096 | -0.0641 | No |
| 401 | PUS1 | PUS1 Entrez, Source | pseudouridylate synthase 1 | 11257 | -0.097 | -0.0673 | No |
| 402 | RNH1 | RNH1 Entrez, Source | ribonuclease/angiogenin inhibitor 1 | 11262 | -0.097 | -0.0658 | No |
| 403 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 11386 | -0.102 | -0.0734 | No |
| 404 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 11395 | -0.102 | -0.0722 | No |
| 405 | IL13RA1 | IL13RA1 Entrez, Source | interleukin 13 receptor, alpha 1 | 11405 | -0.102 | -0.0709 | No |
| 406 | NINJ1 | NINJ1 Entrez, Source | ninjurin 1 | 11455 | -0.104 | -0.0728 | No |
| 407 | RPL18 | RPL18 Entrez, Source | ribosomal protein L18 | 11478 | -0.105 | -0.0726 | No |
| 408 | PSMB6 | PSMB6 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 6 | 11483 | -0.105 | -0.0709 | No |
| 409 | GUCY2F | GUCY2F Entrez, Source | guanylate cyclase 2F, retinal | 11499 | -0.106 | -0.0701 | No |
| 410 | ITGA6 | ITGA6 Entrez, Source | integrin, alpha 6 | 11513 | -0.106 | -0.0692 | No |
| 411 | AKT2 | AKT2 Entrez, Source | v-akt murine thymoma viral oncogene homolog 2 | 11527 | -0.107 | -0.0682 | No |
| 412 | RAN | RAN Entrez, Source | RAN, member RAS oncogene family | 11559 | -0.108 | -0.0686 | No |
| 413 | HDGF | HDGF Entrez, Source | hepatoma-derived growth factor (high-mobility group protein 1-like) | 11565 | -0.108 | -0.0670 | No |
| 414 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 11578 | -0.108 | -0.0659 | No |
| 415 | PRKD1 | PRKD1 Entrez, Source | protein kinase D1 | 11594 | -0.109 | -0.0650 | No |
| 416 | AREG | AREG Entrez, Source | amphiregulin (schwannoma-derived growth factor) | 11612 | -0.110 | -0.0643 | No |
| 417 | SDCCAG10 | SDCCAG10 Entrez, Source | serologically defined colon cancer antigen 10 | 11686 | -0.113 | -0.0679 | No |
| 418 | BYSL | BYSL Entrez, Source | bystin-like | 11704 | -0.113 | -0.0671 | No |
| 419 | EEF1G | EEF1G Entrez, Source | eukaryotic translation elongation factor 1 gamma | 11708 | -0.114 | -0.0652 | No |
| 420 | UNG | UNG Entrez, Source | uracil-DNA glycosylase | 11765 | -0.116 | -0.0674 | No |
| 421 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 11864 | -0.120 | -0.0728 | No |
| 422 | PLAU | PLAU Entrez, Source | plasminogen activator, urokinase | 11891 | -0.121 | -0.0725 | No |
| 423 | PPM1F | PPM1F Entrez, Source | protein phosphatase 1F (PP2C domain containing) | 11918 | -0.122 | -0.0723 | No |
| 424 | MTA1 | MTA1 Entrez, Source | metastasis associated 1 | 11924 | -0.123 | -0.0704 | No |
| 425 | SLC5A5 | SLC5A5 Entrez, Source | solute carrier family 5 (sodium iodide symporter), member 5 | 11933 | -0.123 | -0.0687 | No |
| 426 | FOSL1 | FOSL1 Entrez, Source | FOS-like antigen 1 | 11938 | -0.123 | -0.0668 | No |
| 427 | LGALS3 | LGALS3 Entrez, Source | lectin, galactoside-binding, soluble, 3 (galectin 3) | 11945 | -0.124 | -0.0649 | No |
| 428 | RPS3 | RPS3 Entrez, Source | ribosomal protein S3 | 11970 | -0.125 | -0.0645 | No |
| 429 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12025 | -0.128 | -0.0663 | No |
| 430 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 12057 | -0.129 | -0.0663 | No |
| 431 | ABO | ABO Entrez, Source | ABO blood group (transferase A, alpha 1-3-N-acetylgalactosaminyltransferase; transferase B, alpha 1-3-galactosyltransferase) | 12061 | -0.130 | -0.0641 | No |
| 432 | A2M | A2M Entrez, Source | alpha-2-macroglobulin | 12069 | -0.130 | -0.0622 | No |
| 433 | RPP38 | RPP38 Entrez, Source | ribonuclease P/MRP 38kDa subunit | 12108 | -0.132 | -0.0627 | No |
| 434 | GALR1 | GALR1 Entrez, Source | galanin receptor 1 | 12181 | -0.136 | -0.0658 | No |
| 435 | PCTK2 | PCTK2 Entrez, Source | PCTAIRE protein kinase 2 | 12183 | -0.136 | -0.0633 | No |
| 436 | TSPAN6 | TSPAN6 Entrez, Source | tetraspanin 6 | 12255 | -0.140 | -0.0663 | No |
| 437 | STOML2 | STOML2 Entrez, Source | stomatin (EPB72)-like 2 | 12394 | -0.148 | -0.0742 | No |
| 438 | RPL13A | RPL13A Entrez, Source | ribosomal protein L13a | 12527 | -0.156 | -0.0816 | No |
| 439 | RQCD1 | RQCD1 Entrez, Source | RCD1 required for cell differentiation1 homolog (S. pombe) | 12533 | -0.157 | -0.0790 | No |
| 440 | LRP4 | LRP4 Entrez, Source | low density lipoprotein receptor-related protein 4 | 12582 | -0.161 | -0.0798 | No |
| 441 | PPIC | PPIC Entrez, Source | peptidylprolyl isomerase C (cyclophilin C) | 12653 | -0.168 | -0.0821 | No |
| 442 | PDE5A | PDE5A Entrez, Source | phosphodiesterase 5A, cGMP-specific | 12668 | -0.170 | -0.0800 | No |
| 443 | PDIA5 | PDIA5 Entrez, Source | protein disulfide isomerase family A, member 5 | 12746 | -0.178 | -0.0827 | No |
| 444 | PMVK | PMVK Entrez, Source | phosphomevalonate kinase | 12750 | -0.178 | -0.0796 | No |
| 445 | MAP2K1 | MAP2K1 Entrez, Source | mitogen-activated protein kinase kinase 1 | 12766 | -0.180 | -0.0774 | No |
| 446 | TXNRD1 | TXNRD1 Entrez, Source | thioredoxin reductase 1 | 12775 | -0.181 | -0.0747 | No |
| 447 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 12777 | -0.181 | -0.0714 | No |
| 448 | TAP1 | TAP1 Entrez, Source | transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) | 12854 | -0.192 | -0.0737 | No |
| 449 | EGFR | EGFR Entrez, Source | epidermal growth factor receptor (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) | 12857 | -0.193 | -0.0703 | No |
| 450 | TUBB | TUBB Entrez, Source | tubulin, beta | 12909 | -0.201 | -0.0705 | No |
| 451 | GBP2 | GBP2 Entrez, Source | guanylate binding protein 2, interferon-inducible | 12940 | -0.207 | -0.0690 | No |
| 452 | CUL5 | CUL5 Entrez, Source | cullin 5 | 12964 | -0.212 | -0.0669 | No |
| 453 | E2F5 | E2F5 Entrez, Source | E2F transcription factor 5, p130-binding | 12993 | -0.216 | -0.0650 | No |
| 454 | PPFIBP2 | PPFIBP2 Entrez, Source | PTPRF interacting protein, binding protein 2 (liprin beta 2) | 12995 | -0.216 | -0.0611 | No |
| 455 | NRCAM | NRCAM Entrez, Source | neuronal cell adhesion molecule | 13035 | -0.227 | -0.0599 | No |
| 456 | ILF3 | ILF3 Entrez, Source | interleukin enhancer binding factor 3, 90kDa | 13043 | -0.231 | -0.0561 | No |
| 457 | FUS | FUS Entrez, Source | fusion (involved in t(12;16) in malignant liposarcoma) | 13100 | -0.252 | -0.0558 | No |
| 458 | PKLR | PKLR Entrez, Source | pyruvate kinase, liver and RBC | 13114 | -0.257 | -0.0520 | No |
| 459 | FUBP1 | FUBP1 Entrez, Source | far upstream element (FUSE) binding protein 1 | 13175 | -0.284 | -0.0514 | No |
| 460 | STAT5B | STAT5B Entrez, Source | signal transducer and activator of transcription 5B | 13179 | -0.287 | -0.0463 | No |
| 461 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 13182 | -0.289 | -0.0411 | No |
| 462 | FOXQ1 | FOXQ1 Entrez, Source | forkhead box Q1 | 13187 | -0.293 | -0.0359 | No |
| 463 | CSRP2 | CSRP2 Entrez, Source | cysteine and glycine-rich protein 2 | 13196 | -0.299 | -0.0310 | No |
| 464 | TF | TF Entrez, Source | transferrin | 13216 | -0.315 | -0.0266 | No |
| 465 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 13231 | -0.325 | -0.0216 | No |
| 466 | AOX1 | AOX1 Entrez, Source | aldehyde oxidase 1 | 13251 | -0.347 | -0.0167 | No |
| 467 | CASP4 | CASP4 Entrez, Source | caspase 4, apoptosis-related cysteine peptidase | 13300 | -0.447 | -0.0121 | No |
| 468 | EDG1 | EDG1 Entrez, Source | endothelial differentiation, sphingolipid G-protein-coupled receptor, 1 | 13334 | -0.822 | 0.0006 | No |