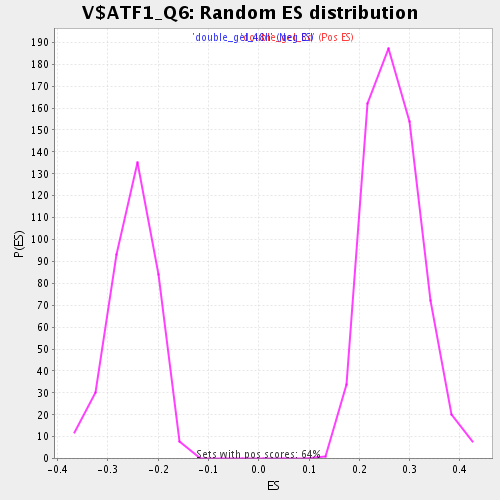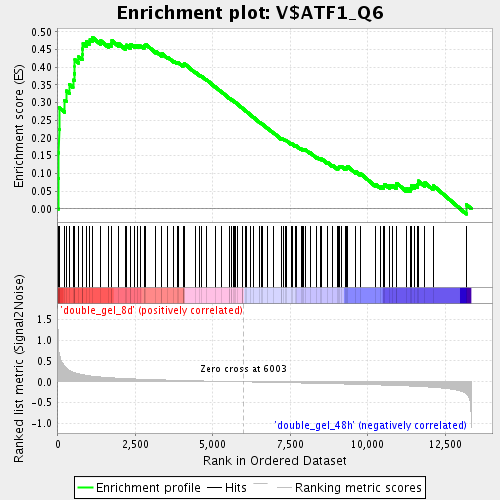
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$ATF1_Q6 |
| Enrichment Score (ES) | 0.48487118 |
| Normalized Enrichment Score (NES) | 1.8125873 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.023045538 |
| FWER p-Value | 0.947 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 9 | 0.936 | 0.0866 | Yes |
| 2 | CLDN7 | CLDN7 Entrez, Source | claudin 7 | 24 | 0.780 | 0.1583 | Yes |
| 3 | PCK1 | PCK1 Entrez, Source | phosphoenolpyruvate carboxykinase 1 (soluble) | 35 | 0.711 | 0.2239 | Yes |
| 4 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 43 | 0.675 | 0.2863 | Yes |
| 5 | HS3ST2 | HS3ST2 Entrez, Source | heparan sulfate (glucosamine) 3-O-sulfotransferase 2 | 219 | 0.370 | 0.3075 | Yes |
| 6 | NCALD | NCALD Entrez, Source | neurocalcin delta | 278 | 0.323 | 0.3332 | Yes |
| 7 | FOXP1 | FOXP1 Entrez, Source | forkhead box P1 | 375 | 0.270 | 0.3512 | Yes |
| 8 | TNFRSF12A | TNFRSF12A Entrez, Source | tumor necrosis factor receptor superfamily, member 12A | 489 | 0.231 | 0.3642 | Yes |
| 9 | SPRED2 | SPRED2 Entrez, Source | sprouty-related, EVH1 domain containing 2 | 528 | 0.221 | 0.3819 | Yes |
| 10 | PDGFA | PDGFA Entrez, Source | platelet-derived growth factor alpha polypeptide | 544 | 0.218 | 0.4011 | Yes |
| 11 | AQP7 | AQP7 Entrez, Source | aquaporin 7 | 548 | 0.217 | 0.4211 | Yes |
| 12 | COL1A1 | COL1A1 Entrez, Source | collagen, type I, alpha 1 | 673 | 0.190 | 0.4295 | Yes |
| 13 | FLT1 | FLT1 Entrez, Source | fms-related tyrosine kinase 1 (vascular endothelial growth factor/vascular permeability factor receptor) | 789 | 0.172 | 0.4368 | Yes |
| 14 | ARG2 | ARG2 Entrez, Source | arginase, type II | 799 | 0.170 | 0.4519 | Yes |
| 15 | LMO4 | LMO4 Entrez, Source | LIM domain only 4 | 808 | 0.169 | 0.4671 | Yes |
| 16 | BNIP3L | BNIP3L Entrez, Source | BCL2/adenovirus E1B 19kDa interacting protein 3-like | 921 | 0.154 | 0.4729 | Yes |
| 17 | PAK1 | PAK1 Entrez, Source | p21/Cdc42/Rac1-activated kinase 1 (STE20 homolog, yeast) | 1034 | 0.144 | 0.4778 | Yes |
| 18 | SLC38A1 | SLC38A1 Entrez, Source | solute carrier family 38, member 1 | 1109 | 0.135 | 0.4849 | Yes |
| 19 | CDX2 | CDX2 Entrez, Source | caudal type homeobox transcription factor 2 | 1384 | 0.114 | 0.4748 | No |
| 20 | PCSK1 | PCSK1 Entrez, Source | proprotein convertase subtilisin/kexin type 1 | 1629 | 0.101 | 0.4658 | No |
| 21 | HSD11B1 | HSD11B1 Entrez, Source | hydroxysteroid (11-beta) dehydrogenase 1 | 1720 | 0.096 | 0.4680 | No |
| 22 | MAP1LC3A | MAP1LC3A Entrez, Source | microtubule-associated protein 1 light chain 3 alpha | 1741 | 0.095 | 0.4754 | No |
| 23 | PER1 | PER1 Entrez, Source | period homolog 1 (Drosophila) | 1956 | 0.087 | 0.4673 | No |
| 24 | MAPK10 | MAPK10 Entrez, Source | mitogen-activated protein kinase 10 | 2176 | 0.078 | 0.4580 | No |
| 25 | TPM4 | TPM4 Entrez, Source | tropomyosin 4 | 2205 | 0.077 | 0.4631 | No |
| 26 | SNF1LK | SNF1LK Entrez, Source | SNF1-like kinase | 2345 | 0.073 | 0.4593 | No |
| 27 | BRE | BRE Entrez, Source | brain and reproductive organ-expressed (TNFRSF1A modulator) | 2350 | 0.072 | 0.4658 | No |
| 28 | CSF3 | CSF3 Entrez, Source | colony stimulating factor 3 (granulocyte) | 2483 | 0.069 | 0.4622 | No |
| 29 | CSF2 | CSF2 Entrez, Source | colony stimulating factor 2 (granulocyte-macrophage) | 2558 | 0.066 | 0.4628 | No |
| 30 | CCND1 | CCND1 Entrez, Source | cyclin D1 | 2668 | 0.063 | 0.4604 | No |
| 31 | ANGPT1 | ANGPT1 Entrez, Source | angiopoietin 1 | 2782 | 0.061 | 0.4575 | No |
| 32 | TRIB1 | TRIB1 Entrez, Source | tribbles homolog 1 (Drosophila) | 2792 | 0.060 | 0.4624 | No |
| 33 | RNF44 | RNF44 Entrez, Source | ring finger protein 44 | 2838 | 0.059 | 0.4646 | No |
| 34 | MAGED2 | MAGED2 Entrez, Source | melanoma antigen family D, 2 | 3164 | 0.050 | 0.4447 | No |
| 35 | ELL2 | ELL2 Entrez, Source | elongation factor, RNA polymerase II, 2 | 3352 | 0.046 | 0.4349 | No |
| 36 | MCAM | MCAM Entrez, Source | melanoma cell adhesion molecule | 3354 | 0.046 | 0.4391 | No |
| 37 | GNB4 | GNB4 Entrez, Source | guanine nucleotide binding protein (G protein), beta polypeptide 4 | 3549 | 0.042 | 0.4283 | No |
| 38 | GDNF | GDNF Entrez, Source | glial cell derived neurotrophic factor | 3737 | 0.038 | 0.4177 | No |
| 39 | JUND | JUND Entrez, Source | jun D proto-oncogene | 3846 | 0.036 | 0.4129 | No |
| 40 | DOK1 | DOK1 Entrez, Source | docking protein 1, 62kDa (downstream of tyrosine kinase 1) | 3881 | 0.035 | 0.4136 | No |
| 41 | ELOVL5 | ELOVL5 Entrez, Source | ELOVL family member 5, elongation of long chain fatty acids (FEN1/Elo2, SUR4/Elo3-like, yeast) | 4037 | 0.032 | 0.4049 | No |
| 42 | GPM6B | GPM6B Entrez, Source | glycoprotein M6B | 4058 | 0.032 | 0.4063 | No |
| 43 | KCNA6 | KCNA6 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 6 | 4066 | 0.032 | 0.4087 | No |
| 44 | ATP6V0C | ATP6V0C Entrez, Source | ATPase, H+ transporting, lysosomal 16kDa, V0 subunit c | 4083 | 0.031 | 0.4104 | No |
| 45 | GABRG2 | GABRG2 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, gamma 2 | 4437 | 0.025 | 0.3860 | No |
| 46 | PARD6A | PARD6A Entrez, Source | par-6 partitioning defective 6 homolog alpha (C.elegans) | 4580 | 0.022 | 0.3773 | No |
| 47 | SYT13 | SYT13 Entrez, Source | synaptotagmin XIII | 4642 | 0.021 | 0.3747 | No |
| 48 | CREM | CREM Entrez, Source | cAMP responsive element modulator | 4794 | 0.019 | 0.3650 | No |
| 49 | DNTTIP1 | DNTTIP1 Entrez, Source | deoxynucleotidyltransferase, terminal, interacting protein 1 | 5076 | 0.014 | 0.3450 | No |
| 50 | WISP1 | WISP1 Entrez, Source | WNT1 inducible signaling pathway protein 1 | 5281 | 0.010 | 0.3306 | No |
| 51 | LDHA | LDHA Entrez, Source | lactate dehydrogenase A | 5536 | 0.007 | 0.3120 | No |
| 52 | CDC42 | CDC42 Entrez, Source | cell division cycle 42 (GTP binding protein, 25kDa) | 5548 | 0.007 | 0.3118 | No |
| 53 | SNAP25 | SNAP25 Entrez, Source | synaptosomal-associated protein, 25kDa | 5609 | 0.006 | 0.3078 | No |
| 54 | SCG2 | SCG2 Entrez, Source | secretogranin II (chromogranin C) | 5614 | 0.005 | 0.3080 | No |
| 55 | CBX8 | CBX8 Entrez, Source | chromobox homolog 8 (Pc class homolog, Drosophila) | 5675 | 0.005 | 0.3039 | No |
| 56 | EHD1 | EHD1 Entrez, Source | EH-domain containing 1 | 5701 | 0.004 | 0.3024 | No |
| 57 | IKBKB | IKBKB Entrez, Source | inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase beta | 5717 | 0.004 | 0.3017 | No |
| 58 | HOXA4 | HOXA4 Entrez, Source | homeobox A4 | 5806 | 0.003 | 0.2953 | No |
| 59 | ADARB2 | ADARB2 Entrez, Source | adenosine deaminase, RNA-specific, B2 (RED2 homolog rat) | 5942 | 0.001 | 0.2851 | No |
| 60 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 5955 | 0.001 | 0.2843 | No |
| 61 | CALD1 | CALD1 Entrez, Source | caldesmon 1 | 5970 | 0.001 | 0.2833 | No |
| 62 | CD37 | CD37 Entrez, Source | CD37 molecule | 6038 | -0.001 | 0.2783 | No |
| 63 | TOP1 | TOP1 Entrez, Source | topoisomerase (DNA) I | 6078 | -0.001 | 0.2754 | No |
| 64 | USP48 | USP48 Entrez, Source | ubiquitin specific peptidase 48 | 6200 | -0.003 | 0.2666 | No |
| 65 | ARL6IP2 | ARL6IP2 Entrez, Source | ADP-ribosylation factor-like 6 interacting protein 2 | 6315 | -0.005 | 0.2584 | No |
| 66 | FOXD3 | FOXD3 Entrez, Source | forkhead box D3 | 6499 | -0.008 | 0.2453 | No |
| 67 | KCNIP2 | KCNIP2 Entrez, Source | Kv channel interacting protein 2 | 6562 | -0.009 | 0.2414 | No |
| 68 | NFIL3 | NFIL3 Entrez, Source | nuclear factor, interleukin 3 regulated | 6595 | -0.009 | 0.2398 | No |
| 69 | TH | TH Entrez, Source | tyrosine hydroxylase | 6761 | -0.012 | 0.2284 | No |
| 70 | UCN | UCN Entrez, Source | urocortin | 6972 | -0.015 | 0.2139 | No |
| 71 | EIF4A2 | EIF4A2 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 2 | 7219 | -0.018 | 0.1971 | No |
| 72 | GRASP | GRASP Entrez, Source | GRP1 (general receptor for phosphoinositides 1)-associated scaffold protein | 7221 | -0.019 | 0.1987 | No |
| 73 | FGF6 | FGF6 Entrez, Source | fibroblast growth factor 6 | 7269 | -0.019 | 0.1969 | No |
| 74 | NRXN1 | NRXN1 Entrez, Source | neurexin 1 | 7337 | -0.020 | 0.1938 | No |
| 75 | CHGB | CHGB Entrez, Source | chromogranin B (secretogranin 1) | 7364 | -0.020 | 0.1937 | No |
| 76 | SOX11 | SOX11 Entrez, Source | SRY (sex determining region Y)-box 11 | 7530 | -0.023 | 0.1834 | No |
| 77 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 7557 | -0.023 | 0.1836 | No |
| 78 | NOVA1 | NOVA1 Entrez, Source | neuro-oncological ventral antigen 1 | 7658 | -0.025 | 0.1784 | No |
| 79 | THRA | THRA Entrez, Source | thyroid hormone receptor, alpha (erythroblastic leukemia viral (v-erb-a) oncogene homolog, avian) | 7692 | -0.026 | 0.1783 | No |
| 80 | EEF2 | EEF2 Entrez, Source | eukaryotic translation elongation factor 2 | 7846 | -0.028 | 0.1693 | No |
| 81 | PPARGC1B | PPARGC1B Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, beta | 7905 | -0.029 | 0.1677 | No |
| 82 | LHX1 | LHX1 Entrez, Source | LIM homeobox 1 | 7941 | -0.030 | 0.1678 | No |
| 83 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 7982 | -0.030 | 0.1676 | No |
| 84 | ESRRG | ESRRG Entrez, Source | estrogen-related receptor gamma | 8140 | -0.033 | 0.1588 | No |
| 85 | VGF | VGF Entrez, Source | VGF nerve growth factor inducible | 8355 | -0.037 | 0.1461 | No |
| 86 | PDE1A | PDE1A Entrez, Source | phosphodiesterase 1A, calmodulin-dependent | 8459 | -0.038 | 0.1418 | No |
| 87 | PTPRU | PTPRU Entrez, Source | protein tyrosine phosphatase, receptor type, U | 8514 | -0.039 | 0.1414 | No |
| 88 | DYRK3 | DYRK3 Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 3 | 8702 | -0.042 | 0.1312 | No |
| 89 | SST | SST Entrez, Source | somatostatin | 8868 | -0.045 | 0.1229 | No |
| 90 | RNPS1 | RNPS1 Entrez, Source | RNA binding protein S1, serine-rich domain | 9038 | -0.048 | 0.1146 | No |
| 91 | CACNA1G | CACNA1G Entrez, Source | calcium channel, voltage-dependent, alpha 1G subunit | 9066 | -0.048 | 0.1171 | No |
| 92 | PPARD | PPARD Entrez, Source | peroxisome proliferative activated receptor, delta | 9075 | -0.048 | 0.1210 | No |
| 93 | SLC18A2 | SLC18A2 Entrez, Source | solute carrier family 18 (vesicular monoamine), member 2 | 9148 | -0.050 | 0.1202 | No |
| 94 | VIP | VIP Entrez, Source | vasoactive intestinal peptide | 9285 | -0.052 | 0.1148 | No |
| 95 | HOXC10 | HOXC10 Entrez, Source | homeobox C10 | 9320 | -0.053 | 0.1171 | No |
| 96 | ISL1 | ISL1 Entrez, Source | ISL1 transcription factor, LIM/homeodomain, (islet-1) | 9336 | -0.053 | 0.1210 | No |
| 97 | TRIM39 | TRIM39 Entrez, Source | tripartite motif-containing 39 | 9616 | -0.058 | 0.1053 | No |
| 98 | YWHAZ | YWHAZ Entrez, Source | tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide | 9753 | -0.061 | 0.1007 | No |
| 99 | KCNA5 | KCNA5 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 5 | 10246 | -0.071 | 0.0702 | No |
| 100 | RPL4 | RPL4 Entrez, Source | ribosomal protein L4 | 10426 | -0.075 | 0.0637 | No |
| 101 | SMAD6 | SMAD6 Entrez, Source | SMAD, mothers against DPP homolog 6 (Drosophila) | 10518 | -0.078 | 0.0641 | No |
| 102 | HAND1 | HAND1 Entrez, Source | heart and neural crest derivatives expressed 1 | 10537 | -0.079 | 0.0701 | No |
| 103 | PPM2C | PPM2C Entrez, Source | protein phosphatase 2C, magnesium-dependent, catalytic subunit | 10699 | -0.083 | 0.0656 | No |
| 104 | CREB3L2 | CREB3L2 Entrez, Source | cAMP responsive element binding protein 3-like 2 | 10782 | -0.085 | 0.0673 | No |
| 105 | HTR2C | HTR2C Entrez, Source | 5-hydroxytryptamine (serotonin) receptor 2C | 10920 | -0.088 | 0.0652 | No |
| 106 | PAFAH1B1 | PAFAH1B1 Entrez, Source | platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa | 10930 | -0.089 | 0.0728 | No |
| 107 | RET | RET Entrez, Source | ret proto-oncogene (multiple endocrine neoplasia and medullary thyroid carcinoma 1, Hirschsprung disease) | 11258 | -0.097 | 0.0572 | No |
| 108 | NTRK2 | NTRK2 Entrez, Source | neurotrophic tyrosine kinase, receptor, type 2 | 11369 | -0.101 | 0.0583 | No |
| 109 | BSND | BSND Entrez, Source | Bartter syndrome, infantile, with sensorineural deafness (Barttin) | 11402 | -0.102 | 0.0654 | No |
| 110 | SULT4A1 | SULT4A1 Entrez, Source | sulfotransferase family 4A, member 1 | 11515 | -0.106 | 0.0668 | No |
| 111 | AREG | AREG Entrez, Source | amphiregulin (schwannoma-derived growth factor) | 11612 | -0.110 | 0.0698 | No |
| 112 | CNN3 | CNN3 Entrez, Source | calponin 3, acidic | 11628 | -0.110 | 0.0790 | No |
| 113 | EIF4A1 | EIF4A1 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 1 | 11843 | -0.119 | 0.0739 | No |
| 114 | SCAMP5 | SCAMP5 Entrez, Source | secretory carrier membrane protein 5 | 12110 | -0.132 | 0.0661 | No |
| 115 | DDIT3 | DDIT3 Entrez, Source | DNA-damage-inducible transcript 3 | 13182 | -0.289 | 0.0121 | No |