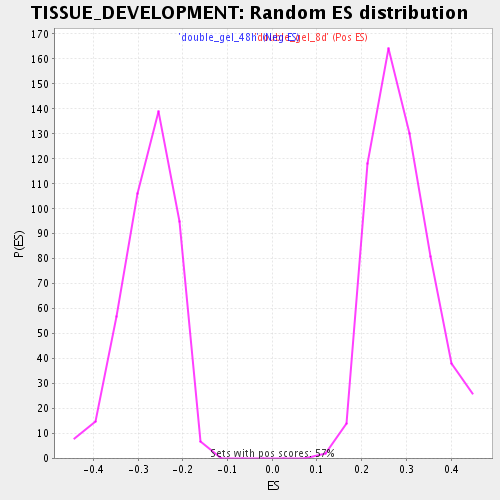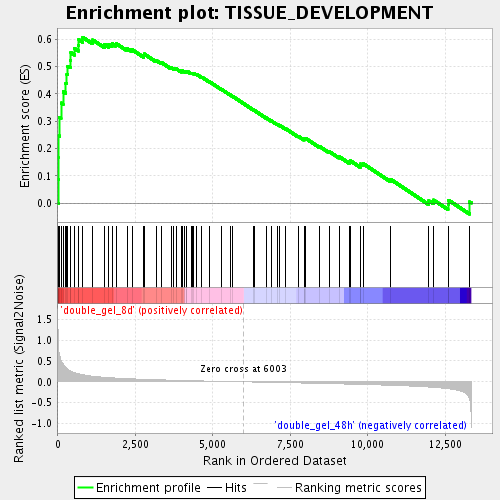
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | TISSUE_DEVELOPMENT |
| Enrichment Score (ES) | 0.6078109 |
| Normalized Enrichment Score (NES) | 2.1051087 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.0010715263 |
| FWER p-Value | 0.027 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LAMC2 | LAMC2 Entrez, Source | laminin, gamma 2 | 19 | 0.791 | 0.0855 | Yes |
| 2 | CTGF | CTGF Entrez, Source | connective tissue growth factor | 27 | 0.757 | 0.1682 | Yes |
| 3 | COL5A2 | COL5A2 Entrez, Source | collagen, type V, alpha 2 | 34 | 0.714 | 0.2462 | Yes |
| 4 | MGP | MGP Entrez, Source | matrix Gla protein | 52 | 0.634 | 0.3145 | Yes |
| 5 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 101 | 0.510 | 0.3669 | Yes |
| 6 | FST | FST Entrez, Source | follistatin | 169 | 0.423 | 0.4084 | Yes |
| 7 | LAMC1 | LAMC1 Entrez, Source | laminin, gamma 1 (formerly LAMB2) | 252 | 0.342 | 0.4398 | Yes |
| 8 | STS | STS Entrez, Source | steroid sulfatase (microsomal), arylsulfatase C, isozyme S | 288 | 0.316 | 0.4719 | Yes |
| 9 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 317 | 0.293 | 0.5021 | Yes |
| 10 | BTD | BTD Entrez, Source | biotinidase | 401 | 0.259 | 0.5242 | Yes |
| 11 | KLF4 | KLF4 Entrez, Source | Kruppel-like factor 4 (gut) | 417 | 0.254 | 0.5510 | Yes |
| 12 | GJB5 | GJB5 Entrez, Source | gap junction protein, beta 5 (connexin 31.1) | 534 | 0.219 | 0.5664 | Yes |
| 13 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 659 | 0.195 | 0.5785 | Yes |
| 14 | COL1A1 | COL1A1 Entrez, Source | collagen, type I, alpha 1 | 673 | 0.190 | 0.5985 | Yes |
| 15 | ALDH3A2 | ALDH3A2 Entrez, Source | aldehyde dehydrogenase 3 family, member A2 | 798 | 0.170 | 0.6078 | Yes |
| 16 | APOA5 | APOA5 Entrez, Source | apolipoprotein A-V | 1126 | 0.134 | 0.5979 | No |
| 17 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 1506 | 0.108 | 0.5812 | No |
| 18 | PLOD1 | PLOD1 Entrez, Source | procollagen-lysine 1, 2-oxoglutarate 5-dioxygenase 1 | 1639 | 0.101 | 0.5824 | No |
| 19 | COL11A1 | COL11A1 Entrez, Source | collagen, type XI, alpha 1 | 1749 | 0.095 | 0.5846 | No |
| 20 | RUNX2 | RUNX2 Entrez, Source | runt-related transcription factor 2 | 1874 | 0.090 | 0.5851 | No |
| 21 | OSTF1 | OSTF1 Entrez, Source | osteoclast stimulating factor 1 | 2235 | 0.076 | 0.5664 | No |
| 22 | FLOT2 | FLOT2 Entrez, Source | flotillin 2 | 2393 | 0.071 | 0.5624 | No |
| 23 | TIE1 | TIE1 Entrez, Source | tyrosine kinase with immunoglobulin-like and EGF-like domains 1 | 2769 | 0.061 | 0.5408 | No |
| 24 | UGCG | UGCG Entrez, Source | UDP-glucose ceramide glucosyltransferase | 2779 | 0.061 | 0.5468 | No |
| 25 | DSPP | DSPP Entrez, Source | dentin sialophosphoprotein | 3167 | 0.050 | 0.5232 | No |
| 26 | TBX3 | TBX3 Entrez, Source | T-box 3 (ulnar mammary syndrome) | 3345 | 0.046 | 0.5149 | No |
| 27 | EYA2 | EYA2 Entrez, Source | eyes absent homolog 2 (Drosophila) | 3653 | 0.040 | 0.4962 | No |
| 28 | IL17F | IL17F Entrez, Source | interleukin 17F | 3733 | 0.038 | 0.4944 | No |
| 29 | RBP2 | RBP2 Entrez, Source | retinol binding protein 2, cellular | 3814 | 0.036 | 0.4923 | No |
| 30 | AHSG | AHSG Entrez, Source | alpha-2-HS-glycoprotein | 3981 | 0.033 | 0.4835 | No |
| 31 | KL | KL Entrez, Source | klotho | 4012 | 0.033 | 0.4848 | No |
| 32 | DSP | DSP Entrez, Source | desmoplakin | 4079 | 0.031 | 0.4833 | No |
| 33 | FGF7 | FGF7 Entrez, Source | fibroblast growth factor 7 (keratinocyte growth factor) | 4156 | 0.030 | 0.4809 | No |
| 34 | GDF11 | GDF11 Entrez, Source | growth differentiation factor 11 | 4158 | 0.030 | 0.4841 | No |
| 35 | LAMA3 | LAMA3 Entrez, Source | laminin, alpha 3 | 4300 | 0.027 | 0.4765 | No |
| 36 | DMP1 | DMP1 Entrez, Source | dentin matrix acidic phosphoprotein | 4341 | 0.026 | 0.4764 | No |
| 37 | BMP1 | BMP1 Entrez, Source | bone morphogenetic protein 1 | 4389 | 0.026 | 0.4756 | No |
| 38 | VAX2 | VAX2 Entrez, Source | ventral anterior homeobox 2 | 4472 | 0.024 | 0.4721 | No |
| 39 | KLK8 | KLK8 Entrez, Source | kallikrein 8 (neuropsin/ovasin) | 4644 | 0.021 | 0.4615 | No |
| 40 | PTF1A | PTF1A Entrez, Source | pancreas specific transcription factor, 1a | 4886 | 0.017 | 0.4452 | No |
| 41 | SLIT2 | SLIT2 Entrez, Source | slit homolog 2 (Drosophila) | 5293 | 0.010 | 0.4158 | No |
| 42 | GHSR | GHSR Entrez, Source | growth hormone secretagogue receptor | 5557 | 0.007 | 0.3967 | No |
| 43 | ATP2C1 | ATP2C1 Entrez, Source | ATPase, Ca++ transporting, type 2C, member 1 | 5638 | 0.005 | 0.3912 | No |
| 44 | MATK | MATK Entrez, Source | megakaryocyte-associated tyrosine kinase | 6298 | -0.005 | 0.3421 | No |
| 45 | GLI1 | GLI1 Entrez, Source | glioma-associated oncogene homolog 1 (zinc finger protein) | 6332 | -0.005 | 0.3402 | No |
| 46 | TCF21 | TCF21 Entrez, Source | transcription factor 21 | 6717 | -0.011 | 0.3125 | No |
| 47 | CRABP2 | CRABP2 Entrez, Source | cellular retinoic acid binding protein 2 | 6879 | -0.013 | 0.3018 | No |
| 48 | ERCC3 | ERCC3 Entrez, Source | excision repair cross-complementing rodent repair deficiency, complementation group 3 (xeroderma pigmentosum group B complementing) | 7085 | -0.016 | 0.2882 | No |
| 49 | GHRL | GHRL Entrez, Source | ghrelin/obestatin preprohormone | 7162 | -0.018 | 0.2844 | No |
| 50 | CDK6 | CDK6 Entrez, Source | cyclin-dependent kinase 6 | 7356 | -0.020 | 0.2721 | No |
| 51 | DHCR24 | DHCR24 Entrez, Source | 24-dehydrocholesterol reductase | 7755 | -0.027 | 0.2450 | No |
| 52 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 7943 | -0.030 | 0.2342 | No |
| 53 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 7971 | -0.030 | 0.2355 | No |
| 54 | POU2F3 | POU2F3 Entrez, Source | POU domain, class 2, transcription factor 3 | 7981 | -0.030 | 0.2382 | No |
| 55 | SMAD2 | SMAD2 Entrez, Source | SMAD, mothers against DPP homolog 2 (Drosophila) | 8430 | -0.038 | 0.2086 | No |
| 56 | PTHLH | PTHLH Entrez, Source | parathyroid hormone-like hormone | 8761 | -0.043 | 0.1885 | No |
| 57 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 9075 | -0.048 | 0.1702 | No |
| 58 | TWIST2 | TWIST2 Entrez, Source | twist homolog 2 (Drosophila) | 9406 | -0.054 | 0.1513 | No |
| 59 | CASR | CASR Entrez, Source | calcium-sensing receptor (hypocalciuric hypercalcemia 1, severe neonatal hyperparathyroidism) | 9439 | -0.055 | 0.1550 | No |
| 60 | SECTM1 | SECTM1 Entrez, Source | secreted and transmembrane 1 | 9756 | -0.061 | 0.1379 | No |
| 61 | NME2 | NME2 Entrez, Source | non-metastatic cells 2, protein (NM23B) expressed in | 9773 | -0.061 | 0.1434 | No |
| 62 | HCK | HCK Entrez, Source | hemopoietic cell kinase | 9848 | -0.063 | 0.1448 | No |
| 63 | NF1 | NF1 Entrez, Source | neurofibromin 1 (neurofibromatosis, von Recklinghausen disease, Watson disease) | 10739 | -0.084 | 0.0869 | No |
| 64 | STX2 | STX2 Entrez, Source | syntaxin 2 | 11971 | -0.125 | 0.0079 | No |
| 65 | FABP5 | FABP5 Entrez, Source | fatty acid binding protein 5 (psoriasis-associated) | 12121 | -0.133 | 0.0113 | No |
| 66 | KLK7 | KLK7 Entrez, Source | kallikrein 7 (chymotryptic, stratum corneum) | 12591 | -0.162 | -0.0062 | No |
| 67 | LAMB3 | LAMB3 Entrez, Source | laminin, beta 3 | 12600 | -0.163 | 0.0111 | No |
| 68 | JAK2 | JAK2 Entrez, Source | Janus kinase 2 (a protein tyrosine kinase) | 13286 | -0.407 | 0.0042 | No |