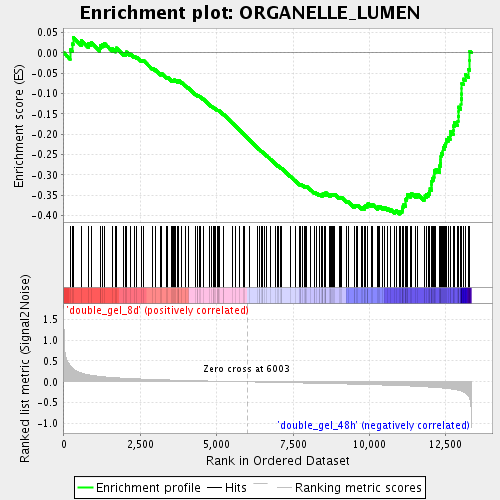
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | ORGANELLE_LUMEN |
| Enrichment Score (ES) | -0.39621153 |
| Normalized Enrichment Score (NES) | -1.7012526 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.08476545 |
| FWER p-Value | 0.998 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | NOL3 | NOL3 Entrez, Source | nucleolar protein 3 (apoptosis repressor with CARD domain) | 202 | 0.385 | 0.0078 | No |
| 2 | BCKDHA | BCKDHA Entrez, Source | branched chain keto acid dehydrogenase E1, alpha polypeptide | 272 | 0.326 | 0.0221 | No |
| 3 | HSD3B1 | HSD3B1 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 1 | 302 | 0.308 | 0.0384 | No |
| 4 | DBT | DBT Entrez, Source | dihydrolipoamide branched chain transacylase E2 | 577 | 0.210 | 0.0302 | No |
| 5 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 808 | 0.169 | 0.0228 | No |
| 6 | DNAJB9 | DNAJB9 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 9 | 886 | 0.157 | 0.0264 | No |
| 7 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1181 | 0.130 | 0.0118 | No |
| 8 | MEN1 | MEN1 Entrez, Source | multiple endocrine neoplasia I | 1194 | 0.128 | 0.0186 | No |
| 9 | BCKDHB | BCKDHB Entrez, Source | branched chain keto acid dehydrogenase E1, beta polypeptide (maple syrup urine disease) | 1252 | 0.124 | 0.0217 | No |
| 10 | EPAS1 | EPAS1 Entrez, Source | endothelial PAS domain protein 1 | 1321 | 0.118 | 0.0237 | No |
| 11 | KLHL7 | KLHL7 Entrez, Source | kelch-like 7 (Drosophila) | 1581 | 0.103 | 0.0102 | No |
| 12 | CIB1 | CIB1 Entrez, Source | calcium and integrin binding 1 (calmyrin) | 1688 | 0.098 | 0.0080 | No |
| 13 | NAP1L2 | NAP1L2 Entrez, Source | nucleosome assembly protein 1-like 2 | 1711 | 0.097 | 0.0122 | No |
| 14 | HDAC7A | HDAC7A Entrez, Source | histone deacetylase 7A | 1959 | 0.087 | -0.0014 | No |
| 15 | TFB2M | TFB2M Entrez, Source | transcription factor B2, mitochondrial | 2021 | 0.084 | -0.0010 | No |
| 16 | DEDD | DEDD Entrez, Source | death effector domain containing | 2043 | 0.083 | 0.0024 | No |
| 17 | HS3ST1 | HS3ST1 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 1 | 2172 | 0.078 | -0.0026 | No |
| 18 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 2300 | 0.074 | -0.0078 | No |
| 19 | BNIP3 | BNIP3 Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3 | 2391 | 0.071 | -0.0104 | No |
| 20 | MRPL41 | MRPL41 Entrez, Source | mitochondrial ribosomal protein L41 | 2546 | 0.066 | -0.0181 | No |
| 21 | SEP15 | SEP15 Entrez, Source | - | 2595 | 0.065 | -0.0179 | No |
| 22 | PPP1R9B | PPP1R9B Entrez, Source | protein phosphatase 1, regulatory subunit 9B, spinophilin | 2900 | 0.057 | -0.0376 | No |
| 23 | NCR1 | NCR1 Entrez, Source | natural cytotoxicity triggering receptor 1 | 2994 | 0.055 | -0.0414 | No |
| 24 | GTF3C2 | GTF3C2 Entrez, Source | general transcription factor IIIC, polypeptide 2, beta 110kDa | 3171 | 0.050 | -0.0517 | No |
| 25 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 3197 | 0.050 | -0.0507 | No |
| 26 | ELL2 | ELL2 Entrez, Source | elongation factor, RNA polymerase II, 2 | 3352 | 0.046 | -0.0596 | No |
| 27 | CDKN2A | CDKN2A Entrez, Source | cyclin-dependent kinase inhibitor 2A (melanoma, p16, inhibits CDK4) | 3401 | 0.045 | -0.0605 | No |
| 28 | NCOA6 | NCOA6 Entrez, Source | nuclear receptor coactivator 6 | 3527 | 0.042 | -0.0675 | No |
| 29 | SMARCD2 | SMARCD2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 2 | 3566 | 0.041 | -0.0679 | No |
| 30 | GRPEL1 | GRPEL1 Entrez, Source | GrpE-like 1, mitochondrial (E. coli) | 3595 | 0.041 | -0.0676 | No |
| 31 | EXOSC9 | EXOSC9 Entrez, Source | exosome component 9 | 3610 | 0.041 | -0.0662 | No |
| 32 | SP140 | SP140 Entrez, Source | SP140 nuclear body protein | 3655 | 0.040 | -0.0672 | No |
| 33 | ISG20 | ISG20 Entrez, Source | interferon stimulated exonuclease gene 20kDa | 3732 | 0.038 | -0.0707 | No |
| 34 | HDAC5 | HDAC5 Entrez, Source | histone deacetylase 5 | 3742 | 0.038 | -0.0691 | No |
| 35 | TAF9 | TAF9 Entrez, Source | TAF9 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 32kDa | 3757 | 0.037 | -0.0679 | No |
| 36 | BCKDK | BCKDK Entrez, Source | branched chain ketoacid dehydrogenase kinase | 3853 | 0.036 | -0.0730 | No |
| 37 | REXO4 | REXO4 Entrez, Source | REX4, RNA exonuclease 4 homolog (S. cerevisiae) | 3989 | 0.033 | -0.0813 | No |
| 38 | HCLS1 | HCLS1 Entrez, Source | hematopoietic cell-specific Lyn substrate 1 | 4075 | 0.031 | -0.0858 | No |
| 39 | MCRS1 | MCRS1 Entrez, Source | microspherule protein 1 | 4309 | 0.027 | -0.1019 | No |
| 40 | NARF | NARF Entrez, Source | nuclear prelamin A recognition factor | 4380 | 0.026 | -0.1057 | No |
| 41 | ETFA | ETFA Entrez, Source | electron-transfer-flavoprotein, alpha polypeptide (glutaric aciduria II) | 4385 | 0.026 | -0.1045 | No |
| 42 | SPTBN4 | SPTBN4 Entrez, Source | spectrin, beta, non-erythrocytic 4 | 4432 | 0.025 | -0.1065 | No |
| 43 | RELA | RELA Entrez, Source | v-rel reticuloendotheliosis viral oncogene homolog A, nuclear factor of kappa light polypeptide gene enhancer in B-cells 3, p65 (avian) | 4483 | 0.024 | -0.1089 | No |
| 44 | CDK9 | CDK9 Entrez, Source | cyclin-dependent kinase 9 (CDC2-related kinase) | 4558 | 0.022 | -0.1132 | No |
| 45 | NFS1 | NFS1 Entrez, Source | NFS1 nitrogen fixation 1 (S. cerevisiae) | 4759 | 0.019 | -0.1272 | No |
| 46 | NFYA | NFYA Entrez, Source | nuclear transcription factor Y, alpha | 4833 | 0.018 | -0.1317 | No |
| 47 | PTF1A | PTF1A Entrez, Source | pancreas specific transcription factor, 1a | 4886 | 0.017 | -0.1346 | No |
| 48 | CYCS | CYCS Entrez, Source | cytochrome c, somatic | 4888 | 0.017 | -0.1337 | No |
| 49 | ACADM | ACADM Entrez, Source | acyl-Coenzyme A dehydrogenase, C-4 to C-12 straight chain | 4941 | 0.016 | -0.1367 | No |
| 50 | HDAC3 | HDAC3 Entrez, Source | histone deacetylase 3 | 4967 | 0.016 | -0.1376 | No |
| 51 | HTRA2 | HTRA2 Entrez, Source | HtrA serine peptidase 2 | 5019 | 0.015 | -0.1406 | No |
| 52 | ACTB | ACTB Entrez, Source | actin, beta | 5041 | 0.014 | -0.1414 | No |
| 53 | ZBTB16 | ZBTB16 Entrez, Source | zinc finger and BTB domain containing 16 | 5042 | 0.014 | -0.1405 | No |
| 54 | NFYB | NFYB Entrez, Source | nuclear transcription factor Y, beta | 5058 | 0.014 | -0.1408 | No |
| 55 | TRPC4AP | TRPC4AP Entrez, Source | transient receptor potential cation channel, subfamily C, member 4 associated protein | 5104 | 0.013 | -0.1434 | No |
| 56 | NR3C1 | NR3C1 Entrez, Source | nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) | 5227 | 0.011 | -0.1520 | No |
| 57 | PARP1 | PARP1 Entrez, Source | poly (ADP-ribose) polymerase family, member 1 | 5231 | 0.011 | -0.1516 | No |
| 58 | HTATIP | HTATIP Entrez, Source | HIV-1 Tat interacting protein, 60kDa | 5528 | 0.007 | -0.1737 | No |
| 59 | MRPS18A | MRPS18A Entrez, Source | mitochondrial ribosomal protein S18A | 5612 | 0.005 | -0.1796 | No |
| 60 | MRPL40 | MRPL40 Entrez, Source | mitochondrial ribosomal protein L40 | 5748 | 0.003 | -0.1897 | No |
| 61 | ASNA1 | ASNA1 Entrez, Source | arsA arsenite transporter, ATP-binding, homolog 1 (bacterial) | 5871 | 0.002 | -0.1989 | No |
| 62 | ZNF148 | ZNF148 Entrez, Source | zinc finger protein 148 | 5916 | 0.001 | -0.2021 | No |
| 63 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 6078 | -0.001 | -0.2143 | No |
| 64 | TXNDC4 | TXNDC4 Entrez, Source | thioredoxin domain containing 4 (endoplasmic reticulum) | 6328 | -0.005 | -0.2329 | No |
| 65 | TBP | TBP Entrez, Source | TATA box binding protein | 6398 | -0.006 | -0.2378 | No |
| 66 | SMARCB1 | SMARCB1 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1 | 6408 | -0.006 | -0.2381 | No |
| 67 | SMARCA4 | SMARCA4 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4 | 6475 | -0.007 | -0.2427 | No |
| 68 | CHAF1B | CHAF1B Entrez, Source | chromatin assembly factor 1, subunit B (p60) | 6494 | -0.008 | -0.2436 | No |
| 69 | ETFB | ETFB Entrez, Source | electron-transfer-flavoprotein, beta polypeptide | 6510 | -0.008 | -0.2443 | No |
| 70 | NUP98 | NUP98 Entrez, Source | nucleoporin 98kDa | 6561 | -0.008 | -0.2476 | No |
| 71 | ACN9 | ACN9 Entrez, Source | ACN9 homolog (S. cerevisiae) | 6645 | -0.010 | -0.2533 | No |
| 72 | PNMA1 | PNMA1 Entrez, Source | paraneoplastic antigen MA1 | 6758 | -0.012 | -0.2611 | No |
| 73 | COIL | COIL Entrez, Source | coilin | 6923 | -0.014 | -0.2728 | No |
| 74 | PRM2 | PRM2 Entrez, Source | protamine 2 | 7003 | -0.015 | -0.2779 | No |
| 75 | NAP1L4 | NAP1L4 Entrez, Source | nucleosome assembly protein 1-like 4 | 7012 | -0.015 | -0.2776 | No |
| 76 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 7085 | -0.016 | -0.2821 | No |
| 77 | SMAD4 | SMAD4 Entrez, Source | SMAD, mothers against DPP homolog 4 (Drosophila) | 7113 | -0.017 | -0.2831 | No |
| 78 | ATP5F1 | ATP5F1 Entrez, Source | ATP synthase, H+ transporting, mitochondrial F0 complex, subunit B1 | 7135 | -0.017 | -0.2837 | No |
| 79 | NFYC | NFYC Entrez, Source | nuclear transcription factor Y, gamma | 7410 | -0.021 | -0.3032 | No |
| 80 | GATAD2A | GATAD2A Entrez, Source | GATA zinc finger domain containing 2A | 7565 | -0.024 | -0.3135 | No |
| 81 | GIT2 | GIT2 Entrez, Source | G protein-coupled receptor kinase interactor 2 | 7717 | -0.026 | -0.3235 | No |
| 82 | TAF6 | TAF6 Entrez, Source | TAF6 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 80kDa | 7744 | -0.027 | -0.3238 | No |
| 83 | HIF1A | HIF1A Entrez, Source | hypoxia-inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | 7751 | -0.027 | -0.3227 | No |
| 84 | KPNA2 | KPNA2 Entrez, Source | karyopherin alpha 2 (RAG cohort 1, importin alpha 1) | 7811 | -0.028 | -0.3255 | No |
| 85 | SUB1 | SUB1 Entrez, Source | SUB1 homolog (S. cerevisiae) | 7882 | -0.029 | -0.3291 | No |
| 86 | PPARGC1B | PPARGC1B Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, beta | 7905 | -0.029 | -0.3290 | No |
| 87 | IKBKAP | IKBKAP Entrez, Source | inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase complex-associated protein | 7916 | -0.029 | -0.3280 | No |
| 88 | SMARCD3 | SMARCD3 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3 | 7948 | -0.030 | -0.3286 | No |
| 89 | RTCD1 | RTCD1 Entrez, Source | RNA terminal phosphate cyclase domain 1 | 8083 | -0.032 | -0.3369 | No |
| 90 | PHF12 | PHF12 Entrez, Source | PHD finger protein 12 | 8193 | -0.034 | -0.3431 | No |
| 91 | TEP1 | TEP1 Entrez, Source | telomerase-associated protein 1 | 8212 | -0.034 | -0.3425 | No |
| 92 | HDAC10 | HDAC10 Entrez, Source | histone deacetylase 10 | 8278 | -0.035 | -0.3453 | No |
| 93 | PITX2 | PITX2 Entrez, Source | paired-like homeodomain transcription factor 2 | 8349 | -0.037 | -0.3484 | No |
| 94 | CPSF3 | CPSF3 Entrez, Source | cleavage and polyadenylation specific factor 3, 73kDa | 8375 | -0.037 | -0.3481 | No |
| 95 | SMAD2 | SMAD2 Entrez, Source | SMAD, mothers against DPP homolog 2 (Drosophila) | 8430 | -0.038 | -0.3499 | No |
| 96 | SC65 | SC65 Entrez, Source | - | 8463 | -0.038 | -0.3500 | No |
| 97 | NUDT21 | NUDT21 Entrez, Source | nudix (nucleoside diphosphate linked moiety X)-type motif 21 | 8466 | -0.038 | -0.3479 | No |
| 98 | OGG1 | OGG1 Entrez, Source | 8-oxoguanine DNA glycosylase | 8478 | -0.039 | -0.3464 | No |
| 99 | THRAP3 | THRAP3 Entrez, Source | thyroid hormone receptor associated protein 3 | 8530 | -0.039 | -0.3479 | No |
| 100 | TAF11 | TAF11 Entrez, Source | TAF11 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 28kDa | 8562 | -0.040 | -0.3479 | No |
| 101 | FOXE3 | FOXE3 Entrez, Source | forkhead box E3 | 8571 | -0.040 | -0.3461 | No |
| 102 | SAP18 | SAP18 Entrez, Source | Sin3A-associated protein, 18kDa | 8573 | -0.040 | -0.3438 | No |
| 103 | DDX24 | DDX24 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 24 | 8687 | -0.042 | -0.3499 | No |
| 104 | UBE2I | UBE2I Entrez, Source | ubiquitin-conjugating enzyme E2I (UBC9 homolog, yeast) | 8725 | -0.043 | -0.3501 | No |
| 105 | CHEK2 | CHEK2 Entrez, Source | CHK2 checkpoint homolog (S. pombe) | 8735 | -0.043 | -0.3482 | No |
| 106 | INTS6 | INTS6 Entrez, Source | integrator complex subunit 6 | 8753 | -0.043 | -0.3469 | No |
| 107 | DDX47 | DDX47 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 | 8805 | -0.044 | -0.3482 | No |
| 108 | SLU7 | SLU7 Entrez, Source | - | 8831 | -0.044 | -0.3474 | No |
| 109 | MBTPS1 | MBTPS1 Entrez, Source | membrane-bound transcription factor peptidase, site 1 | 8862 | -0.045 | -0.3470 | No |
| 110 | RB1 | RB1 Entrez, Source | retinoblastoma 1 (including osteosarcoma) | 9017 | -0.048 | -0.3559 | No |
| 111 | CALR | CALR Entrez, Source | calreticulin | 9052 | -0.048 | -0.3556 | No |
| 112 | MORF4L2 | MORF4L2 Entrez, Source | mortality factor 4 like 2 | 9092 | -0.049 | -0.3556 | No |
| 113 | HDAC6 | HDAC6 Entrez, Source | histone deacetylase 6 | 9263 | -0.052 | -0.3654 | No |
| 114 | NF2 | NF2 Entrez, Source | neurofibromin 2 (bilateral acoustic neuroma) | 9310 | -0.053 | -0.3657 | No |
| 115 | PAX8 | PAX8 Entrez, Source | paired box gene 8 | 9509 | -0.056 | -0.3774 | No |
| 116 | NDUFS1 | NDUFS1 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 1, 75kDa (NADH-coenzyme Q reductase) | 9515 | -0.056 | -0.3744 | No |
| 117 | PHB | PHB Entrez, Source | prohibitin | 9567 | -0.057 | -0.3749 | No |
| 118 | GCM1 | GCM1 Entrez, Source | glial cells missing homolog 1 (Drosophila) | 9613 | -0.058 | -0.3748 | No |
| 119 | RPL3 | RPL3 Entrez, Source | ribosomal protein L3 | 9736 | -0.061 | -0.3805 | No |
| 120 | XRCC6 | XRCC6 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 6 (Ku autoantigen, 70kDa) | 9780 | -0.062 | -0.3800 | No |
| 121 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 9836 | -0.063 | -0.3804 | No |
| 122 | GEMIN6 | GEMIN6 Entrez, Source | gem (nuclear organelle) associated protein 6 | 9841 | -0.063 | -0.3769 | No |
| 123 | MRPL23 | MRPL23 Entrez, Source | mitochondrial ribosomal protein L23 | 9879 | -0.064 | -0.3759 | No |
| 124 | NKX2-5 | NKX2-5 Entrez, Source | NK2 transcription factor related, locus 5 (Drosophila) | 9922 | -0.065 | -0.3752 | No |
| 125 | FGF18 | FGF18 Entrez, Source | fibroblast growth factor 18 | 9933 | -0.065 | -0.3721 | No |
| 126 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 9950 | -0.065 | -0.3694 | No |
| 127 | SMC3 | SMC3 Entrez, Source | structural maintenance of chromosomes 3 | 10069 | -0.068 | -0.3743 | No |
| 128 | PTBP1 | PTBP1 Entrez, Source | polypyrimidine tract binding protein 1 | 10104 | -0.068 | -0.3727 | No |
| 129 | SYMPK | SYMPK Entrez, Source | symplekin | 10270 | -0.072 | -0.3810 | No |
| 130 | NAP1L1 | NAP1L1 Entrez, Source | nucleosome assembly protein 1-like 1 | 10281 | -0.072 | -0.3774 | No |
| 131 | MRPL12 | MRPL12 Entrez, Source | mitochondrial ribosomal protein L12 | 10333 | -0.073 | -0.3769 | No |
| 132 | OPA1 | OPA1 Entrez, Source | optic atrophy 1 (autosomal dominant) | 10430 | -0.076 | -0.3796 | No |
| 133 | MPG | MPG Entrez, Source | N-methylpurine-DNA glycosylase | 10506 | -0.078 | -0.3807 | No |
| 134 | HES6 | HES6 Entrez, Source | hairy and enhancer of split 6 (Drosophila) | 10589 | -0.080 | -0.3821 | No |
| 135 | SFRS2 | SFRS2 Entrez, Source | splicing factor, arginine/serine-rich 2 | 10691 | -0.082 | -0.3849 | No |
| 136 | HNRPK | HNRPK Entrez, Source | heterogeneous nuclear ribonucleoprotein K | 10815 | -0.086 | -0.3891 | No |
| 137 | CLOCK | CLOCK Entrez, Source | clock homolog (mouse) | 10868 | -0.087 | -0.3878 | No |
| 138 | GTF2F1 | GTF2F1 Entrez, Source | general transcription factor IIF, polypeptide 1, 74kDa | 10980 | -0.090 | -0.3908 | Yes |
| 139 | SUPV3L1 | SUPV3L1 Entrez, Source | suppressor of var1, 3-like 1 (S. cerevisiae) | 11025 | -0.091 | -0.3887 | Yes |
| 140 | TOP2A | TOP2A Entrez, Source | topoisomerase (DNA) II alpha 170kDa | 11066 | -0.092 | -0.3862 | Yes |
| 141 | TFIP11 | TFIP11 Entrez, Source | tuftelin interacting protein 11 | 11088 | -0.093 | -0.3822 | Yes |
| 142 | RPL11 | RPL11 Entrez, Source | ribosomal protein L11 | 11095 | -0.093 | -0.3771 | Yes |
| 143 | IMP4 | IMP4 Entrez, Source | IMP4, U3 small nucleolar ribonucleoprotein, homolog (yeast) | 11124 | -0.094 | -0.3736 | Yes |
| 144 | LYAR | LYAR Entrez, Source | - | 11179 | -0.095 | -0.3719 | Yes |
| 145 | GTF3C1 | GTF3C1 Entrez, Source | general transcription factor IIIC, polypeptide 1, alpha 220kDa | 11181 | -0.095 | -0.3663 | Yes |
| 146 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 11182 | -0.095 | -0.3606 | Yes |
| 147 | RBM39 | RBM39 Entrez, Source | RNA binding motif protein 39 | 11211 | -0.096 | -0.3569 | Yes |
| 148 | DKC1 | DKC1 Entrez, Source | dyskeratosis congenita 1, dyskerin | 11229 | -0.096 | -0.3524 | Yes |
| 149 | ARID1B | ARID1B Entrez, Source | AT rich interactive domain 1B (SWI1-like) | 11241 | -0.097 | -0.3475 | Yes |
| 150 | TP53 | TP53 Entrez, Source | tumor protein p53 (Li-Fraumeni syndrome) | 11337 | -0.100 | -0.3487 | Yes |
| 151 | MED4 | MED4 Entrez, Source | mediator of RNA polymerase II transcription, subunit 4 homolog (S. cerevisiae) | 11379 | -0.101 | -0.3457 | Yes |
| 152 | RPL35 | RPL35 Entrez, Source | ribosomal protein L35 | 11505 | -0.106 | -0.3489 | Yes |
| 153 | GNL3 | GNL3 Entrez, Source | guanine nucleotide binding protein-like 3 (nucleolar) | 11580 | -0.109 | -0.3480 | Yes |
| 154 | ATXN3 | ATXN3 Entrez, Source | ataxin 3 | 11788 | -0.116 | -0.3567 | Yes |
| 155 | ACTL6A | ACTL6A Entrez, Source | actin-like 6A | 11816 | -0.118 | -0.3517 | Yes |
| 156 | FMR1 | FMR1 Entrez, Source | fragile X mental retardation 1 | 11863 | -0.120 | -0.3480 | Yes |
| 157 | RPP40 | RPP40 Entrez, Source | ribonuclease P 40kDa subunit | 11919 | -0.122 | -0.3448 | Yes |
| 158 | NAP1L3 | NAP1L3 Entrez, Source | nucleosome assembly protein 1-like 3 | 11966 | -0.125 | -0.3408 | Yes |
| 159 | DNAJA3 | DNAJA3 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 3 | 11979 | -0.125 | -0.3342 | Yes |
| 160 | TAF2 | TAF2 Entrez, Source | TAF2 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 150kDa | 12029 | -0.128 | -0.3303 | Yes |
| 161 | DNMT3A | DNMT3A Entrez, Source | DNA (cytosine-5-)-methyltransferase 3 alpha | 12031 | -0.128 | -0.3227 | Yes |
| 162 | POLR3C | POLR3C Entrez, Source | polymerase (RNA) III (DNA directed) polypeptide C (62kD) | 12038 | -0.128 | -0.3154 | Yes |
| 163 | GTF2H4 | GTF2H4 Entrez, Source | general transcription factor IIH, polypeptide 4, 52kDa | 12050 | -0.129 | -0.3085 | Yes |
| 164 | RPP38 | RPP38 Entrez, Source | ribonuclease P/MRP 38kDa subunit | 12108 | -0.132 | -0.3049 | Yes |
| 165 | RPS19 | RPS19 Entrez, Source | ribosomal protein S19 | 12113 | -0.132 | -0.2973 | Yes |
| 166 | NOC4L | NOC4L Entrez, Source | nucleolar complex associated 4 homolog (S. cerevisiae) | 12118 | -0.133 | -0.2896 | Yes |
| 167 | PDIA4 | PDIA4 Entrez, Source | protein disulfide isomerase family A, member 4 | 12175 | -0.136 | -0.2857 | Yes |
| 168 | MKI67IP | MKI67IP Entrez, Source | MKI67 (FHA domain) interacting nucleolar phosphoprotein | 12289 | -0.142 | -0.2858 | Yes |
| 169 | NUFIP1 | NUFIP1 Entrez, Source | nuclear fragile X mental retardation protein interacting protein 1 | 12295 | -0.142 | -0.2776 | Yes |
| 170 | RUVBL2 | RUVBL2 Entrez, Source | RuvB-like 2 (E. coli) | 12316 | -0.144 | -0.2705 | Yes |
| 171 | MYBBP1A | MYBBP1A Entrez, Source | MYB binding protein (P160) 1a | 12333 | -0.144 | -0.2631 | Yes |
| 172 | DDX56 | DDX56 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 56 | 12337 | -0.145 | -0.2546 | Yes |
| 173 | UBTF | UBTF Entrez, Source | upstream binding transcription factor, RNA polymerase I | 12345 | -0.145 | -0.2464 | Yes |
| 174 | TIMM44 | TIMM44 Entrez, Source | translocase of inner mitochondrial membrane 44 homolog (yeast) | 12404 | -0.149 | -0.2419 | Yes |
| 175 | NCBP1 | NCBP1 Entrez, Source | nuclear cap binding protein subunit 1, 80kDa | 12406 | -0.149 | -0.2330 | Yes |
| 176 | CIRH1A | CIRH1A Entrez, Source | cirrhosis, autosomal recessive 1A (cirhin) | 12459 | -0.152 | -0.2278 | Yes |
| 177 | NOL5A | NOL5A Entrez, Source | nucleolar protein 5A (56kDa with KKE/D repeat) | 12500 | -0.155 | -0.2216 | Yes |
| 178 | NPM1 | NPM1 Entrez, Source | nucleophosmin (nucleolar phosphoprotein B23, numatrin) | 12505 | -0.155 | -0.2126 | Yes |
| 179 | MRPS15 | MRPS15 Entrez, Source | mitochondrial ribosomal protein S15 | 12575 | -0.160 | -0.2082 | Yes |
| 180 | PPIG | PPIG Entrez, Source | peptidylprolyl isomerase G (cyclophilin G) | 12641 | -0.167 | -0.2031 | Yes |
| 181 | NUAK2 | NUAK2 Entrez, Source | NUAK family, SNF1-like kinase, 2 | 12649 | -0.168 | -0.1935 | Yes |
| 182 | CS | CS Entrez, Source | citrate synthase | 12739 | -0.177 | -0.1897 | Yes |
| 183 | PDIA5 | PDIA5 Entrez, Source | protein disulfide isomerase family A, member 5 | 12746 | -0.178 | -0.1795 | Yes |
| 184 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 12773 | -0.181 | -0.1706 | Yes |
| 185 | TDG | TDG Entrez, Source | thymine-DNA glycosylase | 12893 | -0.199 | -0.1677 | Yes |
| 186 | SLC29A2 | SLC29A2 Entrez, Source | solute carrier family 29 (nucleoside transporters), member 2 | 12900 | -0.200 | -0.1562 | Yes |
| 187 | FUSIP1 | FUSIP1 Entrez, Source | FUS interacting protein (serine/arginine-rich) 1 | 12913 | -0.201 | -0.1450 | Yes |
| 188 | NCL | NCL Entrez, Source | nucleolin | 12916 | -0.202 | -0.1330 | Yes |
| 189 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 12994 | -0.216 | -0.1258 | Yes |
| 190 | RUVBL1 | RUVBL1 Entrez, Source | RuvB-like 1 (E. coli) | 12996 | -0.217 | -0.1129 | Yes |
| 191 | NOLC1 | NOLC1 Entrez, Source | nucleolar and coiled-body phosphoprotein 1 | 13015 | -0.221 | -0.1009 | Yes |
| 192 | MCM7 | MCM7 Entrez, Source | MCM7 minichromosome maintenance deficient 7 (S. cerevisiae) | 13021 | -0.223 | -0.0879 | Yes |
| 193 | NUP54 | NUP54 Entrez, Source | nucleoporin 54kDa | 13023 | -0.224 | -0.0745 | Yes |
| 194 | APTX | APTX Entrez, Source | aprataxin | 13079 | -0.242 | -0.0642 | Yes |
| 195 | ARNTL | ARNTL Entrez, Source | aryl hydrocarbon receptor nuclear translocator-like | 13133 | -0.262 | -0.0525 | Yes |
| 196 | MRE11A | MRE11A Entrez, Source | MRE11 meiotic recombination 11 homolog A (S. cerevisiae) | 13244 | -0.339 | -0.0405 | Yes |
| 197 | TAP2 | TAP2 Entrez, Source | transporter 2, ATP-binding cassette, sub-family B (MDR/TAP) | 13278 | -0.393 | -0.0194 | Yes |
| 198 | SDAD1 | SDAD1 Entrez, Source | SDA1 domain containing 1 | 13284 | -0.402 | 0.0044 | Yes |

