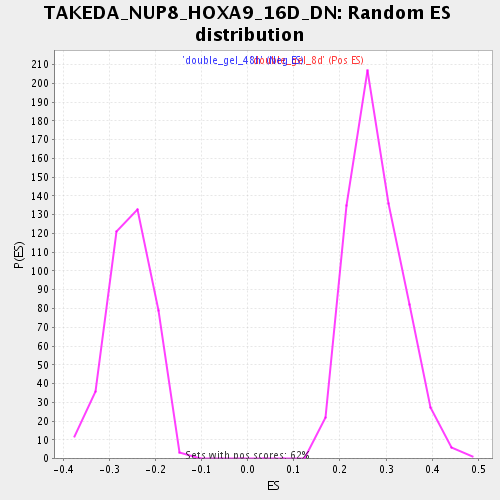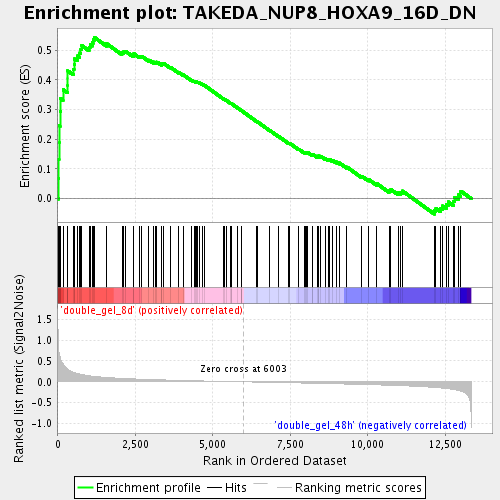
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | TAKEDA_NUP8_HOXA9_16D_DN |
| Enrichment Score (ES) | 0.54306656 |
| Normalized Enrichment Score (NES) | 1.9540858 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0072453604 |
| FWER p-Value | 0.369 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PGLYRP1 | PGLYRP1 Entrez, Source | peptidoglycan recognition protein 1 | 16 | 0.825 | 0.0683 | Yes |
| 2 | CD24 | CD24 Entrez, Source | CD24 molecule | 26 | 0.771 | 0.1325 | Yes |
| 3 | PROCR | PROCR Entrez, Source | protein C receptor, endothelial (EPCR) | 37 | 0.690 | 0.1898 | Yes |
| 4 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 43 | 0.675 | 0.2463 | Yes |
| 5 | BHLHB3 | BHLHB3 Entrez, Source | basic helix-loop-helix domain containing, class B, 3 | 71 | 0.581 | 0.2932 | Yes |
| 6 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 89 | 0.531 | 0.3366 | Yes |
| 7 | AMPD3 | AMPD3 Entrez, Source | adenosine monophosphate deaminase (isoform E) | 167 | 0.426 | 0.3667 | Yes |
| 8 | TSPAN2 | TSPAN2 Entrez, Source | tetraspanin 2 | 311 | 0.298 | 0.3810 | Yes |
| 9 | TFPI | TFPI Entrez, Source | tissue factor pathway inhibitor (lipoprotein-associated coagulation inhibitor) | 316 | 0.294 | 0.4054 | Yes |
| 10 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 317 | 0.293 | 0.4301 | Yes |
| 11 | PQLC1 | PQLC1 Entrez, Source | PQ loop repeat containing 1 | 499 | 0.228 | 0.4356 | Yes |
| 12 | CFLAR | CFLAR Entrez, Source | CASP8 and FADD-like apoptosis regulator | 523 | 0.222 | 0.4525 | Yes |
| 13 | CD59 | CD59 Entrez, Source | CD59 molecule, complement regulatory protein | 533 | 0.219 | 0.4703 | Yes |
| 14 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 627 | 0.202 | 0.4803 | Yes |
| 15 | ARG1 | ARG1 Entrez, Source | arginase, liver | 704 | 0.185 | 0.4901 | Yes |
| 16 | MSRA | MSRA Entrez, Source | methionine sulfoxide reductase A | 738 | 0.179 | 0.5027 | Yes |
| 17 | ITGA9 | ITGA9 Entrez, Source | integrin, alpha 9 | 767 | 0.175 | 0.5152 | Yes |
| 18 | HEXA | HEXA Entrez, Source | hexosaminidase A (alpha polypeptide) | 1007 | 0.146 | 0.5095 | Yes |
| 19 | CXCL1 | CXCL1 Entrez, Source | chemokine (C-X-C motif) ligand 1 (melanoma growth stimulating activity, alpha) | 1041 | 0.142 | 0.5190 | Yes |
| 20 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 1114 | 0.135 | 0.5249 | Yes |
| 21 | SDS | SDS Entrez, Source | serine dehydratase | 1136 | 0.133 | 0.5345 | Yes |
| 22 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 1170 | 0.131 | 0.5431 | Yes |
| 23 | PELI1 | PELI1 Entrez, Source | pellino homolog 1 (Drosophila) | 1568 | 0.104 | 0.5219 | No |
| 24 | KBTBD10 | KBTBD10 Entrez, Source | kelch repeat and BTB (POZ) domain containing 10 | 2069 | 0.082 | 0.4910 | No |
| 25 | APOE | APOE Entrez, Source | apolipoprotein E | 2123 | 0.080 | 0.4938 | No |
| 26 | HS3ST1 | HS3ST1 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 1 | 2172 | 0.078 | 0.4967 | No |
| 27 | LRP3 | LRP3 Entrez, Source | low density lipoprotein receptor-related protein 3 | 2439 | 0.070 | 0.4825 | No |
| 28 | CUGBP2 | CUGBP2 Entrez, Source | CUG triplet repeat, RNA binding protein 2 | 2451 | 0.069 | 0.4875 | No |
| 29 | SCGB3A1 | SCGB3A1 Entrez, Source | secretoglobin, family 3A, member 1 | 2640 | 0.064 | 0.4787 | No |
| 30 | PADI4 | PADI4 Entrez, Source | peptidyl arginine deiminase, type IV | 2700 | 0.063 | 0.4795 | No |
| 31 | ERG | ERG Entrez, Source | v-ets erythroblastosis virus E26 oncogene homolog (avian) | 2935 | 0.057 | 0.4666 | No |
| 32 | ZFHX1B | ZFHX1B Entrez, Source | zinc finger homeobox 1b | 3081 | 0.052 | 0.4601 | No |
| 33 | NPTN | NPTN Entrez, Source | neuroplastin | 3153 | 0.051 | 0.4590 | No |
| 34 | ATXN1 | ATXN1 Entrez, Source | ataxin 1 | 3197 | 0.050 | 0.4599 | No |
| 35 | ELL2 | ELL2 Entrez, Source | elongation factor, RNA polymerase II, 2 | 3352 | 0.046 | 0.4521 | No |
| 36 | MLL5 | MLL5 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia 5 (trithorax homolog, Drosophila) | 3353 | 0.046 | 0.4560 | No |
| 37 | CEACAM1 | CEACAM1 Entrez, Source | carcinoembryonic antigen-related cell adhesion molecule 1 (biliary glycoprotein) | 3419 | 0.045 | 0.4549 | No |
| 38 | SEPP1 | SEPP1 Entrez, Source | selenoprotein P, plasma, 1 | 3625 | 0.040 | 0.4428 | No |
| 39 | GGTA1 | GGTA1 Entrez, Source | glycoprotein, alpha-galactosyltransferase 1 | 3898 | 0.035 | 0.4252 | No |
| 40 | TIA1 | TIA1 Entrez, Source | TIA1 cytotoxic granule-associated RNA binding protein | 4046 | 0.032 | 0.4168 | No |
| 41 | RBMS1 | RBMS1 Entrez, Source | RNA binding motif, single stranded interacting protein 1 | 4322 | 0.027 | 0.3983 | No |
| 42 | HP | HP Entrez, Source | haptoglobin | 4392 | 0.026 | 0.3952 | No |
| 43 | CSTB | CSTB Entrez, Source | cystatin B (stefin B) | 4447 | 0.024 | 0.3932 | No |
| 44 | RPS6KA2 | RPS6KA2 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 2 | 4460 | 0.024 | 0.3943 | No |
| 45 | CYBRD1 | CYBRD1 Entrez, Source | cytochrome b reductase 1 | 4511 | 0.023 | 0.3925 | No |
| 46 | SLC16A10 | SLC16A10 Entrez, Source | solute carrier family 16, member 10 (aromatic amino acid transporter) | 4574 | 0.022 | 0.3896 | No |
| 47 | ALOX5 | ALOX5 Entrez, Source | arachidonate 5-lipoxygenase | 4658 | 0.021 | 0.3851 | No |
| 48 | NRP1 | NRP1 Entrez, Source | neuropilin 1 | 4730 | 0.020 | 0.3814 | No |
| 49 | ADORA3 | ADORA3 Entrez, Source | adenosine A3 receptor | 5346 | 0.009 | 0.3358 | No |
| 50 | SFXN5 | SFXN5 Entrez, Source | sideroflexin 5 | 5377 | 0.009 | 0.3343 | No |
| 51 | RBP1 | RBP1 Entrez, Source | retinol binding protein 1, cellular | 5430 | 0.008 | 0.3310 | No |
| 52 | PRDX2 | PRDX2 Entrez, Source | peroxiredoxin 2 | 5562 | 0.006 | 0.3217 | No |
| 53 | ZFAND3 | ZFAND3 Entrez, Source | zinc finger, AN1-type domain 3 | 5590 | 0.006 | 0.3201 | No |
| 54 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 5599 | 0.006 | 0.3200 | No |
| 55 | MTMR3 | MTMR3 Entrez, Source | myotubularin related protein 3 | 5782 | 0.003 | 0.3065 | No |
| 56 | MMP8 | MMP8 Entrez, Source | matrix metallopeptidase 8 (neutrophil collagenase) | 5909 | 0.001 | 0.2971 | No |
| 57 | CAMK2G | CAMK2G Entrez, Source | calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma | 5913 | 0.001 | 0.2970 | No |
| 58 | ZNF148 | ZNF148 Entrez, Source | zinc finger protein 148 | 5916 | 0.001 | 0.2970 | No |
| 59 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 6403 | -0.006 | 0.2608 | No |
| 60 | SCN11A | SCN11A Entrez, Source | sodium channel, voltage-gated, type XI, alpha | 6451 | -0.007 | 0.2578 | No |
| 61 | CHI3L1 | CHI3L1 Entrez, Source | chitinase 3-like 1 (cartilage glycoprotein-39) | 6833 | -0.013 | 0.2301 | No |
| 62 | CORO2A | CORO2A Entrez, Source | coronin, actin binding protein, 2A | 7122 | -0.017 | 0.2098 | No |
| 63 | OLIG1 | OLIG1 Entrez, Source | oligodendrocyte transcription factor 1 | 7444 | -0.022 | 0.1874 | No |
| 64 | MTUS1 | MTUS1 Entrez, Source | mitochondrial tumor suppressor 1 | 7482 | -0.022 | 0.1865 | No |
| 65 | NRP2 | NRP2 Entrez, Source | neuropilin 2 | 7770 | -0.027 | 0.1671 | No |
| 66 | NCOR1 | NCOR1 Entrez, Source | nuclear receptor co-repressor 1 | 7954 | -0.030 | 0.1558 | No |
| 67 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 7982 | -0.030 | 0.1563 | No |
| 68 | CLTC | CLTC Entrez, Source | clathrin, heavy chain (Hc) | 8030 | -0.031 | 0.1554 | No |
| 69 | NSF | NSF Entrez, Source | N-ethylmaleimide-sensitive factor | 8063 | -0.032 | 0.1557 | No |
| 70 | CENTB2 | CENTB2 Entrez, Source | centaurin, beta 2 | 8209 | -0.034 | 0.1476 | No |
| 71 | GGA1 | GGA1 Entrez, Source | golgi associated, gamma adaptin ear containing, ARF binding protein 1 | 8215 | -0.034 | 0.1501 | No |
| 72 | PSMB2 | PSMB2 Entrez, Source | proteasome (prosome, macropain) subunit, beta type, 2 | 8370 | -0.037 | 0.1415 | No |
| 73 | MGAT5 | MGAT5 Entrez, Source | mannosyl (alpha-1,6-)-glycoprotein beta-1,6-N-acetyl-glucosaminyltransferase | 8412 | -0.038 | 0.1416 | No |
| 74 | LRRFIP2 | LRRFIP2 Entrez, Source | leucine rich repeat (in FLII) interacting protein 2 | 8422 | -0.038 | 0.1441 | No |
| 75 | DNAH11 | DNAH11 Entrez, Source | dynein, axonemal, heavy chain 11 | 8489 | -0.039 | 0.1424 | No |
| 76 | SLC11A2 | SLC11A2 Entrez, Source | solute carrier family 11 (proton-coupled divalent metal ion transporters), member 2 | 8634 | -0.041 | 0.1350 | No |
| 77 | RAP1A | RAP1A Entrez, Source | RAP1A, member of RAS oncogene family | 8724 | -0.043 | 0.1318 | No |
| 78 | ALK | ALK Entrez, Source | anaplastic lymphoma kinase (Ki-1) | 8776 | -0.043 | 0.1316 | No |
| 79 | CAPN3 | CAPN3 Entrez, Source | calpain 3, (p94) | 8869 | -0.045 | 0.1285 | No |
| 80 | DPP4 | DPP4 Entrez, Source | dipeptidyl-peptidase 4 (CD26, adenosine deaminase complexing protein 2) | 8984 | -0.047 | 0.1238 | No |
| 81 | IGFBP4 | IGFBP4 Entrez, Source | insulin-like growth factor binding protein 4 | 9082 | -0.049 | 0.1206 | No |
| 82 | SYNJ2 | SYNJ2 Entrez, Source | synaptojanin 2 | 9325 | -0.053 | 0.1068 | No |
| 83 | TMEM37 | TMEM37 Entrez, Source | transmembrane protein 37 | 9801 | -0.062 | 0.0761 | No |
| 84 | CEBPE | CEBPE Entrez, Source | CCAAT/enhancer binding protein (C/EBP), epsilon | 10021 | -0.067 | 0.0652 | No |
| 85 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 10296 | -0.072 | 0.0506 | No |
| 86 | GLMN | GLMN Entrez, Source | glomulin, FKBP associated protein | 10689 | -0.082 | 0.0279 | No |
| 87 | KPNA3 | KPNA3 Entrez, Source | karyopherin alpha 3 (importin alpha 4) | 10744 | -0.084 | 0.0309 | No |
| 88 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 10976 | -0.090 | 0.0211 | No |
| 89 | GPD2 | GPD2 Entrez, Source | glycerol-3-phosphate dehydrogenase 2 (mitochondrial) | 11070 | -0.092 | 0.0218 | No |
| 90 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 11118 | -0.094 | 0.0262 | No |
| 91 | MMP9 | MMP9 Entrez, Source | matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) | 12150 | -0.134 | -0.0404 | No |
| 92 | EXOSC7 | EXOSC7 Entrez, Source | exosome component 7 | 12198 | -0.137 | -0.0324 | No |
| 93 | QKI | QKI Entrez, Source | quaking homolog, KH domain RNA binding (mouse) | 12354 | -0.146 | -0.0319 | No |
| 94 | PAPD4 | PAPD4 Entrez, Source | PAP associated domain containing 4 | 12421 | -0.150 | -0.0242 | No |
| 95 | TGFBR1 | TGFBR1 Entrez, Source | transforming growth factor, beta receptor I (activin A receptor type II-like kinase, 53kDa) | 12538 | -0.158 | -0.0197 | No |
| 96 | CCDC52 | CCDC52 Entrez, Source | coiled-coil domain containing 52 | 12599 | -0.163 | -0.0105 | No |
| 97 | EDD1 | EDD1 Entrez, Source | E3 ubiquitin protein ligase, HECT domain containing, 1 | 12770 | -0.180 | -0.0082 | No |
| 98 | SULT1C1 | SULT1C1 Entrez, Source | sulfotransferase family, cytosolic, 1C, member 1 | 12806 | -0.186 | 0.0048 | No |
| 99 | RARRES1 | RARRES1 Entrez, Source | retinoic acid receptor responder (tazarotene induced) 1 | 12923 | -0.203 | 0.0132 | No |
| 100 | RNF141 | RNF141 Entrez, Source | ring finger protein 141 | 13001 | -0.218 | 0.0257 | No |