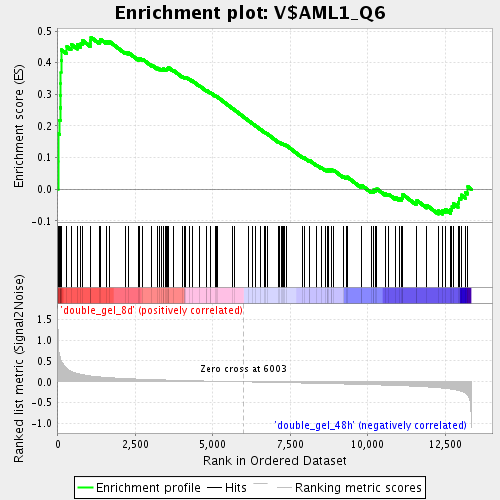
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$AML1_Q6 |
| Enrichment Score (ES) | 0.48036736 |
| Normalized Enrichment Score (NES) | 1.7891184 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.027611742 |
| FWER p-Value | 0.977 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CYP17A1 | CYP17A1 Entrez, Source | cytochrome P450, family 17, subfamily A, polypeptide 1 | 3 | 1.233 | 0.0886 | Yes |
| 2 | VIM | VIM Entrez, Source | vimentin | 4 | 1.225 | 0.1769 | Yes |
| 3 | COL4A1 | COL4A1 Entrez, Source | collagen, type IV, alpha 1 | 58 | 0.625 | 0.2179 | Yes |
| 4 | RGS1 | RGS1 Entrez, Source | regulator of G-protein signalling 1 | 75 | 0.565 | 0.2574 | Yes |
| 5 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 87 | 0.537 | 0.2953 | Yes |
| 6 | ANKRD1 | ANKRD1 Entrez, Source | ankyrin repeat domain 1 (cardiac muscle) | 93 | 0.529 | 0.3331 | Yes |
| 7 | RAB30 | RAB30 Entrez, Source | RAB30, member RAS oncogene family | 98 | 0.518 | 0.3701 | Yes |
| 8 | EMP1 | EMP1 Entrez, Source | epithelial membrane protein 1 | 101 | 0.510 | 0.4067 | Yes |
| 9 | GRK5 | GRK5 Entrez, Source | G protein-coupled receptor kinase 5 | 115 | 0.484 | 0.4406 | Yes |
| 10 | ANXA8 | ANXA8 Entrez, Source | annexin A8 | 289 | 0.316 | 0.4503 | Yes |
| 11 | PXN | PXN Entrez, Source | paxillin | 425 | 0.252 | 0.4583 | Yes |
| 12 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 627 | 0.202 | 0.4576 | Yes |
| 13 | NUCB2 | NUCB2 Entrez, Source | nucleobindin 2 | 743 | 0.177 | 0.4617 | Yes |
| 14 | POU2F1 | POU2F1 Entrez, Source | POU domain, class 2, transcription factor 1 | 797 | 0.170 | 0.4699 | Yes |
| 15 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 1043 | 0.142 | 0.4616 | Yes |
| 16 | DGKA | DGKA Entrez, Source | diacylglycerol kinase, alpha 80kDa | 1049 | 0.141 | 0.4715 | Yes |
| 17 | SLC37A4 | SLC37A4 Entrez, Source | solute carrier family 37 (glycerol-6-phosphate transporter), member 4 | 1065 | 0.139 | 0.4804 | Yes |
| 18 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 1345 | 0.117 | 0.4677 | No |
| 19 | PRICKLE1 | PRICKLE1 Entrez, Source | prickle homolog 1 (Drosophila) | 1377 | 0.115 | 0.4736 | No |
| 20 | RHOG | RHOG Entrez, Source | ras homolog gene family, member G (rho G) | 1566 | 0.104 | 0.4669 | No |
| 21 | IFNG | IFNG Entrez, Source | interferon, gamma | 1653 | 0.100 | 0.4676 | No |
| 22 | SLC6A9 | SLC6A9 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, glycine), member 9 | 2183 | 0.078 | 0.4332 | No |
| 23 | SNX1 | SNX1 Entrez, Source | sorting nexin 1 | 2279 | 0.075 | 0.4314 | No |
| 24 | PACS1 | PACS1 Entrez, Source | phosphofurin acidic cluster sorting protein 1 | 2608 | 0.065 | 0.4113 | No |
| 25 | SCN2B | SCN2B Entrez, Source | sodium channel, voltage-gated, type II, beta | 2628 | 0.064 | 0.4145 | No |
| 26 | RIN1 | RIN1 Entrez, Source | Ras and Rab interactor 1 | 2727 | 0.062 | 0.4115 | No |
| 27 | LMO3 | LMO3 Entrez, Source | LIM domain only 3 (rhombotin-like 2) | 3028 | 0.054 | 0.3927 | No |
| 28 | CRTAC1 | CRTAC1 Entrez, Source | cartilage acidic protein 1 | 3200 | 0.049 | 0.3834 | No |
| 29 | PTK9 | PTK9 Entrez, Source | PTK9 protein tyrosine kinase 9 | 3288 | 0.048 | 0.3802 | No |
| 30 | CHRNA10 | CHRNA10 Entrez, Source | cholinergic receptor, nicotinic, alpha 10 | 3338 | 0.046 | 0.3799 | No |
| 31 | DAB1 | DAB1 Entrez, Source | disabled homolog 1 (Drosophila) | 3399 | 0.045 | 0.3786 | No |
| 32 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 3412 | 0.045 | 0.3809 | No |
| 33 | LUZP1 | LUZP1 Entrez, Source | leucine zipper protein 1 | 3481 | 0.043 | 0.3789 | No |
| 34 | SCN8A | SCN8A Entrez, Source | sodium channel, voltage gated, type VIII, alpha | 3514 | 0.043 | 0.3796 | No |
| 35 | PDE6H | PDE6H Entrez, Source | phosphodiesterase 6H, cGMP-specific, cone, gamma | 3524 | 0.042 | 0.3819 | No |
| 36 | MAP3K11 | MAP3K11 Entrez, Source | mitogen-activated protein kinase kinase kinase 11 | 3563 | 0.042 | 0.3820 | No |
| 37 | MYOD1 | MYOD1 Entrez, Source | myogenic differentiation 1 | 3574 | 0.041 | 0.3843 | No |
| 38 | UBL3 | UBL3 Entrez, Source | ubiquitin-like 3 | 3727 | 0.038 | 0.3755 | No |
| 39 | DIABLO | DIABLO Entrez, Source | diablo homolog (Drosophila) | 4020 | 0.032 | 0.3558 | No |
| 40 | CAP1 | CAP1 Entrez, Source | CAP, adenylate cyclase-associated protein 1 (yeast) | 4084 | 0.031 | 0.3533 | No |
| 41 | CD69 | CD69 Entrez, Source | CD69 molecule | 4095 | 0.031 | 0.3548 | No |
| 42 | NGB | NGB Entrez, Source | neuroglobin | 4127 | 0.030 | 0.3546 | No |
| 43 | GPATC2 | GPATC2 Entrez, Source | G patch domain containing 2 | 4242 | 0.028 | 0.3480 | No |
| 44 | AGTRL1 | AGTRL1 Entrez, Source | angiotensin II receptor-like 1 | 4338 | 0.027 | 0.3428 | No |
| 45 | PSMA1 | PSMA1 Entrez, Source | proteasome (prosome, macropain) subunit, alpha type, 1 | 4575 | 0.022 | 0.3265 | No |
| 46 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 4794 | 0.019 | 0.3114 | No |
| 47 | MMP14 | MMP14 Entrez, Source | matrix metallopeptidase 14 (membrane-inserted) | 4808 | 0.018 | 0.3117 | No |
| 48 | ARF3 | ARF3 Entrez, Source | ADP-ribosylation factor 3 | 4912 | 0.017 | 0.3051 | No |
| 49 | XCL1 | XCL1 Entrez, Source | chemokine (C motif) ligand 1 | 4929 | 0.016 | 0.3051 | No |
| 50 | BLOC1S2 | BLOC1S2 Entrez, Source | biogenesis of lysosome-related organelles complex-1, subunit 2 | 5070 | 0.014 | 0.2955 | No |
| 51 | CD6 | CD6 Entrez, Source | CD6 molecule | 5109 | 0.013 | 0.2936 | No |
| 52 | PGF | PGF Entrez, Source | placental growth factor, vascular endothelial growth factor-related protein | 5143 | 0.012 | 0.2920 | No |
| 53 | PICALM | PICALM Entrez, Source | phosphatidylinositol binding clathrin assembly protein | 5636 | 0.005 | 0.2552 | No |
| 54 | NRAS | NRAS Entrez, Source | neuroblastoma RAS viral (v-ras) oncogene homolog | 5695 | 0.004 | 0.2511 | No |
| 55 | ACSL5 | ACSL5 Entrez, Source | acyl-CoA synthetase long-chain family member 5 | 6137 | -0.002 | 0.2179 | No |
| 56 | KCNJ1 | KCNJ1 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 1 | 6265 | -0.004 | 0.2086 | No |
| 57 | SCN3B | SCN3B Entrez, Source | sodium channel, voltage-gated, type III, beta | 6379 | -0.006 | 0.2005 | No |
| 58 | GZMB | GZMB Entrez, Source | granzyme B (granzyme 2, cytotoxic T-lymphocyte-associated serine esterase 1) | 6537 | -0.008 | 0.1892 | No |
| 59 | ID3 | ID3 Entrez, Source | inhibitor of DNA binding 3, dominant negative helix-loop-helix protein | 6682 | -0.010 | 0.1791 | No |
| 60 | PDE3B | PDE3B Entrez, Source | phosphodiesterase 3B, cGMP-inhibited | 6687 | -0.011 | 0.1795 | No |
| 61 | AKAP3 | AKAP3 Entrez, Source | A kinase (PRKA) anchor protein 3 | 6754 | -0.011 | 0.1754 | No |
| 62 | TPM1 | TPM1 Entrez, Source | tropomyosin 1 (alpha) | 7114 | -0.017 | 0.1495 | No |
| 63 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 7149 | -0.017 | 0.1482 | No |
| 64 | GRASP | GRASP Entrez, Source | GRP1 (general receptor for phosphoinositides 1)-associated scaffold protein | 7221 | -0.019 | 0.1441 | No |
| 65 | DNM3 | DNM3 Entrez, Source | dynamin 3 | 7241 | -0.019 | 0.1441 | No |
| 66 | CUGBP1 | CUGBP1 Entrez, Source | CUG triplet repeat, RNA binding protein 1 | 7294 | -0.019 | 0.1415 | No |
| 67 | MATN1 | MATN1 Entrez, Source | matrilin 1, cartilage matrix protein | 7327 | -0.020 | 0.1405 | No |
| 68 | TNNT2 | TNNT2 Entrez, Source | troponin T type 2 (cardiac) | 7380 | -0.021 | 0.1381 | No |
| 69 | DNAJC14 | DNAJC14 Entrez, Source | DnaJ (Hsp40) homolog, subfamily C, member 14 | 7890 | -0.029 | 0.1017 | No |
| 70 | ATP2A2 | ATP2A2 Entrez, Source | ATPase, Ca++ transporting, cardiac muscle, slow twitch 2 | 7943 | -0.030 | 0.0999 | No |
| 71 | CTSK | CTSK Entrez, Source | cathepsin K (pycnodysostosis) | 8117 | -0.032 | 0.0892 | No |
| 72 | ARHGEF2 | ARHGEF2 Entrez, Source | rho/rac guanine nucleotide exchange factor (GEF) 2 | 8130 | -0.033 | 0.0907 | No |
| 73 | ENTPD1 | ENTPD1 Entrez, Source | ectonucleoside triphosphate diphosphohydrolase 1 | 8353 | -0.037 | 0.0765 | No |
| 74 | IL23A | IL23A Entrez, Source | interleukin 23, alpha subunit p19 | 8504 | -0.039 | 0.0680 | No |
| 75 | MAMDC1 | MAMDC1 Entrez, Source | MAM domain containing 1 | 8637 | -0.041 | 0.0610 | No |
| 76 | LMO2 | LMO2 Entrez, Source | LIM domain only 2 (rhombotin-like 1) | 8708 | -0.042 | 0.0587 | No |
| 77 | NDUFA9 | NDUFA9 Entrez, Source | NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 9, 39kDa | 8712 | -0.042 | 0.0616 | No |
| 78 | TBX5 | TBX5 Entrez, Source | T-box 5 | 8730 | -0.043 | 0.0634 | No |
| 79 | ACIN1 | ACIN1 Entrez, Source | apoptotic chromatin condensation inducer 1 | 8813 | -0.044 | 0.0603 | No |
| 80 | SLC2A3 | SLC2A3 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 3 | 8829 | -0.044 | 0.0624 | No |
| 81 | STAT2 | STAT2 Entrez, Source | signal transducer and activator of transcription 2, 113kDa | 8894 | -0.045 | 0.0608 | No |
| 82 | SLC12A6 | SLC12A6 Entrez, Source | solute carrier family 12 (potassium/chloride transporters), member 6 | 9210 | -0.051 | 0.0407 | No |
| 83 | TITF1 | TITF1 Entrez, Source | thyroid transcription factor 1 | 9298 | -0.052 | 0.0379 | No |
| 84 | FGF4 | FGF4 Entrez, Source | fibroblast growth factor 4 (heparin secretory transforming protein 1, Kaposi sarcoma oncogene) | 9333 | -0.053 | 0.0392 | No |
| 85 | S100A9 | S100A9 Entrez, Source | S100 calcium binding protein A9 | 9785 | -0.062 | 0.0095 | No |
| 86 | SEMA4A | SEMA4A Entrez, Source | sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4A | 9792 | -0.062 | 0.0135 | No |
| 87 | TACC2 | TACC2 Entrez, Source | transforming, acidic coiled-coil containing protein 2 | 10110 | -0.069 | -0.0055 | No |
| 88 | FGD4 | FGD4 Entrez, Source | FYVE, RhoGEF and PH domain containing 4 | 10187 | -0.070 | -0.0062 | No |
| 89 | SCNN1A | SCNN1A Entrez, Source | sodium channel, nonvoltage-gated 1 alpha | 10199 | -0.071 | -0.0019 | No |
| 90 | GPD1 | GPD1 Entrez, Source | glycerol-3-phosphate dehydrogenase 1 (soluble) | 10261 | -0.072 | -0.0014 | No |
| 91 | JMJD1C | JMJD1C Entrez, Source | jumonji domain containing 1C | 10296 | -0.072 | 0.0013 | No |
| 92 | BTG4 | BTG4 Entrez, Source | B-cell translocation gene 4 | 10567 | -0.079 | -0.0134 | No |
| 93 | PAX6 | PAX6 Entrez, Source | paired box gene 6 (aniridia, keratitis) | 10670 | -0.082 | -0.0152 | No |
| 94 | INPPL1 | INPPL1 Entrez, Source | inositol polyphosphate phosphatase-like 1 | 10908 | -0.088 | -0.0268 | No |
| 95 | TNFSF4 | TNFSF4 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 4 (tax-transcriptionally activated glycoprotein 1, 34kDa) | 11022 | -0.091 | -0.0288 | No |
| 96 | EIF2B3 | EIF2B3 Entrez, Source | eukaryotic translation initiation factor 2B, subunit 3 gamma, 58kDa | 11098 | -0.093 | -0.0277 | No |
| 97 | TACSTD2 | TACSTD2 Entrez, Source | tumor-associated calcium signal transducer 2 | 11118 | -0.094 | -0.0224 | No |
| 98 | NRXN3 | NRXN3 Entrez, Source | neurexin 3 | 11130 | -0.094 | -0.0164 | No |
| 99 | HDGF | HDGF Entrez, Source | hepatoma-derived growth factor (high-mobility group protein 1-like) | 11565 | -0.108 | -0.0414 | No |
| 100 | ITGB7 | ITGB7 Entrez, Source | integrin, beta 7 | 11584 | -0.109 | -0.0350 | No |
| 101 | SLC15A3 | SLC15A3 Entrez, Source | solute carrier family 15, member 3 | 11905 | -0.122 | -0.0504 | No |
| 102 | PTPN7 | PTPN7 Entrez, Source | protein tyrosine phosphatase, non-receptor type 7 | 12273 | -0.141 | -0.0680 | No |
| 103 | BMF | BMF Entrez, Source | Bcl2 modifying factor | 12411 | -0.149 | -0.0676 | No |
| 104 | PDZK1 | PDZK1 Entrez, Source | PDZ domain containing 1 | 12501 | -0.155 | -0.0632 | No |
| 105 | ABCD4 | ABCD4 Entrez, Source | ATP-binding cassette, sub-family D (ALD), member 4 | 12676 | -0.171 | -0.0640 | No |
| 106 | TCF12 | TCF12 Entrez, Source | transcription factor 12 (HTF4, helix-loop-helix transcription factors 4) | 12718 | -0.175 | -0.0545 | No |
| 107 | NOTCH2 | NOTCH2 Entrez, Source | Notch homolog 2 (Drosophila) | 12769 | -0.180 | -0.0453 | No |
| 108 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 12915 | -0.202 | -0.0417 | No |
| 109 | VRK1 | VRK1 Entrez, Source | vaccinia related kinase 1 | 12945 | -0.208 | -0.0289 | No |
| 110 | GTF2A1 | GTF2A1 Entrez, Source | general transcription factor IIA, 1, 19/37kDa | 13010 | -0.220 | -0.0179 | No |
| 111 | ARNT | ARNT Entrez, Source | aryl hydrocarbon receptor nuclear translocator | 13148 | -0.269 | -0.0089 | No |
| 112 | KIF1B | KIF1B Entrez, Source | kinesin family member 1B | 13231 | -0.325 | 0.0084 | No |

