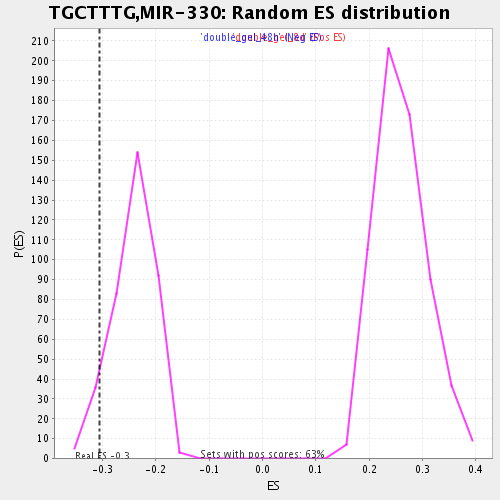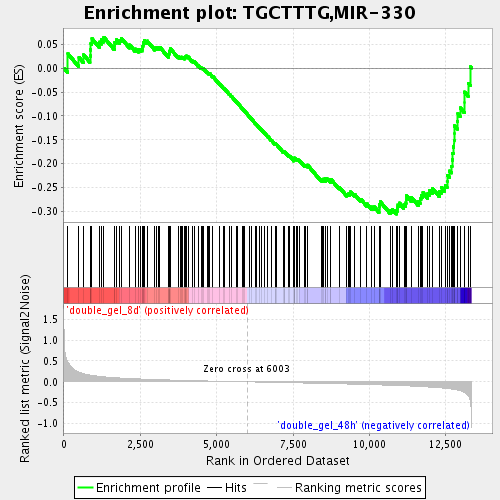
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_48h |
| GeneSet | TGCTTTG,MIR-330 |
| Enrichment Score (ES) | -0.30546054 |
| Normalized Enrichment Score (NES) | -1.2613751 |
| Nominal p-value | 0.077747986 |
| FDR q-value | 0.5213534 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | GPM6A | GPM6A Entrez, Source | glycoprotein M6A | 130 | 0.466 | 0.0300 | No |
| 2 | ITM2C | ITM2C Entrez, Source | integral membrane protein 2C | 491 | 0.231 | 0.0224 | No |
| 3 | CCND3 | CCND3 Entrez, Source | cyclin D3 | 643 | 0.198 | 0.0279 | No |
| 4 | TRIM3 | TRIM3 Entrez, Source | tripartite motif-containing 3 | 854 | 0.162 | 0.0258 | No |
| 5 | GPHN | GPHN Entrez, Source | gephyrin | 868 | 0.160 | 0.0385 | No |
| 6 | TRAK2 | TRAK2 Entrez, Source | trafficking protein, kinesin binding 2 | 882 | 0.158 | 0.0510 | No |
| 7 | DPYSL3 | DPYSL3 Entrez, Source | dihydropyrimidinase-like 3 | 917 | 0.154 | 0.0616 | No |
| 8 | IRX2 | IRX2 Entrez, Source | iroquois homeobox protein 2 | 1157 | 0.132 | 0.0547 | No |
| 9 | SYT7 | SYT7 Entrez, Source | synaptotagmin VII | 1219 | 0.126 | 0.0609 | No |
| 10 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 1305 | 0.119 | 0.0647 | No |
| 11 | RUNX3 | RUNX3 Entrez, Source | runt-related transcription factor 3 | 1657 | 0.100 | 0.0466 | No |
| 12 | BIRC4 | BIRC4 Entrez, Source | baculoviral IAP repeat-containing 4 | 1664 | 0.099 | 0.0546 | No |
| 13 | DCX | DCX Entrez, Source | doublecortex; lissencephaly, X-linked (doublecortin) | 1708 | 0.097 | 0.0596 | No |
| 14 | CEP170 | CEP170 Entrez, Source | centrosomal protein 170kDa | 1823 | 0.092 | 0.0588 | No |
| 15 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 1874 | 0.090 | 0.0627 | No |
| 16 | MTCH1 | MTCH1 Entrez, Source | mitochondrial carrier homolog 1 (C. elegans) | 2149 | 0.079 | 0.0487 | No |
| 17 | FRK | FRK Entrez, Source | fyn-related kinase | 2344 | 0.073 | 0.0402 | No |
| 18 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 2434 | 0.070 | 0.0394 | No |
| 19 | D4S234E | D4S234E Entrez, Source | - | 2507 | 0.067 | 0.0397 | No |
| 20 | PHC2 | PHC2 Entrez, Source | polyhomeotic homolog 2 (Drosophila) | 2559 | 0.066 | 0.0414 | No |
| 21 | TLOC1 | TLOC1 Entrez, Source | translocation protein 1 | 2571 | 0.066 | 0.0462 | No |
| 22 | PSCD3 | PSCD3 Entrez, Source | pleckstrin homology, Sec7 and coiled-coil domains 3 | 2590 | 0.065 | 0.0504 | No |
| 23 | DAG1 | DAG1 Entrez, Source | dystroglycan 1 (dystrophin-associated glycoprotein 1) | 2610 | 0.065 | 0.0545 | No |
| 24 | BTG1 | BTG1 Entrez, Source | B-cell translocation gene 1, anti-proliferative | 2629 | 0.064 | 0.0586 | No |
| 25 | CREBBP | CREBBP Entrez, Source | CREB binding protein (Rubinstein-Taybi syndrome) | 2722 | 0.062 | 0.0569 | No |
| 26 | SH3GL2 | SH3GL2 Entrez, Source | SH3-domain GRB2-like 2 | 2978 | 0.055 | 0.0423 | No |
| 27 | TGFBR3 | TGFBR3 Entrez, Source | transforming growth factor, beta receptor III (betaglycan, 300kDa) | 3034 | 0.054 | 0.0427 | No |
| 28 | BICD2 | BICD2 Entrez, Source | bicaudal D homolog 2 (Drosophila) | 3088 | 0.052 | 0.0431 | No |
| 29 | ARMCX6 | ARMCX6 Entrez, Source | armadillo repeat containing, X-linked 6 | 3136 | 0.051 | 0.0439 | No |
| 30 | NR4A3 | NR4A3 Entrez, Source | nuclear receptor subfamily 4, group A, member 3 | 3417 | 0.045 | 0.0265 | No |
| 31 | CHP | CHP Entrez, Source | - | 3438 | 0.044 | 0.0288 | No |
| 32 | PDGFD | PDGFD Entrez, Source | platelet derived growth factor D | 3442 | 0.044 | 0.0323 | No |
| 33 | PTGFRN | PTGFRN Entrez, Source | prostaglandin F2 receptor negative regulator | 3455 | 0.044 | 0.0352 | No |
| 34 | CLCN5 | CLCN5 Entrez, Source | chloride channel 5 (nephrolithiasis 2, X-linked, Dent disease) | 3470 | 0.044 | 0.0379 | No |
| 35 | LUZP1 | LUZP1 Entrez, Source | leucine zipper protein 1 | 3481 | 0.043 | 0.0408 | No |
| 36 | HDAC5 | HDAC5 Entrez, Source | histone deacetylase 5 | 3742 | 0.038 | 0.0243 | No |
| 37 | MTPN | MTPN Entrez, Source | myotrophin | 3810 | 0.036 | 0.0223 | No |
| 38 | DPP10 | DPP10 Entrez, Source | dipeptidyl-peptidase 10 | 3838 | 0.036 | 0.0234 | No |
| 39 | SLC4A10 | SLC4A10 Entrez, Source | solute carrier family 4, sodium bicarbonate transporter-like, member 10 | 3892 | 0.035 | 0.0223 | No |
| 40 | RND3 | RND3 Entrez, Source | Rho family GTPase 3 | 3937 | 0.034 | 0.0219 | No |
| 41 | PCAF | PCAF Entrez, Source | p300/CBP-associated factor | 3962 | 0.034 | 0.0229 | No |
| 42 | GRID1 | GRID1 Entrez, Source | glutamate receptor, ionotropic, delta 1 | 3987 | 0.033 | 0.0239 | No |
| 43 | EGR4 | EGR4 Entrez, Source | early growth response 4 | 4003 | 0.033 | 0.0256 | No |
| 44 | KCNA6 | KCNA6 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 6 | 4066 | 0.032 | 0.0236 | No |
| 45 | TEAD1 | TEAD1 Entrez, Source | TEA domain family member 1 (SV40 transcriptional enhancer factor) | 4210 | 0.029 | 0.0152 | No |
| 46 | GALNT7 | GALNT7 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 7 (GalNAc-T7) | 4259 | 0.028 | 0.0140 | No |
| 47 | BTRC | BTRC Entrez, Source | beta-transducin repeat containing | 4396 | 0.025 | 0.0059 | No |
| 48 | POU3F1 | POU3F1 Entrez, Source | POU domain, class 3, transcription factor 1 | 4491 | 0.024 | 0.0007 | No |
| 49 | SYT5 | SYT5 Entrez, Source | synaptotagmin V | 4526 | 0.023 | 0.0001 | No |
| 50 | PPP1R1B | PPP1R1B Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 1B (dopamine and cAMP regulated phosphoprotein, DARPP-32) | 4556 | 0.022 | -0.0002 | No |
| 51 | PPM1B | PPM1B Entrez, Source | protein phosphatase 1B (formerly 2C), magnesium-dependent, beta isoform | 4685 | 0.020 | -0.0081 | No |
| 52 | FNTB | FNTB Entrez, Source | farnesyltransferase, CAAX box, beta | 4744 | 0.019 | -0.0109 | No |
| 53 | PLCB1 | PLCB1 Entrez, Source | phospholipase C, beta 1 (phosphoinositide-specific) | 4764 | 0.019 | -0.0107 | No |
| 54 | TFEB | TFEB Entrez, Source | transcription factor EB | 4873 | 0.017 | -0.0174 | No |
| 55 | CMPK | CMPK Entrez, Source | cytidylate kinase | 5102 | 0.013 | -0.0336 | No |
| 56 | CSNK1G3 | CSNK1G3 Entrez, Source | casein kinase 1, gamma 3 | 5235 | 0.011 | -0.0426 | No |
| 57 | PTPN4 | PTPN4 Entrez, Source | protein tyrosine phosphatase, non-receptor type 4 (megakaryocyte) | 5255 | 0.011 | -0.0431 | No |
| 58 | PCDHA11 | PCDHA11 Entrez, Source | protocadherin alpha 11 | 5427 | 0.008 | -0.0554 | No |
| 59 | PCDHA4 | PCDHA4 Entrez, Source | protocadherin alpha 4 | 5482 | 0.008 | -0.0588 | No |
| 60 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 5636 | 0.005 | -0.0700 | No |
| 61 | E2F1 | E2F1 Entrez, Source | E2F transcription factor 1 | 5669 | 0.005 | -0.0720 | No |
| 62 | FYTTD1 | FYTTD1 Entrez, Source | forty-two-three domain containing 1 | 5831 | 0.002 | -0.0840 | No |
| 63 | MAP2K5 | MAP2K5 Entrez, Source | mitogen-activated protein kinase kinase 5 | 5887 | 0.002 | -0.0880 | No |
| 64 | STAU1 | STAU1 Entrez, Source | staufen, RNA binding protein, homolog 1 (Drosophila) | 5908 | 0.001 | -0.0894 | No |
| 65 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 6060 | -0.001 | -0.1008 | No |
| 66 | ACBD3 | ACBD3 Entrez, Source | acyl-Coenzyme A binding domain containing 3 | 6131 | -0.002 | -0.1059 | No |
| 67 | STXBP3 | STXBP3 Entrez, Source | syntaxin binding protein 3 | 6260 | -0.004 | -0.1153 | No |
| 68 | ESR1 | ESR1 Entrez, Source | estrogen receptor 1 | 6300 | -0.005 | -0.1179 | No |
| 69 | MBNL1 | MBNL1 Entrez, Source | muscleblind-like (Drosophila) | 6403 | -0.006 | -0.1250 | No |
| 70 | GCG | GCG Entrez, Source | glucagon | 6455 | -0.007 | -0.1283 | No |
| 71 | KCNIP2 | KCNIP2 Entrez, Source | Kv channel interacting protein 2 | 6562 | -0.009 | -0.1356 | No |
| 72 | ARHGEF9 | ARHGEF9 Entrez, Source | Cdc42 guanine nucleotide exchange factor (GEF) 9 | 6676 | -0.010 | -0.1433 | No |
| 73 | MTMR9 | MTMR9 Entrez, Source | myotubularin related protein 9 | 6787 | -0.012 | -0.1506 | No |
| 74 | RAB6A | RAB6A Entrez, Source | RAB6A, member RAS oncogene family | 6910 | -0.014 | -0.1587 | No |
| 75 | STRBP | STRBP Entrez, Source | spermatid perinuclear RNA binding protein | 6915 | -0.014 | -0.1578 | No |
| 76 | EIF4E | EIF4E Entrez, Source | eukaryotic translation initiation factor 4E | 6949 | -0.014 | -0.1591 | No |
| 77 | PCDHA13 | PCDHA13 Entrez, Source | protocadherin alpha 13 | 7173 | -0.018 | -0.1745 | No |
| 78 | GRM5 | GRM5 Entrez, Source | glutamate receptor, metabotropic 5 | 7203 | -0.018 | -0.1751 | No |
| 79 | SLC17A6 | SLC17A6 Entrez, Source | solute carrier family 17 (sodium-dependent inorganic phosphate cotransporter), member 6 | 7230 | -0.019 | -0.1755 | No |
| 80 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 7356 | -0.020 | -0.1832 | No |
| 81 | SLC25A3 | SLC25A3 Entrez, Source | solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3 | 7388 | -0.021 | -0.1838 | No |
| 82 | DHX40 | DHX40 Entrez, Source | DEAH (Asp-Glu-Ala-His) box polypeptide 40 | 7529 | -0.023 | -0.1925 | No |
| 83 | SOX11 | SOX11 Entrez, Source | SRY (sex determining region Y)-box 11 | 7530 | -0.023 | -0.1905 | No |
| 84 | MRFAP1 | MRFAP1 Entrez, Source | Mof4 family associated protein 1 | 7544 | -0.023 | -0.1895 | No |
| 85 | GNL1 | GNL1 Entrez, Source | guanine nucleotide binding protein-like 1 | 7550 | -0.023 | -0.1879 | No |
| 86 | AP2M1 | AP2M1 Entrez, Source | adaptor-related protein complex 2, mu 1 subunit | 7615 | -0.024 | -0.1906 | No |
| 87 | HIVEP2 | HIVEP2 Entrez, Source | human immunodeficiency virus type I enhancer binding protein 2 | 7650 | -0.025 | -0.1911 | No |
| 88 | PDCD4 | PDCD4 Entrez, Source | programmed cell death 4 (neoplastic transformation inhibitor) | 7707 | -0.026 | -0.1931 | No |
| 89 | AGTR2 | AGTR2 Entrez, Source | angiotensin II receptor, type 2 | 7888 | -0.029 | -0.2043 | No |
| 90 | GNAO1 | GNAO1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O | 7919 | -0.029 | -0.2041 | No |
| 91 | DLX1 | DLX1 Entrez, Source | distal-less homeobox 1 | 7960 | -0.030 | -0.2045 | No |
| 92 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 7971 | -0.030 | -0.2027 | No |
| 93 | SOSTDC1 | SOSTDC1 Entrez, Source | sclerostin domain containing 1 | 8420 | -0.038 | -0.2334 | No |
| 94 | MAPK1 | MAPK1 Entrez, Source | mitogen-activated protein kinase 1 | 8472 | -0.038 | -0.2340 | No |
| 95 | PCYT1B | PCYT1B Entrez, Source | phosphate cytidylyltransferase 1, choline, beta | 8490 | -0.039 | -0.2320 | No |
| 96 | RAP2A | RAP2A Entrez, Source | RAP2A, member of RAS oncogene family | 8557 | -0.040 | -0.2336 | No |
| 97 | MTDH | MTDH Entrez, Source | metadherin | 8564 | -0.040 | -0.2306 | No |
| 98 | SP4 | SP4 Entrez, Source | Sp4 transcription factor | 8627 | -0.041 | -0.2318 | No |
| 99 | RAP1A | RAP1A Entrez, Source | RAP1A, member of RAS oncogene family | 8724 | -0.043 | -0.2355 | No |
| 100 | GATAD2B | GATAD2B Entrez, Source | GATA zinc finger domain containing 2B | 8739 | -0.043 | -0.2329 | No |
| 101 | LYN | LYN Entrez, Source | v-yes-1 Yamaguchi sarcoma viral related oncogene homolog | 9007 | -0.047 | -0.2491 | No |
| 102 | SLC8A3 | SLC8A3 Entrez, Source | solute carrier family 8 (sodium-calcium exchanger), member 3 | 9258 | -0.052 | -0.2636 | No |
| 103 | NF2 | NF2 Entrez, Source | neurofibromin 2 (bilateral acoustic neuroma) | 9310 | -0.053 | -0.2630 | No |
| 104 | AMPH | AMPH Entrez, Source | amphiphysin (Stiff-Man syndrome with breast cancer 128kDa autoantigen) | 9358 | -0.053 | -0.2620 | No |
| 105 | PCMT1 | PCMT1 Entrez, Source | protein-L-isoaspartate (D-aspartate) O-methyltransferase | 9378 | -0.054 | -0.2588 | No |
| 106 | IFIT1 | IFIT1 Entrez, Source | interferon-induced protein with tetratricopeptide repeats 1 | 9511 | -0.056 | -0.2640 | No |
| 107 | DCAMKL1 | DCAMKL1 Entrez, Source | doublecortin and CaM kinase-like 1 | 9719 | -0.060 | -0.2746 | No |
| 108 | GRIA3 | GRIA3 Entrez, Source | glutamate receptor, ionotrophic, AMPA 3 | 9917 | -0.065 | -0.2840 | No |
| 109 | APC | APC Entrez, Source | adenomatosis polyposis coli | 10080 | -0.068 | -0.2904 | No |
| 110 | DEK | DEK Entrez, Source | DEK oncogene (DNA binding) | 10155 | -0.069 | -0.2901 | No |
| 111 | ATP2B1 | ATP2B1 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 1 | 10314 | -0.073 | -0.2959 | No |
| 112 | PUM2 | PUM2 Entrez, Source | pumilio homolog 2 (Drosophila) | 10315 | -0.073 | -0.2896 | No |
| 113 | BMP2K | BMP2K Entrez, Source | BMP2 inducible kinase | 10329 | -0.073 | -0.2844 | No |
| 114 | METAP2 | METAP2 Entrez, Source | methionyl aminopeptidase 2 | 10345 | -0.074 | -0.2792 | No |
| 115 | ATP2B2 | ATP2B2 Entrez, Source | ATPase, Ca++ transporting, plasma membrane 2 | 10683 | -0.082 | -0.2978 | No |
| 116 | FIGF | FIGF Entrez, Source | c-fos induced growth factor (vascular endothelial growth factor D) | 10740 | -0.084 | -0.2948 | No |
| 117 | MLLT10 | MLLT10 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 10 | 10881 | -0.088 | -0.2980 | Yes |
| 118 | RNF126 | RNF126 Entrez, Source | ring finger protein 126 | 10907 | -0.088 | -0.2923 | Yes |
| 119 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 10930 | -0.089 | -0.2864 | Yes |
| 120 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 10976 | -0.090 | -0.2821 | Yes |
| 121 | NRXN3 | NRXN3 Entrez, Source | neurexin 3 | 11130 | -0.094 | -0.2857 | Yes |
| 122 | PRKCB1 | PRKCB1 Entrez, Source | protein kinase C, beta 1 | 11192 | -0.096 | -0.2822 | Yes |
| 123 | PRR3 | PRR3 Entrez, Source | proline rich 3 | 11198 | -0.096 | -0.2744 | Yes |
| 124 | TPM3 | TPM3 Entrez, Source | tropomyosin 3 | 11208 | -0.096 | -0.2668 | Yes |
| 125 | SHOX2 | SHOX2 Entrez, Source | short stature homeobox 2 | 11372 | -0.101 | -0.2706 | Yes |
| 126 | NPTXR | NPTXR Entrez, Source | neuronal pentraxin receptor | 11606 | -0.110 | -0.2789 | Yes |
| 127 | NCOA2 | NCOA2 Entrez, Source | nuclear receptor coactivator 2 | 11659 | -0.111 | -0.2733 | Yes |
| 128 | EFNA3 | EFNA3 Entrez, Source | ephrin-A3 | 11688 | -0.113 | -0.2658 | Yes |
| 129 | OGT | OGT Entrez, Source | O-linked N-acetylglucosamine (GlcNAc) transferase (UDP-N-acetylglucosamine:polypeptide-N-acetylglucosaminyl transferase) | 11749 | -0.115 | -0.2605 | Yes |
| 130 | PDLIM5 | PDLIM5 Entrez, Source | PDZ and LIM domain 5 | 11910 | -0.122 | -0.2622 | Yes |
| 131 | TRIP12 | TRIP12 Entrez, Source | thyroid hormone receptor interactor 12 | 11969 | -0.125 | -0.2560 | Yes |
| 132 | CSNK1A1 | CSNK1A1 Entrez, Source | casein kinase 1, alpha 1 | 12054 | -0.129 | -0.2513 | Yes |
| 133 | TARDBP | TARDBP Entrez, Source | TAR DNA binding protein | 12292 | -0.142 | -0.2572 | Yes |
| 134 | QKI | QKI Entrez, Source | quaking homolog, KH domain RNA binding (mouse) | 12354 | -0.146 | -0.2493 | Yes |
| 135 | BMPR2 | BMPR2 Entrez, Source | bone morphogenetic protein receptor, type II (serine/threonine kinase) | 12471 | -0.153 | -0.2451 | Yes |
| 136 | MTM1 | MTM1 Entrez, Source | myotubularin 1 | 12542 | -0.158 | -0.2369 | Yes |
| 137 | SFRS10 | SFRS10 Entrez, Source | splicing factor, arginine/serine-rich 10 (transformer 2 homolog, Drosophila) | 12549 | -0.158 | -0.2238 | Yes |
| 138 | RBM12 | RBM12 Entrez, Source | RNA binding motif protein 12 | 12622 | -0.165 | -0.2151 | Yes |
| 139 | ETF1 | ETF1 Entrez, Source | eukaryotic translation termination factor 1 | 12680 | -0.171 | -0.2049 | Yes |
| 140 | NUPL1 | NUPL1 Entrez, Source | nucleoporin like 1 | 12703 | -0.173 | -0.1917 | Yes |
| 141 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 12718 | -0.175 | -0.1778 | Yes |
| 142 | NDNL2 | NDNL2 Entrez, Source | necdin-like 2 | 12742 | -0.177 | -0.1644 | Yes |
| 143 | MAP2K1 | MAP2K1 Entrez, Source | mitogen-activated protein kinase kinase 1 | 12766 | -0.180 | -0.1508 | Yes |
| 144 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 12773 | -0.181 | -0.1358 | Yes |
| 145 | ID2 | ID2 Entrez, Source | inhibitor of DNA binding 2, dominant negative helix-loop-helix protein | 12777 | -0.181 | -0.1205 | Yes |
| 146 | HNRPU | HNRPU Entrez, Source | heterogeneous nuclear ribonucleoprotein U (scaffold attachment factor A) | 12874 | -0.196 | -0.1111 | Yes |
| 147 | TDG | TDG Entrez, Source | thymine-DNA glycosylase | 12893 | -0.199 | -0.0955 | Yes |
| 148 | DICER1 | DICER1 Entrez, Source | Dicer1, Dcr-1 homolog (Drosophila) | 12970 | -0.213 | -0.0830 | Yes |
| 149 | FNBP4 | FNBP4 Entrez, Source | formin binding protein 4 | 13108 | -0.255 | -0.0716 | Yes |
| 150 | GPAM | GPAM Entrez, Source | glycerol-3-phosphate acyltransferase, mitochondrial | 13110 | -0.256 | -0.0498 | Yes |
| 151 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 13231 | -0.325 | -0.0311 | Yes |
| 152 | OTUD4 | OTUD4 Entrez, Source | OTU domain containing 4 | 13303 | -0.462 | 0.0030 | Yes |