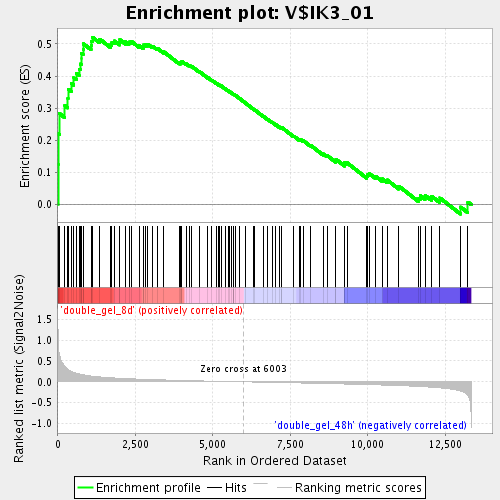
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$IK3_01 |
| Enrichment Score (ES) | 0.52041745 |
| Normalized Enrichment Score (NES) | 1.8445117 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.019013308 |
| FWER p-Value | 0.858 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CYP17A1 | CYP17A1 Entrez, Source | cytochrome P450, family 17, subfamily A, polypeptide 1 | 3 | 1.233 | 0.1257 | Yes |
| 2 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 9 | 0.936 | 0.2210 | Yes |
| 3 | CCL20 | CCL20 Entrez, Source | chemokine (C-C motif) ligand 20 | 50 | 0.641 | 0.2834 | Yes |
| 4 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 222 | 0.366 | 0.3079 | Yes |
| 5 | TYRO3 | TYRO3 Entrez, Source | TYRO3 protein tyrosine kinase | 321 | 0.292 | 0.3304 | Yes |
| 6 | TIMP3 | TIMP3 Entrez, Source | TIMP metallopeptidase inhibitor 3 (Sorsby fundus dystrophy, pseudoinflammatory) | 329 | 0.291 | 0.3595 | Yes |
| 7 | RHBDF1 | RHBDF1 Entrez, Source | rhomboid 5 homolog 1 (Drosophila) | 431 | 0.250 | 0.3775 | Yes |
| 8 | RGS5 | RGS5 Entrez, Source | regulator of G-protein signalling 5 | 513 | 0.224 | 0.3942 | Yes |
| 9 | CYR61 | CYR61 Entrez, Source | cysteine-rich, angiogenic inducer, 61 | 600 | 0.206 | 0.4088 | Yes |
| 10 | MKNK2 | MKNK2 Entrez, Source | MAP kinase interacting serine/threonine kinase 2 | 687 | 0.187 | 0.4214 | Yes |
| 11 | GABBR1 | GABBR1 Entrez, Source | gamma-aminobutyric acid (GABA) B receptor, 1 | 722 | 0.182 | 0.4375 | Yes |
| 12 | NFKBIA | NFKBIA Entrez, Source | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | 754 | 0.176 | 0.4531 | Yes |
| 13 | EGR1 | EGR1 Entrez, Source | early growth response 1 | 756 | 0.176 | 0.4710 | Yes |
| 14 | HCST | HCST Entrez, Source | hematopoietic cell signal transducer | 815 | 0.167 | 0.4837 | Yes |
| 15 | ESM1 | ESM1 Entrez, Source | endothelial cell-specific molecule 1 | 824 | 0.166 | 0.5001 | Yes |
| 16 | ADAM15 | ADAM15 Entrez, Source | ADAM metallopeptidase domain 15 (metargidin) | 1071 | 0.139 | 0.4957 | Yes |
| 17 | KIFC3 | KIFC3 Entrez, Source | kinesin family member C3 | 1087 | 0.137 | 0.5086 | Yes |
| 18 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 1114 | 0.135 | 0.5204 | Yes |
| 19 | IGSF4 | IGSF4 Entrez, Source | immunoglobulin superfamily, member 4 | 1345 | 0.117 | 0.5150 | No |
| 20 | DCX | DCX Entrez, Source | doublecortex; lissencephaly, X-linked (doublecortin) | 1708 | 0.097 | 0.4975 | No |
| 21 | MAP1LC3A | MAP1LC3A Entrez, Source | microtubule-associated protein 1 light chain 3 alpha | 1741 | 0.095 | 0.5049 | No |
| 22 | CEPT1 | CEPT1 Entrez, Source | choline/ethanolamine phosphotransferase 1 | 1813 | 0.092 | 0.5089 | No |
| 23 | SEMA3A | SEMA3A Entrez, Source | sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A | 1982 | 0.086 | 0.5050 | No |
| 24 | RAMP2 | RAMP2 Entrez, Source | receptor (calcitonin) activity modifying protein 2 | 1999 | 0.085 | 0.5125 | No |
| 25 | SLC6A9 | SLC6A9 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, glycine), member 9 | 2183 | 0.078 | 0.5066 | No |
| 26 | FGF10 | FGF10 Entrez, Source | fibroblast growth factor 10 | 2295 | 0.074 | 0.5058 | No |
| 27 | SNX16 | SNX16 Entrez, Source | sorting nexin 16 | 2361 | 0.072 | 0.5083 | No |
| 28 | SERPINI1 | SERPINI1 Entrez, Source | serpin peptidase inhibitor, clade I (neuroserpin), member 1 | 2621 | 0.064 | 0.4953 | No |
| 29 | EGR3 | EGR3 Entrez, Source | early growth response 3 | 2754 | 0.061 | 0.4916 | No |
| 30 | RAB11B | RAB11B Entrez, Source | RAB11B, member RAS oncogene family | 2773 | 0.061 | 0.4965 | No |
| 31 | C7ORF10 | C7ORF10 Entrez, Source | chromosome 7 open reading frame 10 | 2833 | 0.059 | 0.4981 | No |
| 32 | VAMP3 | VAMP3 Entrez, Source | vesicle-associated membrane protein 3 (cellubrevin) | 2906 | 0.057 | 0.4985 | No |
| 33 | TRAPPC3 | TRAPPC3 Entrez, Source | trafficking protein particle complex 3 | 3049 | 0.053 | 0.4932 | No |
| 34 | ACTN1 | ACTN1 Entrez, Source | actinin, alpha 1 | 3213 | 0.049 | 0.4859 | No |
| 35 | ANK3 | ANK3 Entrez, Source | ankyrin 3, node of Ranvier (ankyrin G) | 3412 | 0.045 | 0.4755 | No |
| 36 | ACCN2 | ACCN2 Entrez, Source | amiloride-sensitive cation channel 2, neuronal | 3921 | 0.034 | 0.4407 | No |
| 37 | MAG | MAG Entrez, Source | myelin associated glycoprotein | 3969 | 0.033 | 0.4405 | No |
| 38 | KCNJ13 | KCNJ13 Entrez, Source | potassium inwardly-rectifying channel, subfamily J, member 13 | 3972 | 0.033 | 0.4438 | No |
| 39 | VSNL1 | VSNL1 Entrez, Source | visinin-like 1 | 4001 | 0.033 | 0.4450 | No |
| 40 | NAPG | NAPG Entrez, Source | N-ethylmaleimide-sensitive factor attachment protein, gamma | 4138 | 0.030 | 0.4379 | No |
| 41 | FLNC | FLNC Entrez, Source | filamin C, gamma (actin binding protein 280) | 4230 | 0.029 | 0.4339 | No |
| 42 | SIRT6 | SIRT6 Entrez, Source | sirtuin (silent mating type information regulation 2 homolog) 6 (S. cerevisiae) | 4310 | 0.027 | 0.4307 | No |
| 43 | CDK9 | CDK9 Entrez, Source | cyclin-dependent kinase 9 (CDC2-related kinase) | 4558 | 0.022 | 0.4144 | No |
| 44 | PROK2 | PROK2 Entrez, Source | prokineticin 2 | 4842 | 0.018 | 0.3948 | No |
| 45 | ACE2 | ACE2 Entrez, Source | angiotensin I converting enzyme (peptidyl-dipeptidase A) 2 | 4970 | 0.016 | 0.3868 | No |
| 46 | GATA3 | GATA3 Entrez, Source | GATA binding protein 3 | 5124 | 0.013 | 0.3766 | No |
| 47 | SMPX | SMPX Entrez, Source | small muscle protein, X-linked | 5198 | 0.012 | 0.3723 | No |
| 48 | USP52 | USP52 Entrez, Source | ubiquitin specific peptidase 52 | 5230 | 0.011 | 0.3711 | No |
| 49 | ASCL3 | ASCL3 Entrez, Source | achaete-scute complex (Drosophila) homolog-like 3 | 5263 | 0.011 | 0.3698 | No |
| 50 | CNTN4 | CNTN4 Entrez, Source | contactin 4 | 5393 | 0.009 | 0.3609 | No |
| 51 | FGF20 | FGF20 Entrez, Source | fibroblast growth factor 20 | 5495 | 0.007 | 0.3541 | No |
| 52 | CHRNB2 | CHRNB2 Entrez, Source | cholinergic receptor, nicotinic, beta 2 (neuronal) | 5551 | 0.007 | 0.3506 | No |
| 53 | CXCL5 | CXCL5 Entrez, Source | chemokine (C-X-C motif) ligand 5 | 5599 | 0.006 | 0.3476 | No |
| 54 | AMBN | AMBN Entrez, Source | ameloblastin (enamel matrix protein) | 5664 | 0.005 | 0.3433 | No |
| 55 | CBX8 | CBX8 Entrez, Source | chromobox homolog 8 (Pc class homolog, Drosophila) | 5675 | 0.005 | 0.3430 | No |
| 56 | WSB2 | WSB2 Entrez, Source | WD repeat and SOCS box-containing 2 | 5679 | 0.005 | 0.3433 | No |
| 57 | DIO2 | DIO2 Entrez, Source | deiodinase, iodothyronine, type II | 5720 | 0.004 | 0.3407 | No |
| 58 | GABRB1 | GABRB1 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 1 | 5863 | 0.002 | 0.3301 | No |
| 59 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 6060 | -0.001 | 0.3155 | No |
| 60 | RBBP9 | RBBP9 Entrez, Source | retinoblastoma binding protein 9 | 6318 | -0.005 | 0.2966 | No |
| 61 | PTGES | PTGES Entrez, Source | prostaglandin E synthase | 6358 | -0.006 | 0.2942 | No |
| 62 | KCNA4 | KCNA4 Entrez, Source | potassium voltage-gated channel, shaker-related subfamily, member 4 | 6619 | -0.009 | 0.2755 | No |
| 63 | APBA1 | APBA1 Entrez, Source | amyloid beta (A4) precursor protein-binding, family A, member 1 (X11) | 6771 | -0.012 | 0.2654 | No |
| 64 | OTX1 | OTX1 Entrez, Source | orthodenticle homolog 1 (Drosophila) | 6913 | -0.014 | 0.2561 | No |
| 65 | SLC38A5 | SLC38A5 Entrez, Source | solute carrier family 38, member 5 | 7031 | -0.016 | 0.2489 | No |
| 66 | MADD | MADD Entrez, Source | MAP-kinase activating death domain | 7149 | -0.017 | 0.2418 | No |
| 67 | EIF4A2 | EIF4A2 Entrez, Source | eukaryotic translation initiation factor 4A, isoform 2 | 7219 | -0.018 | 0.2385 | No |
| 68 | PPP1CB | PPP1CB Entrez, Source | protein phosphatase 1, catalytic subunit, beta isoform | 7228 | -0.019 | 0.2398 | No |
| 69 | HMGA2 | HMGA2 Entrez, Source | high mobility group AT-hook 2 | 7610 | -0.024 | 0.2135 | No |
| 70 | C2 | C2 Entrez, Source | complement component 2 | 7795 | -0.027 | 0.2024 | No |
| 71 | HCN3 | HCN3 Entrez, Source | hyperpolarization activated cyclic nucleotide-gated potassium channel 3 | 7843 | -0.028 | 0.2017 | No |
| 72 | GNAO1 | GNAO1 Entrez, Source | guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O | 7919 | -0.029 | 0.1991 | No |
| 73 | MSX1 | MSX1 Entrez, Source | msh homeobox homolog 1 (Drosophila) | 8144 | -0.033 | 0.1855 | No |
| 74 | TMC4 | TMC4 Entrez, Source | transmembrane channel-like 4 | 8565 | -0.040 | 0.1579 | No |
| 75 | SLIT3 | SLIT3 Entrez, Source | slit homolog 3 (Drosophila) | 8691 | -0.042 | 0.1527 | No |
| 76 | TCF8 | TCF8 Entrez, Source | transcription factor 8 (represses interleukin 2 expression) | 8965 | -0.047 | 0.1369 | No |
| 77 | PRRX1 | PRRX1 Entrez, Source | paired related homeobox 1 | 8971 | -0.047 | 0.1413 | No |
| 78 | PSD | PSD Entrez, Source | pleckstrin and Sec7 domain containing | 9254 | -0.052 | 0.1253 | No |
| 79 | KCNH2 | KCNH2 Entrez, Source | potassium voltage-gated channel, subfamily H (eag-related), member 2 | 9256 | -0.052 | 0.1305 | No |
| 80 | CHAD | CHAD Entrez, Source | chondroadherin | 9330 | -0.053 | 0.1304 | No |
| 81 | UBXD2 | UBXD2 Entrez, Source | UBX domain containing 2 | 9969 | -0.066 | 0.0890 | No |
| 82 | PPP2R2B | PPP2R2B Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), beta isoform | 10000 | -0.066 | 0.0935 | No |
| 83 | SLC30A3 | SLC30A3 Entrez, Source | solute carrier family 30 (zinc transporter), member 3 | 10045 | -0.067 | 0.0971 | No |
| 84 | TNNI2 | TNNI2 Entrez, Source | troponin I type 2 (skeletal, fast) | 10252 | -0.072 | 0.0888 | No |
| 85 | LIFR | LIFR Entrez, Source | leukemia inhibitory factor receptor alpha | 10469 | -0.077 | 0.0804 | No |
| 86 | DHRS4 | DHRS4 Entrez, Source | dehydrogenase/reductase (SDR family) member 4 | 10624 | -0.080 | 0.0770 | No |
| 87 | FGF17 | FGF17 Entrez, Source | fibroblast growth factor 17 | 11001 | -0.090 | 0.0579 | No |
| 88 | RHOQ | RHOQ Entrez, Source | ras homolog gene family, member Q | 11646 | -0.111 | 0.0206 | No |
| 89 | SYNPR | SYNPR Entrez, Source | synaptoporin | 11694 | -0.113 | 0.0286 | No |
| 90 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 11850 | -0.119 | 0.0291 | No |
| 91 | RBX1 | RBX1 Entrez, Source | ring-box 1 | 12058 | -0.129 | 0.0267 | No |
| 92 | MECP2 | MECP2 Entrez, Source | methyl CpG binding protein 2 (Rett syndrome) | 12329 | -0.144 | 0.0210 | No |
| 93 | XPO1 | XPO1 Entrez, Source | exportin 1 (CRM1 homolog, yeast) | 12994 | -0.216 | -0.0070 | No |
| 94 | IGFALS | IGFALS Entrez, Source | insulin-like growth factor binding protein, acid labile subunit | 13230 | -0.325 | 0.0085 | No |

