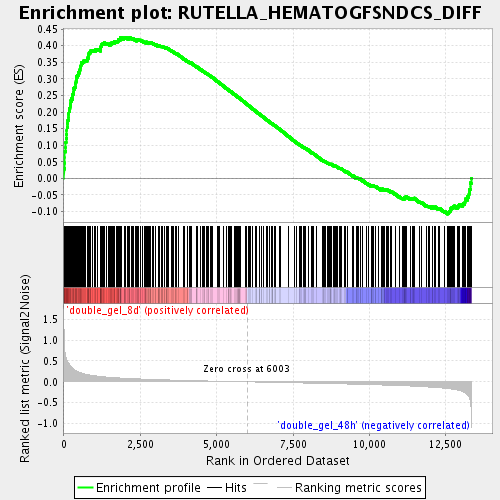
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | RUTELLA_HEMATOGFSNDCS_DIFF |
| Enrichment Score (ES) | 0.42567095 |
| Normalized Enrichment Score (NES) | 1.8138505 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.023008399 |
| FWER p-Value | 0.945 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGFBP7 | IGFBP7 Entrez, Source | insulin-like growth factor binding protein 7 | 2 | 1.254 | 0.0291 | Yes |
| 2 | DAB2 | DAB2 Entrez, Source | disabled homolog 2, mitogen-responsive phosphoprotein (Drosophila) | 18 | 0.797 | 0.0465 | Yes |
| 3 | RGS2 | RGS2 Entrez, Source | regulator of G-protein signalling 2, 24kDa | 25 | 0.771 | 0.0640 | Yes |
| 4 | TIMP2 | TIMP2 Entrez, Source | TIMP metallopeptidase inhibitor 2 | 28 | 0.756 | 0.0815 | Yes |
| 5 | SERPINE1 | SERPINE1 Entrez, Source | serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 | 56 | 0.628 | 0.0941 | Yes |
| 6 | RAMP1 | RAMP1 Entrez, Source | receptor (calcitonin) activity modifying protein 1 | 62 | 0.606 | 0.1078 | Yes |
| 7 | BHLHB3 | BHLHB3 Entrez, Source | basic helix-loop-helix domain containing, class B, 3 | 71 | 0.581 | 0.1207 | Yes |
| 8 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 89 | 0.531 | 0.1318 | Yes |
| 9 | VNN1 | VNN1 Entrez, Source | vanin 1 | 97 | 0.522 | 0.1435 | Yes |
| 10 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 101 | 0.510 | 0.1551 | Yes |
| 11 | TNFAIP6 | TNFAIP6 Entrez, Source | tumor necrosis factor, alpha-induced protein 6 | 116 | 0.483 | 0.1653 | Yes |
| 12 | PVRL2 | PVRL2 Entrez, Source | poliovirus receptor-related 2 (herpesvirus entry mediator B) | 127 | 0.468 | 0.1755 | Yes |
| 13 | CCL2 | CCL2 Entrez, Source | chemokine (C-C motif) ligand 2 | 145 | 0.452 | 0.1847 | Yes |
| 14 | APOC1 | APOC1 Entrez, Source | apolipoprotein C-I | 151 | 0.445 | 0.1947 | Yes |
| 15 | MARCKSL1 | MARCKSL1 Entrez, Source | MARCKS-like 1 | 166 | 0.428 | 0.2036 | Yes |
| 16 | ABCG1 | ABCG1 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 1 | 187 | 0.400 | 0.2114 | Yes |
| 17 | SGK | SGK Entrez, Source | serum/glucocorticoid regulated kinase | 200 | 0.388 | 0.2195 | Yes |
| 18 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 222 | 0.366 | 0.2264 | Yes |
| 19 | CFD | CFD Entrez, Source | complement factor D (adipsin) | 228 | 0.365 | 0.2345 | Yes |
| 20 | IFITM2 | IFITM2 Entrez, Source | interferon induced transmembrane protein 2 (1-8D) | 254 | 0.339 | 0.2405 | Yes |
| 21 | CD9 | CD9 Entrez, Source | CD9 molecule | 283 | 0.320 | 0.2458 | Yes |
| 22 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 288 | 0.316 | 0.2529 | Yes |
| 23 | CLIC2 | CLIC2 Entrez, Source | chloride intracellular channel 2 | 304 | 0.304 | 0.2588 | Yes |
| 24 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 317 | 0.293 | 0.2648 | Yes |
| 25 | NME3 | NME3 Entrez, Source | non-metastatic cells 3, protein expressed in | 318 | 0.293 | 0.2716 | Yes |
| 26 | CD48 | CD48 Entrez, Source | CD48 molecule | 352 | 0.281 | 0.2756 | Yes |
| 27 | PLSCR1 | PLSCR1 Entrez, Source | phospholipid scramblase 1 | 376 | 0.270 | 0.2801 | Yes |
| 28 | DECR1 | DECR1 Entrez, Source | 2,4-dienoyl CoA reductase 1, mitochondrial | 378 | 0.269 | 0.2863 | Yes |
| 29 | CYP27A1 | CYP27A1 Entrez, Source | cytochrome P450, family 27, subfamily A, polypeptide 1 | 394 | 0.263 | 0.2913 | Yes |
| 30 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 404 | 0.258 | 0.2966 | Yes |
| 31 | CCNL1 | CCNL1 Entrez, Source | cyclin L1 | 405 | 0.257 | 0.3026 | Yes |
| 32 | TPST1 | TPST1 Entrez, Source | tyrosylprotein sulfotransferase 1 | 418 | 0.254 | 0.3076 | Yes |
| 33 | HERPUD1 | HERPUD1 Entrez, Source | homocysteine-inducible, endoplasmic reticulum stress-inducible, ubiquitin-like domain member 1 | 448 | 0.245 | 0.3111 | Yes |
| 34 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 470 | 0.238 | 0.3150 | Yes |
| 35 | DUSP5 | DUSP5 Entrez, Source | dual specificity phosphatase 5 | 472 | 0.237 | 0.3204 | Yes |
| 36 | MGLL | MGLL Entrez, Source | monoglyceride lipase | 510 | 0.224 | 0.3228 | Yes |
| 37 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 523 | 0.222 | 0.3271 | Yes |
| 38 | SPRED2 | SPRED2 Entrez, Source | sprouty-related, EVH1 domain containing 2 | 528 | 0.221 | 0.3319 | Yes |
| 39 | CD59 | CD59 Entrez, Source | CD59 molecule, complement regulatory protein | 533 | 0.219 | 0.3367 | Yes |
| 40 | BCAT1 | BCAT1 Entrez, Source | branched chain aminotransferase 1, cytosolic | 558 | 0.215 | 0.3399 | Yes |
| 41 | TSPAN3 | TSPAN3 Entrez, Source | tetraspanin 3 | 563 | 0.213 | 0.3446 | Yes |
| 42 | CCR1 | CCR1 Entrez, Source | chemokine (C-C motif) receptor 1 | 564 | 0.213 | 0.3495 | Yes |
| 43 | PHYH | PHYH Entrez, Source | phytanoyl-CoA 2-hydroxylase | 614 | 0.204 | 0.3505 | Yes |
| 44 | TNK2 | TNK2 Entrez, Source | tyrosine kinase, non-receptor, 2 | 629 | 0.201 | 0.3541 | Yes |
| 45 | LMBR1L | LMBR1L Entrez, Source | limb region 1 homolog (mouse)-like | 671 | 0.191 | 0.3554 | Yes |
| 46 | FBP1 | FBP1 Entrez, Source | fructose-1,6-bisphosphatase 1 | 720 | 0.182 | 0.3559 | Yes |
| 47 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 756 | 0.176 | 0.3574 | Yes |
| 48 | CREBL2 | CREBL2 Entrez, Source | cAMP responsive element binding protein-like 2 | 764 | 0.175 | 0.3609 | Yes |
| 49 | LPXN | LPXN Entrez, Source | leupaxin | 785 | 0.172 | 0.3634 | Yes |
| 50 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 798 | 0.170 | 0.3664 | Yes |
| 51 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 809 | 0.168 | 0.3696 | Yes |
| 52 | ITPR1 | ITPR1 Entrez, Source | inositol 1,4,5-triphosphate receptor, type 1 | 816 | 0.167 | 0.3730 | Yes |
| 53 | CYB5A | CYB5A Entrez, Source | cytochrome b5 type A (microsomal) | 817 | 0.167 | 0.3769 | Yes |
| 54 | IL10 | IL10 Entrez, Source | interleukin 10 | 842 | 0.163 | 0.3789 | Yes |
| 55 | SLC16A6 | SLC16A6 Entrez, Source | solute carrier family 16, member 6 (monocarboxylic acid transporter 7) | 855 | 0.162 | 0.3817 | Yes |
| 56 | TMEM34 | TMEM34 Entrez, Source | transmembrane protein 34 | 875 | 0.159 | 0.3840 | Yes |
| 57 | GAS6 | GAS6 Entrez, Source | growth arrest-specific 6 | 883 | 0.158 | 0.3871 | Yes |
| 58 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 934 | 0.153 | 0.3868 | Yes |
| 59 | KLF9 | KLF9 Entrez, Source | Kruppel-like factor 9 | 998 | 0.147 | 0.3854 | Yes |
| 60 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 1041 | 0.142 | 0.3855 | Yes |
| 61 | ST8SIA1 | ST8SIA1 Entrez, Source | ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 1 | 1042 | 0.142 | 0.3888 | Yes |
| 62 | ACSL3 | ACSL3 Entrez, Source | acyl-CoA synthetase long-chain family member 3 | 1086 | 0.137 | 0.3887 | Yes |
| 63 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 1114 | 0.135 | 0.3897 | Yes |
| 64 | ANXA5 | ANXA5 Entrez, Source | annexin A5 | 1191 | 0.129 | 0.3869 | Yes |
| 65 | IL1B | IL1B Entrez, Source | interleukin 1, beta | 1197 | 0.128 | 0.3895 | Yes |
| 66 | RAB9 | RAB9 Entrez, Source | RAB9, member RAS oncogene family | 1201 | 0.128 | 0.3922 | Yes |
| 67 | F2RL1 | F2RL1 Entrez, Source | coagulation factor II (thrombin) receptor-like 1 | 1205 | 0.127 | 0.3950 | Yes |
| 68 | GADD45B | GADD45B Entrez, Source | growth arrest and DNA-damage-inducible, beta | 1210 | 0.127 | 0.3976 | Yes |
| 69 | C1QA | C1QA Entrez, Source | complement component 1, q subcomponent, A chain | 1220 | 0.126 | 0.3999 | Yes |
| 70 | CHPT1 | CHPT1 Entrez, Source | choline phosphotransferase 1 | 1231 | 0.125 | 0.4020 | Yes |
| 71 | PSCD1 | PSCD1 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 1(cytohesin 1) | 1237 | 0.125 | 0.4045 | Yes |
| 72 | ZFP36 | ZFP36 Entrez, Source | zinc finger protein 36, C3H type, homolog (mouse) | 1260 | 0.123 | 0.4057 | Yes |
| 73 | YWHAQ | YWHAQ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide | 1294 | 0.120 | 0.4059 | Yes |
| 74 | ASAH1 | ASAH1 Entrez, Source | N-acylsphingosine amidohydrolase (acid ceramidase) 1 | 1310 | 0.119 | 0.4076 | Yes |
| 75 | FAH | FAH Entrez, Source | fumarylacetoacetate hydrolase (fumarylacetoacetase) | 1315 | 0.119 | 0.4100 | Yes |
| 76 | PIR | PIR Entrez, Source | pirin (iron-binding nuclear protein) | 1402 | 0.113 | 0.4060 | Yes |
| 77 | SP110 | SP110 Entrez, Source | SP110 nuclear body protein | 1446 | 0.111 | 0.4053 | Yes |
| 78 | DUSP6 | DUSP6 Entrez, Source | dual specificity phosphatase 6 | 1504 | 0.108 | 0.4034 | Yes |
| 79 | RNASE4 | RNASE4 Entrez, Source | ribonuclease, RNase A family, 4 | 1525 | 0.107 | 0.4044 | Yes |
| 80 | OSBPL1A | OSBPL1A Entrez, Source | oxysterol binding protein-like 1A | 1527 | 0.107 | 0.4068 | Yes |
| 81 | TMEM23 | TMEM23 Entrez, Source | transmembrane protein 23 | 1555 | 0.105 | 0.4072 | Yes |
| 82 | PELI1 | PELI1 Entrez, Source | pellino homolog 1 (Drosophila) | 1568 | 0.104 | 0.4087 | Yes |
| 83 | HADHA | HADHA Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase/3-ketoacyl-Coenzyme A thiolase/enoyl-Coenzyme A hydratase (trifunctional protein), alpha subunit | 1594 | 0.103 | 0.4092 | Yes |
| 84 | ADFP | ADFP Entrez, Source | adipose differentiation-related protein | 1626 | 0.101 | 0.4091 | Yes |
| 85 | TGM2 | TGM2 Entrez, Source | transglutaminase 2 (C polypeptide, protein-glutamine-gamma-glutamyltransferase) | 1650 | 0.100 | 0.4097 | Yes |
| 86 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 1657 | 0.100 | 0.4115 | Yes |
| 87 | SH3BP5 | SH3BP5 Entrez, Source | SH3-domain binding protein 5 (BTK-associated) | 1659 | 0.100 | 0.4138 | Yes |
| 88 | HSD11B1 | HSD11B1 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 1 | 1720 | 0.096 | 0.4114 | Yes |
| 89 | LAMP2 | LAMP2 Entrez, Source | lysosomal-associated membrane protein 2 | 1744 | 0.095 | 0.4119 | Yes |
| 90 | NAT1 | NAT1 Entrez, Source | N-acetyltransferase 1 (arylamine N-acetyltransferase) | 1745 | 0.095 | 0.4141 | Yes |
| 91 | SPP1 | SPP1 Entrez, Source | secreted phosphoprotein 1 (osteopontin, bone sialoprotein I, early T-lymphocyte activation 1) | 1776 | 0.094 | 0.4140 | Yes |
| 92 | ADRBK2 | ADRBK2 Entrez, Source | adrenergic, beta, receptor kinase 2 | 1778 | 0.094 | 0.4161 | Yes |
| 93 | OPTN | OPTN Entrez, Source | optineurin | 1779 | 0.094 | 0.4183 | Yes |
| 94 | SPHK1 | SPHK1 Entrez, Source | sphingosine kinase 1 | 1821 | 0.092 | 0.4173 | Yes |
| 95 | CTSD | CTSD Entrez, Source | cathepsin D (lysosomal aspartyl peptidase) | 1839 | 0.091 | 0.4181 | Yes |
| 96 | TLE4 | TLE4 Entrez, Source | transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) | 1841 | 0.091 | 0.4201 | Yes |
| 97 | HBEGF | HBEGF Entrez, Source | heparin-binding EGF-like growth factor | 1849 | 0.091 | 0.4217 | Yes |
| 98 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 1850 | 0.091 | 0.4238 | Yes |
| 99 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 1854 | 0.090 | 0.4257 | Yes |
| 100 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 1887 | 0.089 | 0.4253 | No |
| 101 | ELL3 | ELL3 Entrez, Source | elongation factor RNA polymerase II-like 3 | 1966 | 0.086 | 0.4213 | No |
| 102 | CEBPD | CEBPD Entrez, Source | CCAAT/enhancer binding protein (C/EBP), delta | 1977 | 0.086 | 0.4225 | No |
| 103 | WASL | WASL Entrez, Source | Wiskott-Aldrich syndrome-like | 1998 | 0.085 | 0.4230 | No |
| 104 | ST6GAL1 | ST6GAL1 Entrez, Source | ST6 beta-galactosamide alpha-2,6-sialyltranferase 1 | 2013 | 0.085 | 0.4238 | No |
| 105 | PTGER2 | PTGER2 Entrez, Source | prostaglandin E receptor 2 (subtype EP2), 53kDa | 2027 | 0.084 | 0.4248 | No |
| 106 | G6PC3 | G6PC3 Entrez, Source | glucose 6 phosphatase, catalytic, 3 | 2090 | 0.081 | 0.4219 | No |
| 107 | APOE | APOE Entrez, Source | apolipoprotein E | 2123 | 0.080 | 0.4213 | No |
| 108 | HEXB | HEXB Entrez, Source | hexosaminidase B (beta polypeptide) | 2133 | 0.080 | 0.4225 | No |
| 109 | WNT5B | WNT5B Entrez, Source | wingless-type MMTV integration site family, member 5B | 2142 | 0.079 | 0.4237 | No |
| 110 | GNA12 | GNA12 Entrez, Source | guanine nucleotide binding protein (G protein) alpha 12 | 2156 | 0.079 | 0.4245 | No |
| 111 | FCER1A | FCER1A Entrez, Source | Fc fragment of IgE, high affinity I, receptor for; alpha polypeptide | 2214 | 0.077 | 0.4219 | No |
| 112 | LTA4H | LTA4H Entrez, Source | leukotriene A4 hydrolase | 2256 | 0.075 | 0.4205 | No |
| 113 | TRADD | TRADD Entrez, Source | TNFRSF1A-associated via death domain | 2264 | 0.075 | 0.4218 | No |
| 114 | MYD88 | MYD88 Entrez, Source | myeloid differentiation primary response gene (88) | 2332 | 0.073 | 0.4183 | No |
| 115 | ATP6V1H | ATP6V1H Entrez, Source | ATPase, H+ transporting, lysosomal 50/57kDa, V1 subunit H | 2389 | 0.071 | 0.4156 | No |
| 116 | MPZL1 | MPZL1 Entrez, Source | myelin protein zero-like 1 | 2397 | 0.071 | 0.4167 | No |
| 117 | MSRB2 | MSRB2 Entrez, Source | methionine sulfoxide reductase B2 | 2405 | 0.071 | 0.4179 | No |
| 118 | NDUFS2 | NDUFS2 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 2, 49kDa (NADH-coenzyme Q reductase) | 2448 | 0.069 | 0.4162 | No |
| 119 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 2451 | 0.069 | 0.4177 | No |
| 120 | CRYZ | CRYZ Entrez, Source | crystallin, zeta (quinone reductase) | 2492 | 0.068 | 0.4162 | No |
| 121 | SEPT9 | SEPT9 Entrez, Source | septin 9 | 2498 | 0.068 | 0.4174 | No |
| 122 | NIT2 | NIT2 Entrez, Source | nitrilase family, member 2 | 2557 | 0.066 | 0.4145 | No |
| 123 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 2629 | 0.064 | 0.4105 | No |
| 124 | HK2 | HK2 Entrez, Source | hexokinase 2 | 2661 | 0.063 | 0.4096 | No |
| 125 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 2668 | 0.063 | 0.4106 | No |
| 126 | CPD | CPD Entrez, Source | carboxypeptidase D | 2683 | 0.063 | 0.4110 | No |
| 127 | RERE | RERE Entrez, Source | arginine-glutamic acid dipeptide (RE) repeats | 2695 | 0.063 | 0.4116 | No |
| 128 | C3AR1 | C3AR1 Entrez, Source | complement component 3a receptor 1 | 2746 | 0.061 | 0.4092 | No |
| 129 | ITGAL | ITGAL Entrez, Source | integrin, alpha L (antigen CD11A (p180), lymphocyte function-associated antigen 1; alpha polypeptide) | 2766 | 0.061 | 0.4091 | No |
| 130 | FKBP1B | FKBP1B Entrez, Source | FK506 binding protein 1B, 12.6 kDa | 2776 | 0.061 | 0.4099 | No |
| 131 | BIRC3 | BIRC3 Entrez, Source | baculoviral IAP repeat-containing 3 | 2803 | 0.060 | 0.4092 | No |
| 132 | LAMP1 | LAMP1 Entrez, Source | lysosomal-associated membrane protein 1 | 2821 | 0.060 | 0.4093 | No |
| 133 | LITAF | LITAF Entrez, Source | lipopolysaccharide-induced TNF factor | 2826 | 0.059 | 0.4104 | No |
| 134 | IMPA2 | IMPA2 Entrez, Source | inositol(myo)-1(or 4)-monophosphatase 2 | 2844 | 0.059 | 0.4105 | No |
| 135 | ME1 | ME1 Entrez, Source | malic enzyme 1, NADP(+)-dependent, cytosolic | 2902 | 0.057 | 0.4074 | No |
| 136 | ARL6IP5 | ARL6IP5 Entrez, Source | ADP-ribosylation-like factor 6 interacting protein 5 | 2947 | 0.056 | 0.4053 | No |
| 137 | CTBS | CTBS Entrez, Source | chitobiase, di-N-acetyl- | 2990 | 0.055 | 0.4034 | No |
| 138 | EREG | EREG Entrez, Source | epiregulin | 3011 | 0.054 | 0.4031 | No |
| 139 | SQLE | SQLE Entrez, Source | squalene epoxidase | 3091 | 0.052 | 0.3982 | No |
| 140 | ACO1 | ACO1 Entrez, Source | aconitase 1, soluble | 3097 | 0.052 | 0.3990 | No |
| 141 | LTBP2 | LTBP2 Entrez, Source | latent transforming growth factor beta binding protein 2 | 3107 | 0.052 | 0.3995 | No |
| 142 | MAFB | MAFB Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog B (avian) | 3122 | 0.051 | 0.3997 | No |
| 143 | PVR | PVR Entrez, Source | poliovirus receptor | 3141 | 0.051 | 0.3995 | No |
| 144 | STOM | STOM Entrez, Source | stomatin | 3184 | 0.050 | 0.3974 | No |
| 145 | SCAMP2 | SCAMP2 Entrez, Source | secretory carrier membrane protein 2 | 3199 | 0.049 | 0.3975 | No |
| 146 | SPINT2 | SPINT2 Entrez, Source | serine peptidase inhibitor, Kunitz type, 2 | 3215 | 0.049 | 0.3975 | No |
| 147 | PPAP2A | PPAP2A Entrez, Source | phosphatidic acid phosphatase type 2A | 3216 | 0.049 | 0.3986 | No |
| 148 | MMP12 | MMP12 Entrez, Source | matrix metallopeptidase 12 (macrophage elastase) | 3285 | 0.048 | 0.3945 | No |
| 149 | CASP3 | CASP3 Entrez, Source | caspase 3, apoptosis-related cysteine peptidase | 3300 | 0.047 | 0.3945 | No |
| 150 | TNFRSF4 | TNFRSF4 Entrez, Source | tumor necrosis factor receptor superfamily, member 4 | 3342 | 0.046 | 0.3924 | No |
| 151 | IL18 | IL18 Entrez, Source | interleukin 18 (interferon-gamma-inducing factor) | 3347 | 0.046 | 0.3932 | No |
| 152 | TSEN34 | TSEN34 Entrez, Source | tRNA splicing endonuclease 34 homolog (S. cerevisiae) | 3348 | 0.046 | 0.3943 | No |
| 153 | SDC4 | SDC4 Entrez, Source | syndecan 4 (amphiglycan, ryudocan) | 3385 | 0.045 | 0.3925 | No |
| 154 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 3417 | 0.045 | 0.3912 | No |
| 155 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 3506 | 0.043 | 0.3854 | No |
| 156 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 3540 | 0.042 | 0.3839 | No |
| 157 | GRPEL1 | GRPEL1 Entrez, Source | GrpE-like 1, mitochondrial (E. coli) | 3595 | 0.041 | 0.3806 | No |
| 158 | GRSF1 | GRSF1 Entrez, Source | G-rich RNA sequence binding factor 1 | 3661 | 0.040 | 0.3766 | No |
| 159 | FYN | FYN Entrez, Source | FYN oncogene related to SRC, FGR, YES | 3662 | 0.040 | 0.3775 | No |
| 160 | CD47 | CD47 Entrez, Source | CD47 molecule | 3700 | 0.039 | 0.3755 | No |
| 161 | LEPROT | LEPROT Entrez, Source | leptin receptor overlapping transcript | 3751 | 0.038 | 0.3726 | No |
| 162 | DAD1 | DAD1 Entrez, Source | defender against cell death 1 | 3918 | 0.034 | 0.3606 | No |
| 163 | ANXA2 | ANXA2 Entrez, Source | annexin A2 | 3949 | 0.034 | 0.3590 | No |
| 164 | TIA1 | TIA1 Entrez, Source | TIA1 cytotoxic granule-associated RNA binding protein | 4046 | 0.032 | 0.3524 | No |
| 165 | ACP5 | ACP5 Entrez, Source | acid phosphatase 5, tartrate resistant | 4056 | 0.032 | 0.3524 | No |
| 166 | RGC32 | RGC32 Entrez, Source | - | 4104 | 0.031 | 0.3495 | No |
| 167 | MDH1 | MDH1 Entrez, Source | malate dehydrogenase 1, NAD (soluble) | 4125 | 0.030 | 0.3487 | No |
| 168 | APLP2 | APLP2 Entrez, Source | amyloid beta (A4) precursor-like protein 2 | 4130 | 0.030 | 0.3491 | No |
| 169 | SLC12A8 | SLC12A8 Entrez, Source | solute carrier family 12 (potassium/chloride transporters), member 8 | 4151 | 0.030 | 0.3482 | No |
| 170 | RAB13 | RAB13 Entrez, Source | RAB13, member RAS oncogene family | 4159 | 0.030 | 0.3484 | No |
| 171 | NQO1 | NQO1 Entrez, Source | NAD(P)H dehydrogenase, quinone 1 | 4179 | 0.030 | 0.3476 | No |
| 172 | PPIF | PPIF Entrez, Source | peptidylprolyl isomerase F (cyclophilin F) | 4337 | 0.027 | 0.3361 | No |
| 173 | CSNK1E | CSNK1E Entrez, Source | casein kinase 1, epsilon | 4346 | 0.026 | 0.3361 | No |
| 174 | MCL1 | MCL1 Entrez, Source | myeloid cell leukemia sequence 1 (BCL2-related) | 4349 | 0.026 | 0.3366 | No |
| 175 | NARF | NARF Entrez, Source | nuclear prelamin A recognition factor | 4380 | 0.026 | 0.3349 | No |
| 176 | ACADVL | ACADVL Entrez, Source | acyl-Coenzyme A dehydrogenase, very long chain | 4485 | 0.024 | 0.3274 | No |
| 177 | NRG1 | NRG1 Entrez, Source | neuregulin 1 | 4536 | 0.023 | 0.3241 | No |
| 178 | STK17B | STK17B Entrez, Source | serine/threonine kinase 17b (apoptosis-inducing) | 4570 | 0.022 | 0.3221 | No |
| 179 | SLC1A2 | SLC1A2 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 2 | 4596 | 0.022 | 0.3206 | No |
| 180 | VDR | VDR Entrez, Source | vitamin D (1,25- dihydroxyvitamin D3) receptor | 4613 | 0.021 | 0.3199 | No |
| 181 | ALOX5 | ALOX5 Entrez, Source | arachidonate 5-lipoxygenase | 4658 | 0.021 | 0.3170 | No |
| 182 | SERPINF1 | SERPINF1 Entrez, Source | serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 | 4697 | 0.020 | 0.3145 | No |
| 183 | SMAD3 | SMAD3 Entrez, Source | SMAD, mothers against DPP homolog 3 (Drosophila) | 4720 | 0.020 | 0.3133 | No |
| 184 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 4730 | 0.020 | 0.3131 | No |
| 185 | DYNC1LI1 | DYNC1LI1 Entrez, Source | dynein, cytoplasmic 1, light intermediate chain 1 | 4807 | 0.018 | 0.3076 | No |
| 186 | PLEKHA5 | PLEKHA5 Entrez, Source | pleckstrin homology domain containing, family A member 5 | 4824 | 0.018 | 0.3068 | No |
| 187 | TFEB | TFEB Entrez, Source | transcription factor EB | 4873 | 0.017 | 0.3035 | No |
| 188 | NET1 | NET1 Entrez, Source | neuroepithelial cell transforming gene 1 | 5025 | 0.015 | 0.2922 | No |
| 189 | ALAS1 | ALAS1 Entrez, Source | aminolevulinate, delta-, synthase 1 | 5059 | 0.014 | 0.2900 | No |
| 190 | INHBA | INHBA Entrez, Source | inhibin, beta A (activin A, activin AB alpha polypeptide) | 5081 | 0.014 | 0.2887 | No |
| 191 | BET1 | BET1 Entrez, Source | BET1 homolog (S. cerevisiae) | 5206 | 0.012 | 0.2794 | No |
| 192 | SON | SON Entrez, Source | SON DNA binding protein | 5330 | 0.010 | 0.2702 | No |
| 193 | LYPD3 | LYPD3 Entrez, Source | LY6/PLAUR domain containing 3 | 5376 | 0.009 | 0.2669 | No |
| 194 | TRIP10 | TRIP10 Entrez, Source | thyroid hormone receptor interactor 10 | 5381 | 0.009 | 0.2668 | No |
| 195 | CCL17 | CCL17 Entrez, Source | chemokine (C-C motif) ligand 17 | 5384 | 0.009 | 0.2669 | No |
| 196 | CACNA2D3 | CACNA2D3 Entrez, Source | calcium channel, voltage-dependent, alpha 2/delta 3 subunit | 5434 | 0.008 | 0.2633 | No |
| 197 | ICAM2 | ICAM2 Entrez, Source | intercellular adhesion molecule 2 | 5448 | 0.008 | 0.2625 | No |
| 198 | CTSL | CTSL Entrez, Source | cathepsin L | 5451 | 0.008 | 0.2625 | No |
| 199 | CD80 | CD80 Entrez, Source | CD80 molecule | 5469 | 0.008 | 0.2614 | No |
| 200 | LDLR | LDLR Entrez, Source | low density lipoprotein receptor (familial hypercholesterolemia) | 5574 | 0.006 | 0.2535 | No |
| 201 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 5599 | 0.006 | 0.2518 | No |
| 202 | NKG7 | NKG7 Entrez, Source | natural killer cell group 7 sequence | 5655 | 0.005 | 0.2477 | No |
| 203 | STAT1 | STAT1 Entrez, Source | signal transducer and activator of transcription 1, 91kDa | 5682 | 0.005 | 0.2458 | No |
| 204 | C5AR1 | C5AR1 Entrez, Source | complement component 5a receptor 1 | 5691 | 0.004 | 0.2453 | No |
| 205 | SIGIRR | SIGIRR Entrez, Source | single immunoglobulin and toll-interleukin 1 receptor (TIR) domain | 5702 | 0.004 | 0.2446 | No |
| 206 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 5704 | 0.004 | 0.2446 | No |
| 207 | AKT3 | AKT3 Entrez, Source | v-akt murine thymoma viral oncogene homolog 3 (protein kinase B, gamma) | 5706 | 0.004 | 0.2446 | No |
| 208 | SLC36A1 | SLC36A1 Entrez, Source | solute carrier family 36 (proton/amino acid symporter), member 1 | 5733 | 0.004 | 0.2427 | No |
| 209 | ENSA | ENSA Entrez, Source | endosulfine alpha | 5749 | 0.003 | 0.2416 | No |
| 210 | MAOA | MAOA Entrez, Source | monoamine oxidase A | 5751 | 0.003 | 0.2416 | No |
| 211 | MMP2 | MMP2 Entrez, Source | matrix metallopeptidase 2 (gelatinase A, 72kDa gelatinase, 72kDa type IV collagenase) | 5780 | 0.003 | 0.2395 | No |
| 212 | GLS | GLS Entrez, Source | glutaminase | 5935 | 0.001 | 0.2277 | No |
| 213 | PKNOX1 | PKNOX1 Entrez, Source | PBX/knotted 1 homeobox 1 | 5978 | 0.000 | 0.2245 | No |
| 214 | CXCR4 | CXCR4 Entrez, Source | chemokine (C-X-C motif) receptor 4 | 5981 | 0.000 | 0.2243 | No |
| 215 | CD81 | CD81 Entrez, Source | CD81 molecule | 6049 | -0.001 | 0.2192 | No |
| 216 | CCL22 | CCL22 Entrez, Source | chemokine (C-C motif) ligand 22 | 6089 | -0.001 | 0.2162 | No |
| 217 | SLC1A4 | SLC1A4 Entrez, Source | solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 | 6096 | -0.002 | 0.2158 | No |
| 218 | SQSTM1 | SQSTM1 Entrez, Source | sequestosome 1 | 6184 | -0.003 | 0.2091 | No |
| 219 | ADAM19 | ADAM19 Entrez, Source | ADAM metallopeptidase domain 19 (meltrin beta) | 6281 | -0.004 | 0.2018 | No |
| 220 | MATK | MATK Entrez, Source | megakaryocyte-associated tyrosine kinase | 6298 | -0.005 | 0.2007 | No |
| 221 | CD86 | CD86 Entrez, Source | CD86 molecule | 6302 | -0.005 | 0.2006 | No |
| 222 | SYF2 | SYF2 Entrez, Source | SYF2 homolog, RNA splicing factor (S. cerevisiae) | 6387 | -0.006 | 0.1943 | No |
| 223 | TFG | TFG Entrez, Source | TRK-fused gene | 6388 | -0.006 | 0.1944 | No |
| 224 | SPOCK1 | SPOCK1 Entrez, Source | sparc/osteonectin, cwcv and kazal-like domains proteoglycan (testican) 1 | 6415 | -0.006 | 0.1926 | No |
| 225 | SCN11A | SCN11A Entrez, Source | sodium channel, voltage-gated, type XI, alpha | 6451 | -0.007 | 0.1900 | No |
| 226 | SEC22B | SEC22B Entrez, Source | SEC22 vesicle trafficking protein homolog B (S. cerevisiae) | 6468 | -0.007 | 0.1889 | No |
| 227 | ACPP | ACPP Entrez, Source | acid phosphatase, prostate | 6543 | -0.008 | 0.1834 | No |
| 228 | NINJ2 | NINJ2 Entrez, Source | ninjurin 2 | 6616 | -0.009 | 0.1781 | No |
| 229 | SLC35B1 | SLC35B1 Entrez, Source | solute carrier family 35, member B1 | 6663 | -0.010 | 0.1748 | No |
| 230 | DNAJB6 | DNAJB6 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 6 | 6729 | -0.011 | 0.1700 | No |
| 231 | PHACTR1 | PHACTR1 Entrez, Source | phosphatase and actin regulator 1 | 6730 | -0.011 | 0.1703 | No |
| 232 | ATP6V0A1 | ATP6V0A1 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a1 | 6799 | -0.012 | 0.1653 | No |
| 233 | SEPHS1 | SEPHS1 Entrez, Source | selenophosphate synthetase 1 | 6820 | -0.012 | 0.1641 | No |
| 234 | BCAP31 | BCAP31 Entrez, Source | B-cell receptor-associated protein 31 | 6822 | -0.012 | 0.1643 | No |
| 235 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 6833 | -0.013 | 0.1638 | No |
| 236 | CMKLR1 | CMKLR1 Entrez, Source | chemokine-like receptor 1 | 6876 | -0.013 | 0.1609 | No |
| 237 | PLOD3 | PLOD3 Entrez, Source | procollagen-lysine, 2-oxoglutarate 5-dioxygenase 3 | 6920 | -0.014 | 0.1579 | No |
| 238 | FTH1 | FTH1 Entrez, Source | ferritin, heavy polypeptide 1 | 6933 | -0.014 | 0.1573 | No |
| 239 | QPRT | QPRT Entrez, Source | quinolinate phosphoribosyltransferase (nicotinate-nucleotide pyrophosphorylase (carboxylating)) | 7045 | -0.016 | 0.1491 | No |
| 240 | CEP350 | CEP350 Entrez, Source | centrosomal protein 350kDa | 7098 | -0.017 | 0.1455 | No |
| 241 | CIRBP | CIRBP Entrez, Source | cold inducible RNA binding protein | 7346 | -0.020 | 0.1269 | No |
| 242 | IL1A | IL1A Entrez, Source | interleukin 1, alpha | 7552 | -0.023 | 0.1117 | No |
| 243 | RAB38 | RAB38 Entrez, Source | RAB38, member RAS oncogene family | 7621 | -0.025 | 0.1070 | No |
| 244 | PPP1R12A | PPP1R12A Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 12A | 7696 | -0.026 | 0.1019 | No |
| 245 | DPEP2 | DPEP2 Entrez, Source | dipeptidase 2 | 7712 | -0.026 | 0.1014 | No |
| 246 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 7755 | -0.027 | 0.0988 | No |
| 247 | AIF1 | AIF1 Entrez, Source | allograft inflammatory factor 1 | 7788 | -0.027 | 0.0969 | No |
| 248 | TMEM97 | TMEM97 Entrez, Source | transmembrane protein 97 | 7845 | -0.028 | 0.0933 | No |
| 249 | MTX2 | MTX2 Entrez, Source | metaxin 2 | 7869 | -0.028 | 0.0921 | No |
| 250 | LXN | LXN Entrez, Source | latexin | 7879 | -0.029 | 0.0921 | No |
| 251 | NEDD9 | NEDD9 Entrez, Source | neural precursor cell expressed, developmentally down-regulated 9 | 7899 | -0.029 | 0.0913 | No |
| 252 | CX3CR1 | CX3CR1 Entrez, Source | chemokine (C-X3-C motif) receptor 1 | 7909 | -0.029 | 0.0913 | No |
| 253 | CYP1B1 | CYP1B1 Entrez, Source | cytochrome P450, family 1, subfamily B, polypeptide 1 | 7989 | -0.031 | 0.0860 | No |
| 254 | CTSK | CTSK Entrez, Source | cathepsin K (pycnodysostosis) | 8117 | -0.032 | 0.0769 | No |
| 255 | PTGDS | PTGDS Entrez, Source | prostaglandin D2 synthase 21kDa (brain) | 8146 | -0.033 | 0.0755 | No |
| 256 | CSRP1 | CSRP1 Entrez, Source | cysteine and glycine-rich protein 1 | 8162 | -0.033 | 0.0752 | No |
| 257 | CYBB | CYBB Entrez, Source | cytochrome b-245, beta polypeptide (chronic granulomatous disease) | 8263 | -0.035 | 0.0683 | No |
| 258 | TTN | TTN Entrez, Source | titin | 8450 | -0.038 | 0.0548 | No |
| 259 | GOT2 | GOT2 Entrez, Source | glutamic-oxaloacetic transaminase 2, mitochondrial (aspartate aminotransferase 2) | 8497 | -0.039 | 0.0522 | No |
| 260 | TNIP2 | TNIP2 Entrez, Source | TNFAIP3 interacting protein 2 | 8505 | -0.039 | 0.0525 | No |
| 261 | IER5 | IER5 Entrez, Source | immediate early response 5 | 8527 | -0.039 | 0.0518 | No |
| 262 | TCIRG1 | TCIRG1 Entrez, Source | T-cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 subunit A3 | 8560 | -0.040 | 0.0503 | No |
| 263 | SLC11A2 | SLC11A2 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2 | 8634 | -0.041 | 0.0456 | No |
| 264 | ANP32A | ANP32A Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member A | 8654 | -0.041 | 0.0451 | No |
| 265 | CD14 | CD14 Entrez, Source | CD14 molecule | 8706 | -0.042 | 0.0422 | No |
| 266 | LMO2 | LMO2 Entrez, Source | LIM domain only 2 (rhombotin-like 1) | 8708 | -0.042 | 0.0431 | No |
| 267 | ST3GAL6 | ST3GAL6 Entrez, Source | ST3 beta-galactoside alpha-2,3-sialyltransferase 6 | 8734 | -0.043 | 0.0422 | No |
| 268 | RAB8B | RAB8B Entrez, Source | RAB8B, member RAS oncogene family | 8738 | -0.043 | 0.0429 | No |
| 269 | LTB4R | LTB4R Entrez, Source | leukotriene B4 receptor | 8747 | -0.043 | 0.0433 | No |
| 270 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 8829 | -0.044 | 0.0381 | No |
| 271 | NR4A2 | NR4A2 Entrez, Source | nuclear receptor subfamily 4, group A, member 2 | 8840 | -0.044 | 0.0384 | No |
| 272 | INDO | INDO Entrez, Source | indoleamine-pyrrole 2,3 dioxygenase | 8856 | -0.045 | 0.0383 | No |
| 273 | REPIN1 | REPIN1 Entrez, Source | replication initiator 1 | 8879 | -0.045 | 0.0376 | No |
| 274 | SDC2 | SDC2 Entrez, Source | syndecan 2 (heparan sulfate proteoglycan 1, cell surface-associated, fibroglycan) | 8914 | -0.046 | 0.0361 | No |
| 275 | MARCKS | MARCKS Entrez, Source | myristoylated alanine-rich protein kinase C substrate | 8955 | -0.047 | 0.0341 | No |
| 276 | SNX10 | SNX10 Entrez, Source | sorting nexin 10 | 9028 | -0.048 | 0.0296 | No |
| 277 | IL6ST | IL6ST Entrez, Source | interleukin 6 signal transducer (gp130, oncostatin M receptor) | 9035 | -0.048 | 0.0303 | No |
| 278 | SCCPDH | SCCPDH Entrez, Source | saccharopine dehydrogenase (putative) | 9045 | -0.048 | 0.0307 | No |
| 279 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 9075 | -0.048 | 0.0296 | No |
| 280 | GPNMB | GPNMB Entrez, Source | glycoprotein (transmembrane) nmb | 9192 | -0.050 | 0.0218 | No |
| 281 | ATP1B3 | ATP1B3 Entrez, Source | ATPase, Na+/K+ transporting, beta 3 polypeptide | 9230 | -0.051 | 0.0202 | No |
| 282 | FNBP1 | FNBP1 Entrez, Source | formin binding protein 1 | 9270 | -0.052 | 0.0184 | No |
| 283 | PSEN2 | PSEN2 Entrez, Source | presenilin 2 (Alzheimer disease 4) | 9275 | -0.052 | 0.0193 | No |
| 284 | NR1H3 | NR1H3 Entrez, Source | nuclear receptor subfamily 1, group H, member 3 | 9459 | -0.055 | 0.0065 | No |
| 285 | LAMP3 | LAMP3 Entrez, Source | lysosomal-associated membrane protein 3 | 9461 | -0.055 | 0.0077 | No |
| 286 | RPL5 | RPL5 Entrez, Source | ribosomal protein L5 | 9572 | -0.057 | 0.0006 | No |
| 287 | TEGT | TEGT Entrez, Source | testis enhanced gene transcript (BAX inhibitor 1) | 9576 | -0.057 | 0.0017 | No |
| 288 | PDCD6IP | PDCD6IP Entrez, Source | programmed cell death 6 interacting protein | 9600 | -0.058 | 0.0012 | No |
| 289 | NDE1 | NDE1 Entrez, Source | nudE nuclear distribution gene E homolog 1 (A. nidulans) | 9639 | -0.058 | -0.0003 | No |
| 290 | CSPG2 | CSPG2 Entrez, Source | chondroitin sulfate proteoglycan 2 (versican) | 9690 | -0.060 | -0.0028 | No |
| 291 | SELL | SELL Entrez, Source | selectin L (lymphocyte adhesion molecule 1) | 9693 | -0.060 | -0.0016 | No |
| 292 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 9785 | -0.062 | -0.0071 | No |
| 293 | PSMB5 | PSMB5 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 5 | 9898 | -0.064 | -0.0143 | No |
| 294 | SC5DL | SC5DL Entrez, Source | sterol-C5-desaturase (ERG3 delta-5-desaturase homolog, fungal)-like | 9978 | -0.066 | -0.0188 | No |
| 295 | HSPB1 | HSPB1 Entrez, Source | heat shock 27kDa protein 1 | 10057 | -0.068 | -0.0233 | No |
| 296 | CD63 | CD63 Entrez, Source | CD63 molecule | 10073 | -0.068 | -0.0228 | No |
| 297 | KCNMA1 | KCNMA1 Entrez, Source | potassium large conductance calcium-activated channel, subfamily M, alpha member 1 | 10077 | -0.068 | -0.0215 | No |
| 298 | CD52 | CD52 Entrez, Source | CD52 molecule | 10093 | -0.068 | -0.0210 | No |
| 299 | RPL28 | RPL28 Entrez, Source | ribosomal protein L28 | 10134 | -0.069 | -0.0225 | No |
| 300 | CASP1 | CASP1 Entrez, Source | caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) | 10182 | -0.070 | -0.0245 | No |
| 301 | SLC1A3 | SLC1A3 Entrez, Source | solute carrier family 1 (glial high affinity glutamate transporter), member 3 | 10194 | -0.070 | -0.0237 | No |
| 302 | PRDX1 | PRDX1 Entrez, Source | peroxiredoxin 1 | 10293 | -0.072 | -0.0296 | No |
| 303 | RDX | RDX Entrez, Source | radixin | 10377 | -0.074 | -0.0342 | No |
| 304 | RDH11 | RDH11 Entrez, Source | retinol dehydrogenase 11 (all-trans/9-cis/11-cis) | 10402 | -0.075 | -0.0344 | No |
| 305 | PACSIN2 | PACSIN2 Entrez, Source | protein kinase C and casein kinase substrate in neurons 2 | 10406 | -0.075 | -0.0328 | No |
| 306 | TXNDC13 | TXNDC13 Entrez, Source | thioredoxin domain containing 13 | 10415 | -0.075 | -0.0317 | No |
| 307 | NARS | NARS Entrez, Source | asparaginyl-tRNA synthetase | 10470 | -0.077 | -0.0341 | No |
| 308 | CYP27B1 | CYP27B1 Entrez, Source | cytochrome P450, family 27, subfamily B, polypeptide 1 | 10490 | -0.077 | -0.0337 | No |
| 309 | SMS | SMS Entrez, Source | spermine synthase | 10501 | -0.078 | -0.0327 | No |
| 310 | FKBP4 | FKBP4 Entrez, Source | FK506 binding protein 4, 59kDa | 10541 | -0.079 | -0.0339 | No |
| 311 | JAK1 | JAK1 Entrez, Source | Janus kinase 1 (a protein tyrosine kinase) | 10575 | -0.079 | -0.0345 | No |
| 312 | DHRS4 | DHRS4 Entrez, Source | dehydrogenase/reductase (SDR family) member 4 | 10624 | -0.080 | -0.0364 | No |
| 313 | TEX2 | TEX2 Entrez, Source | testis expressed sequence 2 | 10702 | -0.083 | -0.0404 | No |
| 314 | ATP6V1C1 | ATP6V1C1 Entrez, Source | ATPase, H+ transporting, lysosomal 42kDa, V1 subunit C1 | 10713 | -0.083 | -0.0392 | No |
| 315 | CTSB | CTSB Entrez, Source | cathepsin B | 10843 | -0.086 | -0.0471 | No |
| 316 | FDX1 | FDX1 Entrez, Source | ferredoxin 1 | 10969 | -0.090 | -0.0547 | No |
| 317 | GPD2 | GPD2 Entrez, Source | glycerol-3-phosphate dehydrogenase 2 (mitochondrial) | 11070 | -0.092 | -0.0602 | No |
| 318 | EIF2B3 | EIF2B3 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 3 gamma, 58kDa | 11098 | -0.093 | -0.0601 | No |
| 319 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 11137 | -0.094 | -0.0609 | No |
| 320 | DHCR7 | DHCR7 Entrez, Source | 7-dehydrocholesterol reductase | 11141 | -0.094 | -0.0589 | No |
| 321 | SLC29A3 | SLC29A3 Entrez, Source | solute carrier family 29 (nucleoside transporters), member 3 | 11153 | -0.095 | -0.0576 | No |
| 322 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 11182 | -0.095 | -0.0575 | No |
| 323 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 11192 | -0.096 | -0.0560 | No |
| 324 | SLC31A1 | SLC31A1 Entrez, Source | solute carrier family 31 (copper transporters), member 1 | 11210 | -0.096 | -0.0550 | No |
| 325 | RUNX1 | RUNX1 Entrez, Source | runt-related transcription factor 1 (acute myeloid leukemia 1; aml1 oncogene) | 11329 | -0.100 | -0.0618 | No |
| 326 | LMNB1 | LMNB1 Entrez, Source | lamin B1 | 11330 | -0.100 | -0.0595 | No |
| 327 | IL13RA1 | IL13RA1 Entrez, Source | interleukin 13 receptor, alpha 1 | 11405 | -0.102 | -0.0628 | No |
| 328 | TSC22D1 | TSC22D1 Entrez, Source | TSC22 domain family, member 1 | 11438 | -0.104 | -0.0628 | No |
| 329 | GFPT1 | GFPT1 Entrez, Source | glutamine-fructose-6-phosphate transaminase 1 | 11452 | -0.104 | -0.0614 | No |
| 330 | GTF3A | GTF3A Entrez, Source | general transcription factor IIIA | 11457 | -0.104 | -0.0593 | No |
| 331 | DDIT4 | DDIT4 Entrez, Source | DNA-damage-inducible transcript 4 | 11645 | -0.111 | -0.0711 | No |
| 332 | SOD2 | SOD2 Entrez, Source | superoxide dismutase 2, mitochondrial | 11710 | -0.114 | -0.0734 | No |
| 333 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 11864 | -0.120 | -0.0824 | No |
| 334 | PPM1F | PPM1F Entrez, Source | protein phosphatase 1F (PP2C domain containing) | 11918 | -0.122 | -0.0836 | No |
| 335 | RPL22 | RPL22 Entrez, Source | ribosomal protein L22 | 11961 | -0.125 | -0.0840 | No |
| 336 | GM2A | GM2A Entrez, Source | GM2 ganglioside activator | 12057 | -0.129 | -0.0883 | No |
| 337 | EIF2S1 | EIF2S1 Entrez, Source | eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa | 12063 | -0.130 | -0.0856 | No |
| 338 | ROBO1 | ROBO1 Entrez, Source | roundabout, axon guidance receptor, homolog 1 (Drosophila) | 12116 | -0.133 | -0.0865 | No |
| 339 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 12150 | -0.134 | -0.0860 | No |
| 340 | GCLM | GCLM Entrez, Source | glutamate-cysteine ligase, modifier subunit | 12248 | -0.139 | -0.0902 | No |
| 341 | VAT1 | VAT1 Entrez, Source | vesicle amine transport protein 1 homolog (T californica) | 12302 | -0.143 | -0.0909 | No |
| 342 | KLF10 | KLF10 Entrez, Source | Kruppel-like factor 10 | 12453 | -0.151 | -0.0990 | No |
| 343 | C1ORF24 | C1ORF24 Entrez, Source | chromosome 1 open reading frame 24 | 12569 | -0.160 | -0.1041 | No |
| 344 | RNF138 | RNF138 Entrez, Source | ring finger protein 138 | 12593 | -0.163 | -0.1021 | No |
| 345 | TM9SF4 | TM9SF4 Entrez, Source | transmembrane 9 superfamily protein member 4 | 12607 | -0.164 | -0.0993 | No |
| 346 | ZDHHC18 | ZDHHC18 Entrez, Source | zinc finger, DHHC-type containing 18 | 12635 | -0.167 | -0.0975 | No |
| 347 | CD68 | CD68 Entrez, Source | CD68 molecule | 12657 | -0.169 | -0.0952 | No |
| 348 | SGPL1 | SGPL1 Entrez, Source | sphingosine-1-phosphate lyase 1 | 12660 | -0.169 | -0.0914 | No |
| 349 | CDS2 | CDS2 Entrez, Source | CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 2 | 12681 | -0.171 | -0.0889 | No |
| 350 | AQP9 | AQP9 Entrez, Source | aquaporin 9 | 12727 | -0.176 | -0.0883 | No |
| 351 | PDIA5 | PDIA5 Entrez, Source | protein disulfide isomerase family A, member 5 | 12746 | -0.178 | -0.0855 | No |
| 352 | NOTCH2 | NOTCH2 Entrez, Source | Notch homolog 2 (Drosophila) | 12769 | -0.180 | -0.0830 | No |
| 353 | MX2 | MX2 Entrez, Source | myxovirus (influenza virus) resistance 2 (mouse) | 12876 | -0.196 | -0.0866 | No |
| 354 | GGH | GGH Entrez, Source | gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) | 12902 | -0.200 | -0.0839 | No |
| 355 | RARRES1 | RARRES1 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 1 | 12923 | -0.203 | -0.0807 | No |
| 356 | ABCC3 | ABCC3 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 3 | 12946 | -0.209 | -0.0775 | No |
| 357 | SCARB2 | SCARB2 Entrez, Source | scavenger receptor class B, member 2 | 13055 | -0.235 | -0.0804 | No |
| 358 | SLC7A7 | SLC7A7 Entrez, Source | solute carrier family 7 (cationic amino acid transporter, y+ system), member 7 | 13061 | -0.237 | -0.0752 | No |
| 359 | GSR | GSR Entrez, Source | glutathione reductase | 13122 | -0.259 | -0.0738 | No |
| 360 | HOMER2 | HOMER2 Entrez, Source | homer homolog 2 (Drosophila) | 13128 | -0.260 | -0.0681 | No |
| 361 | SLC30A1 | SLC30A1 Entrez, Source | solute carrier family 30 (zinc transporter), member 1 | 13129 | -0.260 | -0.0621 | No |
| 362 | TFRC | TFRC Entrez, Source | transferrin receptor (p90, CD71) | 13197 | -0.299 | -0.0603 | No |
| 363 | SFPQ | SFPQ Entrez, Source | splicing factor proline/glutamine-rich (polypyrimidine tract binding protein associated) | 13213 | -0.310 | -0.0542 | No |
| 364 | SLC3A2 | SLC3A2 Entrez, Source | solute carrier family 3 (activators of dibasic and neutral amino acid transport), member 2 | 13255 | -0.353 | -0.0491 | No |
| 365 | KMO | KMO Entrez, Source | kynurenine 3-monooxygenase (kynurenine 3-hydroxylase) | 13261 | -0.360 | -0.0411 | No |
| 366 | CES1 | CES1 Entrez, Source | carboxylesterase 1 (monocyte/macrophage serine esterase 1) | 13281 | -0.399 | -0.0333 | No |
| 367 | CXCL10 | CXCL10 Entrez, Source | chemokine (C-X-C motif) ligand 10 | 13301 | -0.450 | -0.0242 | No |
| 368 | GADD45G | GADD45G Entrez, Source | growth arrest and DNA-damage-inducible, gamma | 13302 | -0.454 | -0.0136 | No |
| 369 | CXCL9 | CXCL9 Entrez, Source | chemokine (C-X-C motif) ligand 9 | 13330 | -0.714 | 0.0009 | No |

