Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
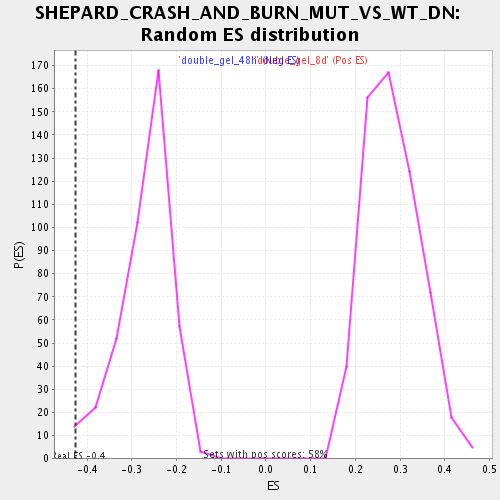
| GeneSet | SHEPARD_CRASH_AND_BURN_MUT_VS_WT_DN |
| Enrichment Score (ES) | -0.42517915 |
| Normalized Enrichment Score (NES) | -1.5762393 |
| Nominal p-value | 0.011961723 |
| FDR q-value | 0.1609706 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 222 | 0.366 | 0.0274 | No |
| 2 | UCP2 | UCP2 Entrez, Source | uncoupling protein 2 (mitochondrial, proton carrier) | 398 | 0.260 | 0.0456 | No |
| 3 | DPYSL3 | DPYSL3 Entrez, Source | dihydropyrimidinase-like 3 | 917 | 0.154 | 0.0252 | No |
| 4 | AKAP9 | AKAP9 Entrez, Source | A kinase (PRKA) anchor protein (yotiao) 9 | 1346 | 0.117 | 0.0069 | No |
| 5 | RTKN | RTKN Entrez, Source | rhotekin | 1496 | 0.108 | 0.0088 | No |
| 6 | SFRP1 | SFRP1 Entrez, Source | secreted frizzled-related protein 1 | 1546 | 0.105 | 0.0178 | No |
| 7 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 1773 | 0.094 | 0.0121 | No |
| 8 | LGMN | LGMN Entrez, Source | legumain | 1864 | 0.090 | 0.0161 | No |
| 9 | POU3F3 | POU3F3 Entrez, Source | POU domain, class 3, transcription factor 3 | 2053 | 0.083 | 0.0120 | No |
| 10 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 2173 | 0.078 | 0.0124 | No |
| 11 | NELL2 | NELL2 Entrez, Source | NEL-like 2 (chicken) | 2324 | 0.073 | 0.0099 | No |
| 12 | TACC3 | TACC3 Entrez, Source | transforming, acidic coiled-coil containing protein 3 | 2708 | 0.062 | -0.0114 | No |
| 13 | NR2F1 | NR2F1 Entrez, Source | nuclear receptor subfamily 2, group F, member 1 | 2825 | 0.059 | -0.0130 | No |
| 14 | GOLGB1 | GOLGB1 Entrez, Source | golgi autoantigen, golgin subfamily b, macrogolgin (with transmembrane signal), 1 | 3576 | 0.041 | -0.0646 | No |
| 15 | RHO | RHO Entrez, Source | rhodopsin (opsin 2, rod pigment) (retinitis pigmentosa 4, autosomal dominant) | 3754 | 0.037 | -0.0734 | No |
| 16 | MYH8 | MYH8 Entrez, Source | myosin, heavy chain 8, skeletal muscle, perinatal | 3786 | 0.037 | -0.0713 | No |
| 17 | PTPRO | PTPRO Entrez, Source | protein tyrosine phosphatase, receptor type, O | 3944 | 0.034 | -0.0790 | No |
| 18 | ATP1A1 | ATP1A1 Entrez, Source | ATPase, Na+/K+ transporting, alpha 1 polypeptide | 4072 | 0.031 | -0.0848 | No |
| 19 | NEUROG1 | NEUROG1 Entrez, Source | neurogenin 1 | 4377 | 0.026 | -0.1046 | No |
| 20 | OTP | OTP Entrez, Source | orthopedia homolog (Drosophila) | 4482 | 0.024 | -0.1096 | No |
| 21 | POU3F1 | POU3F1 Entrez, Source | POU domain, class 3, transcription factor 1 | 4491 | 0.024 | -0.1074 | No |
| 22 | GDF10 | GDF10 Entrez, Source | growth differentiation factor 10 | 4564 | 0.022 | -0.1101 | No |
| 23 | SMTN | SMTN Entrez, Source | smoothelin | 4588 | 0.022 | -0.1092 | No |
| 24 | BUB1B | BUB1B Entrez, Source | BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) | 4680 | 0.020 | -0.1136 | No |
| 25 | GRIK3 | GRIK3 Entrez, Source | glutamate receptor, ionotropic, kainate 3 | 5046 | 0.014 | -0.1394 | No |
| 26 | HMGN3 | HMGN3 Entrez, Source | high mobility group nucleosomal binding domain 3 | 5116 | 0.013 | -0.1431 | No |
| 27 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 5399 | 0.009 | -0.1633 | No |
| 28 | CTSL | CTSL Entrez, Source | cathepsin L | 5451 | 0.008 | -0.1662 | No |
| 29 | DIO2 | DIO2 Entrez, Source | deiodinase, iodothyronine, type II | 5720 | 0.004 | -0.1859 | No |
| 30 | CDCA2 | CDCA2 Entrez, Source | cell division cycle associated 2 | 5754 | 0.003 | -0.1880 | No |
| 31 | ADARB2 | ADARB2 Entrez, Source | adenosine deaminase, RNA-specific, B2 (RED2 homolog rat) | 5942 | 0.001 | -0.2020 | No |
| 32 | ARNT2 | ARNT2 Entrez, Source | aryl-hydrocarbon receptor nuclear translocator 2 | 6133 | -0.002 | -0.2160 | No |
| 33 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 6595 | -0.009 | -0.2497 | No |
| 34 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 6785 | -0.012 | -0.2625 | No |
| 35 | TM7SF2 | TM7SF2 Entrez, Source | transmembrane 7 superfamily member 2 | 6881 | -0.013 | -0.2681 | No |
| 36 | CRYBB1 | CRYBB1 Entrez, Source | crystallin, beta B1 | 6934 | -0.014 | -0.2703 | No |
| 37 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 7015 | -0.015 | -0.2745 | No |
| 38 | GPSM1 | GPSM1 Entrez, Source | G-protein signalling modulator 1 (AGS3-like, C. elegans) | 7414 | -0.021 | -0.3020 | No |
| 39 | SOX11 | SOX11 Entrez, Source | SRY (sex determining region Y)-box 11 | 7530 | -0.023 | -0.3079 | No |
| 40 | LHX1 | LHX1 Entrez, Source | LIM homeobox 1 | 7941 | -0.030 | -0.3352 | No |
| 41 | DDX17 | DDX17 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 17 | 8274 | -0.035 | -0.3560 | No |
| 42 | ZP2 | ZP2 Entrez, Source | zona pellucida glycoprotein 2 (sperm receptor) | 8403 | -0.037 | -0.3611 | No |
| 43 | TTN | TTN Entrez, Source | titin | 8450 | -0.038 | -0.3600 | No |
| 44 | ENO3 | ENO3 Entrez, Source | enolase 3 (beta, muscle) | 8467 | -0.038 | -0.3566 | No |
| 45 | AURKB | AURKB Entrez, Source | aurora kinase B | 8506 | -0.039 | -0.3547 | No |
| 46 | SLIT3 | SLIT3 Entrez, Source | slit homolog 3 (Drosophila) | 8691 | -0.042 | -0.3635 | No |
| 47 | NUMA1 | NUMA1 Entrez, Source | nuclear mitotic apparatus protein 1 | 8995 | -0.047 | -0.3807 | No |
| 48 | TMPO | TMPO Entrez, Source | thymopoietin | 9189 | -0.050 | -0.3892 | No |
| 49 | GFAP | GFAP Entrez, Source | glial fibrillary acidic protein | 9399 | -0.054 | -0.3984 | No |
| 50 | TRIM39 | TRIM39 Entrez, Source | tripartite motif-containing 39 | 9616 | -0.058 | -0.4077 | No |
| 51 | PTPRN | PTPRN Entrez, Source | protein tyrosine phosphatase, receptor type, N | 9636 | -0.058 | -0.4021 | No |
| 52 | NDE1 | NDE1 Entrez, Source | nudE nuclear distribution gene E homolog 1 (A. nidulans) | 9639 | -0.058 | -0.3952 | No |
| 53 | FGFR4 | FGFR4 Entrez, Source | fibroblast growth factor receptor 4 | 9846 | -0.063 | -0.4031 | No |
| 54 | HMGA1 | HMGA1 Entrez, Source | high mobility group AT-hook 1 | 9950 | -0.065 | -0.4030 | No |
| 55 | PLEKHF1 | PLEKHF1 Entrez, Source | pleckstrin homology domain containing, family F (with FYVE domain) member 1 | 9956 | -0.066 | -0.3954 | No |
| 56 | DEK | DEK Entrez, Source | DEK oncogene (DNA binding) | 10155 | -0.069 | -0.4020 | No |
| 57 | BUB3 | BUB3 Entrez, Source | BUB3 budding uninhibited by benzimidazoles 3 homolog (yeast) | 10355 | -0.074 | -0.4081 | No |
| 58 | KHDRBS1 | KHDRBS1 Entrez, Source | KH domain containing, RNA binding, signal transduction associated 1 | 10557 | -0.079 | -0.4137 | Yes |
| 59 | HES6 | HES6 Entrez, Source | hairy and enhancer of split 6 (Drosophila) | 10589 | -0.080 | -0.4065 | Yes |
| 60 | POU3F2 | POU3F2 Entrez, Source | POU domain, class 3, transcription factor 2 | 10612 | -0.080 | -0.3985 | Yes |
| 61 | MDK | MDK Entrez, Source | midkine (neurite growth-promoting factor 2) | 10921 | -0.089 | -0.4110 | Yes |
| 62 | PRIM2A | PRIM2A Entrez, Source | primase, polypeptide 2A, 58kDa | 10985 | -0.090 | -0.4049 | Yes |
| 63 | CRYBA4 | CRYBA4 Entrez, Source | crystallin, beta A4 | 11054 | -0.092 | -0.3990 | Yes |
| 64 | HDAC2 | HDAC2 Entrez, Source | histone deacetylase 2 | 11182 | -0.095 | -0.3971 | Yes |
| 65 | BMS1L | BMS1L Entrez, Source | BMS1-like, ribosome assembly protein (yeast) | 11556 | -0.108 | -0.4122 | Yes |
| 66 | PTEN | PTEN Entrez, Source | phosphatase and tensin homolog (mutated in multiple advanced cancers 1) | 11578 | -0.108 | -0.4007 | Yes |
| 67 | FZD5 | FZD5 Entrez, Source | frizzled homolog 5 (Drosophila) | 11677 | -0.112 | -0.3945 | Yes |
| 68 | HEY1 | HEY1 Entrez, Source | hairy/enhancer-of-split related with YRPW motif 1 | 11915 | -0.122 | -0.3976 | Yes |
| 69 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 12025 | -0.128 | -0.3905 | Yes |
| 70 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 12218 | -0.137 | -0.3884 | Yes |
| 71 | ANP32B | ANP32B Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member B | 12389 | -0.148 | -0.3834 | Yes |
| 72 | DDX20 | DDX20 Entrez, Source | DEAD (Asp-Glu-Ala-Asp) box polypeptide 20 | 12437 | -0.150 | -0.3688 | Yes |
| 73 | CDC27 | CDC27 Entrez, Source | cell division cycle 27 | 12571 | -0.160 | -0.3595 | Yes |
| 74 | SYNCRIP | SYNCRIP Entrez, Source | synaptotagmin binding, cytoplasmic RNA interacting protein | 12730 | -0.176 | -0.3502 | Yes |
| 75 | NOTCH2 | NOTCH2 Entrez, Source | Notch homolog 2 (Drosophila) | 12769 | -0.180 | -0.3314 | Yes |
| 76 | FUBP1 | FUBP1 Entrez, Source | far upstream element (FUSE) binding protein 1 | 13175 | -0.284 | -0.3276 | Yes |
| 77 | SLC34A1 | SLC34A1 Entrez, Source | solute carrier family 34 (sodium phosphate), member 1 | 13245 | -0.339 | -0.2919 | Yes |
| 78 | PON1 | PON1 Entrez, Source | paraoxonase 1 | 13268 | -0.372 | -0.2487 | Yes |
| 79 | PPP1R3C | PPP1R3C Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 3C | 13309 | -0.475 | -0.1945 | Yes |
| 80 | ASCL1 | ASCL1 Entrez, Source | achaete-scute complex-like 1 (Drosophila) | 13333 | -0.811 | -0.0985 | Yes |
| 81 | EDG1 | EDG1 Entrez, Source | endothelial differentiation, sphingolipid G-protein-coupled receptor, 1 | 13334 | -0.822 | 0.0006 | Yes |