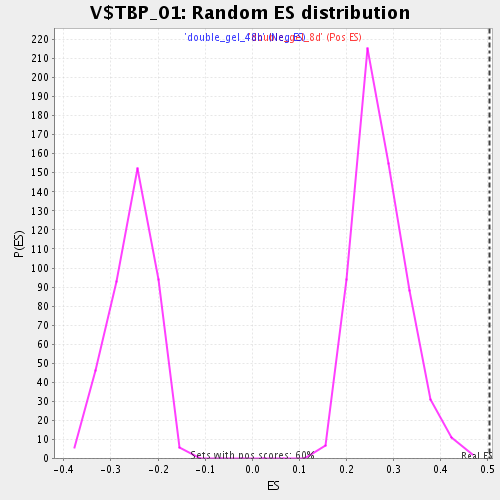
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_48h.class.cls #double_gel_8d_versus_double_gel_48h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_48h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | V$TBP_01 |
| Enrichment Score (ES) | 0.503625 |
| Normalized Enrichment Score (NES) | 1.8518142 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.01814381 |
| FWER p-Value | 0.839 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 6 | 1.154 | 0.0944 | Yes |
| 2 | FOS | FOS Entrez, Source | v-fos FBJ murine osteosarcoma viral oncogene homolog | 9 | 0.936 | 0.1712 | Yes |
| 3 | SLC25A4 | SLC25A4 Entrez, Source | solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 4 | 14 | 0.830 | 0.2391 | Yes |
| 4 | EGR2 | EGR2 Entrez, Source | early growth response 2 (Krox-20 homolog, Drosophila) | 87 | 0.537 | 0.2779 | Yes |
| 5 | TRPM4 | TRPM4 Entrez, Source | transient receptor potential cation channel, subfamily M, member 4 | 137 | 0.461 | 0.3120 | Yes |
| 6 | P2RX5 | P2RX5 Entrez, Source | purinergic receptor P2X, ligand-gated ion channel, 5 | 205 | 0.383 | 0.3384 | Yes |
| 7 | SCHIP1 | SCHIP1 Entrez, Source | schwannomin interacting protein 1 | 231 | 0.364 | 0.3665 | Yes |
| 8 | SLC3A1 | SLC3A1 Entrez, Source | 1 | 374 | 0.271 | 0.3780 | Yes |
| 9 | NR0B2 | NR0B2 Entrez, Source | nuclear receptor subfamily 0, group B, member 2 | 485 | 0.232 | 0.3887 | Yes |
| 10 | FLI1 | FLI1 Entrez, Source | Friend leukemia virus integration 1 | 627 | 0.202 | 0.3946 | Yes |
| 11 | TEF | TEF Entrez, Source | thyrotrophic embryonic factor | 701 | 0.186 | 0.4044 | Yes |
| 12 | SSH3 | SSH3 Entrez, Source | slingshot homolog 3 (Drosophila) | 792 | 0.170 | 0.4116 | Yes |
| 13 | CKB | CKB Entrez, Source | creatine kinase, brain | 820 | 0.166 | 0.4232 | Yes |
| 14 | PRKAG1 | PRKAG1 Entrez, Source | protein kinase, AMP-activated, gamma 1 non-catalytic subunit | 942 | 0.152 | 0.4266 | Yes |
| 15 | S100A4 | S100A4 Entrez, Source | S100 calcium binding protein A4 | 962 | 0.151 | 0.4375 | Yes |
| 16 | NR4A1 | NR4A1 Entrez, Source | nuclear receptor subfamily 4, group A, member 1 | 1043 | 0.142 | 0.4432 | Yes |
| 17 | TAX1BP3 | TAX1BP3 Entrez, Source | Tax1 (human T-cell leukemia virus type I) binding protein 3 | 1166 | 0.131 | 0.4447 | Yes |
| 18 | PPARGC1A | PPARGC1A Entrez, Source | peroxisome proliferative activated receptor, gamma, coactivator 1, alpha | 1181 | 0.130 | 0.4543 | Yes |
| 19 | AMPD1 | AMPD1 Entrez, Source | adenosine monophosphate deaminase 1 (isoform M) | 1199 | 0.128 | 0.4635 | Yes |
| 20 | ACTA1 | ACTA1 Entrez, Source | actin, alpha 1, skeletal muscle | 1218 | 0.126 | 0.4725 | Yes |
| 21 | SH3GLB2 | SH3GLB2 Entrez, Source | SH3-domain GRB2-like endophilin B2 | 1376 | 0.115 | 0.4701 | Yes |
| 22 | GPSN2 | GPSN2 Entrez, Source | glycoprotein, synaptic 2 | 1434 | 0.112 | 0.4750 | Yes |
| 23 | HRC | HRC Entrez, Source | histidine rich calcium binding protein | 1486 | 0.109 | 0.4801 | Yes |
| 24 | PRKAA2 | PRKAA2 Entrez, Source | protein kinase, AMP-activated, alpha 2 catalytic subunit | 1575 | 0.104 | 0.4820 | Yes |
| 25 | HSPB3 | HSPB3 Entrez, Source | heat shock 27kDa protein 3 | 1739 | 0.096 | 0.4775 | Yes |
| 26 | FXYD1 | FXYD1 Entrez, Source | FXYD domain containing ion transport regulator 1 (phospholemman) | 1781 | 0.094 | 0.4821 | Yes |
| 27 | CDH20 | CDH20 Entrez, Source | cadherin 20, type 2 | 1782 | 0.094 | 0.4898 | Yes |
| 28 | SEMA4B | SEMA4B Entrez, Source | sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4B | 1786 | 0.093 | 0.4973 | Yes |
| 29 | BRAF | BRAF Entrez, Source | v-raf murine sarcoma viral oncogene homolog B1 | 1804 | 0.093 | 0.5036 | Yes |
| 30 | HDAC7A | HDAC7A Entrez, Source | histone deacetylase 7A | 1959 | 0.087 | 0.4991 | No |
| 31 | ABCC6 | ABCC6 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 6 | 2073 | 0.082 | 0.4973 | No |
| 32 | CHRDL1 | CHRDL1 Entrez, Source | chordin-like 1 | 2170 | 0.078 | 0.4965 | No |
| 33 | ZFP36L1 | ZFP36L1 Entrez, Source | zinc finger protein 36, C3H type-like 1 | 2445 | 0.070 | 0.4815 | No |
| 34 | GIT1 | GIT1 Entrez, Source | G protein-coupled receptor kinase interactor 1 | 2623 | 0.064 | 0.4734 | No |
| 35 | TRDN | TRDN Entrez, Source | triadin | 2851 | 0.059 | 0.4611 | No |
| 36 | DES | DES Entrez, Source | desmin | 2879 | 0.058 | 0.4639 | No |
| 37 | HIST1H2BA | HIST1H2BA Entrez, Source | histone cluster 1, H2ba | 3230 | 0.049 | 0.4414 | No |
| 38 | CYP26B1 | CYP26B1 Entrez, Source | cytochrome P450, family 26, subfamily B, polypeptide 1 | 3231 | 0.049 | 0.4454 | No |
| 39 | ASB16 | ASB16 Entrez, Source | ankyrin repeat and SOCS box-containing 16 | 3250 | 0.048 | 0.4480 | No |
| 40 | DNM2 | DNM2 Entrez, Source | dynamin 2 | 3302 | 0.047 | 0.4481 | No |
| 41 | GRIK4 | GRIK4 Entrez, Source | glutamate receptor, ionotropic, kainate 4 | 3356 | 0.046 | 0.4479 | No |
| 42 | GPR153 | GPR153 Entrez, Source | G protein-coupled receptor 153 | 3535 | 0.042 | 0.4379 | No |
| 43 | SLC2A4 | SLC2A4 Entrez, Source | solute carrier family 2 (facilitated glucose transporter), member 4 | 3677 | 0.039 | 0.4304 | No |
| 44 | GABRB2 | GABRB2 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, beta 2 | 3779 | 0.037 | 0.4258 | No |
| 45 | MYH8 | MYH8 Entrez, Source | myosin, heavy chain 8, skeletal muscle, perinatal | 3786 | 0.037 | 0.4284 | No |
| 46 | CSNK1E | CSNK1E Entrez, Source | casein kinase 1, epsilon | 4346 | 0.026 | 0.3883 | No |
| 47 | SULT2A1 | SULT2A1 Entrez, Source | sulfotransferase family, cytosolic, 2A, dehydroepiandrosterone (DHEA)-preferring, member 1 | 4352 | 0.026 | 0.3901 | No |
| 48 | FGF21 | FGF21 Entrez, Source | fibroblast growth factor 21 | 4452 | 0.024 | 0.3847 | No |
| 49 | TSC1 | TSC1 Entrez, Source | tuberous sclerosis 1 | 4477 | 0.024 | 0.3848 | No |
| 50 | TCF1 | TCF1 Entrez, Source | transcription factor 1, hepatic; LF-B1, hepatic nuclear factor (HNF1), albumin proximal factor | 4517 | 0.023 | 0.3837 | No |
| 51 | GALNT1 | GALNT1 Entrez, Source | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 1 (GalNAc-T1) | 4813 | 0.018 | 0.3630 | No |
| 52 | PTF1A | PTF1A Entrez, Source | pancreas specific transcription factor, 1a | 4886 | 0.017 | 0.3589 | No |
| 53 | DNAJA4 | DNAJA4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 4 | 5030 | 0.015 | 0.3493 | No |
| 54 | HAND2 | HAND2 Entrez, Source | heart and neural crest derivatives expressed 2 | 5313 | 0.010 | 0.3288 | No |
| 55 | AQP1 | AQP1 Entrez, Source | aquaporin 1 (Colton blood group) | 5400 | 0.009 | 0.3231 | No |
| 56 | EHD1 | EHD1 Entrez, Source | EH-domain containing 1 | 5701 | 0.004 | 0.3008 | No |
| 57 | SLC35C2 | SLC35C2 Entrez, Source | solute carrier family 35, member C2 | 5747 | 0.004 | 0.2976 | No |
| 58 | MAP2K5 | MAP2K5 Entrez, Source | mitogen-activated protein kinase kinase 5 | 5887 | 0.002 | 0.2873 | No |
| 59 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 6060 | -0.001 | 0.2744 | No |
| 60 | LUC7L | LUC7L Entrez, Source | LUC7-like (S. cerevisiae) | 6242 | -0.004 | 0.2610 | No |
| 61 | GCG | GCG Entrez, Source | glucagon | 6455 | -0.007 | 0.2456 | No |
| 62 | PAX3 | PAX3 Entrez, Source | paired box gene 3 (Waardenburg syndrome 1) | 6540 | -0.008 | 0.2399 | No |
| 63 | POFUT1 | POFUT1 Entrez, Source | protein O-fucosyltransferase 1 | 6632 | -0.010 | 0.2338 | No |
| 64 | PPP2R2A | PPP2R2A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B (PR 52), alpha isoform | 6690 | -0.011 | 0.2304 | No |
| 65 | THBS2 | THBS2 Entrez, Source | thrombospondin 2 | 6731 | -0.011 | 0.2283 | No |
| 66 | PNMA1 | PNMA1 Entrez, Source | paraneoplastic antigen MA1 | 6758 | -0.012 | 0.2273 | No |
| 67 | CTCF | CTCF Entrez, Source | CCCTC-binding factor (zinc finger protein) | 6862 | -0.013 | 0.2205 | No |
| 68 | ART5 | ART5 Entrez, Source | ADP-ribosyltransferase 5 | 6905 | -0.014 | 0.2185 | No |
| 69 | FGF5 | FGF5 Entrez, Source | fibroblast growth factor 5 | 6936 | -0.014 | 0.2174 | No |
| 70 | TTC17 | TTC17 Entrez, Source | tetratricopeptide repeat domain 17 | 7081 | -0.016 | 0.2078 | No |
| 71 | HAPLN1 | HAPLN1 Entrez, Source | hyaluronan and proteoglycan link protein 1 | 7300 | -0.020 | 0.1930 | No |
| 72 | MGAT3 | MGAT3 Entrez, Source | mannosyl (beta-1,4-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase | 7454 | -0.022 | 0.1832 | No |
| 73 | HIST1H2AA | HIST1H2AA Entrez, Source | histone cluster 1, H2aa | 7576 | -0.024 | 0.1760 | No |
| 74 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 7950 | -0.030 | 0.1503 | No |
| 75 | NCOR1 | NCOR1 Entrez, Source | nuclear receptor co-repressor 1 | 7954 | -0.030 | 0.1526 | No |
| 76 | ESRRG | ESRRG Entrez, Source | estrogen-related receptor gamma | 8140 | -0.033 | 0.1413 | No |
| 77 | NOG | NOG Entrez, Source | noggin | 8217 | -0.034 | 0.1384 | No |
| 78 | CRISP1 | CRISP1 Entrez, Source | cysteine-rich secretory protein 1 | 8354 | -0.037 | 0.1311 | No |
| 79 | AGT | AGT Entrez, Source | angiotensinogen (serpin peptidase inhibitor, clade A, member 8) | 8544 | -0.039 | 0.1201 | No |
| 80 | CLCN1 | CLCN1 Entrez, Source | chloride channel 1, skeletal muscle (Thomsen disease, autosomal dominant) | 8606 | -0.041 | 0.1188 | No |
| 81 | PACSIN3 | PACSIN3 Entrez, Source | protein kinase C and casein kinase substrate in neurons 3 | 8824 | -0.044 | 0.1060 | No |
| 82 | MYOCD | MYOCD Entrez, Source | myocardin | 9057 | -0.048 | 0.0925 | No |
| 83 | FHL2 | FHL2 Entrez, Source | four and a half LIM domains 2 | 9173 | -0.050 | 0.0879 | No |
| 84 | EPHA7 | EPHA7 Entrez, Source | EPH receptor A7 | 9340 | -0.053 | 0.0797 | No |
| 85 | LGI1 | LGI1 Entrez, Source | leucine-rich, glioma inactivated 1 | 9443 | -0.055 | 0.0766 | No |
| 86 | LHX5 | LHX5 Entrez, Source | LIM homeobox 5 | 9552 | -0.057 | 0.0731 | No |
| 87 | MYST2 | MYST2 Entrez, Source | MYST histone acetyltransferase 2 | 9642 | -0.059 | 0.0712 | No |
| 88 | RARA | RARA Entrez, Source | retinoic acid receptor, alpha | 9679 | -0.059 | 0.0733 | No |
| 89 | COL10A1 | COL10A1 Entrez, Source | collagen, type X, alpha 1(Schmid metaphyseal chondrodysplasia) | 9784 | -0.062 | 0.0705 | No |
| 90 | ITGA7 | ITGA7 Entrez, Source | integrin, alpha 7 | 9876 | -0.064 | 0.0689 | No |
| 91 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 10099 | -0.068 | 0.0577 | No |
| 92 | TUBA4 | TUBA4 Entrez, Source | tubulin, alpha 4 | 10207 | -0.071 | 0.0555 | No |
| 93 | MEOX2 | MEOX2 Entrez, Source | mesenchyme homeobox 2 | 10416 | -0.075 | 0.0459 | No |
| 94 | NR0B1 | NR0B1 Entrez, Source | nuclear receptor subfamily 0, group B, member 1 | 10457 | -0.077 | 0.0492 | No |
| 95 | PLEC1 | PLEC1 Entrez, Source | plectin 1, intermediate filament binding protein 500kDa | 10661 | -0.081 | 0.0405 | No |
| 96 | KPNA3 | KPNA3 Entrez, Source | karyopherin alpha 3 (importin alpha 4) | 10744 | -0.084 | 0.0413 | No |
| 97 | ART3 | ART3 Entrez, Source | ADP-ribosyltransferase 3 | 10863 | -0.087 | 0.0395 | No |
| 98 | IFNB1 | IFNB1 Entrez, Source | interferon, beta 1, fibroblast | 10923 | -0.089 | 0.0423 | No |
| 99 | ITGB3BP | ITGB3BP Entrez, Source | integrin beta 3 binding protein (beta3-endonexin) | 10965 | -0.089 | 0.0466 | No |
| 100 | RPS19 | RPS19 Entrez, Source | ribosomal protein S19 | 12113 | -0.132 | -0.0292 | No |
| 101 | NDRG2 | NDRG2 Entrez, Source | NDRG family member 2 | 12174 | -0.135 | -0.0226 | No |
| 102 | SOX9 | SOX9 Entrez, Source | SRY (sex determining region Y)-box 9 (campomelic dysplasia, autosomal sex-reversal) | 12407 | -0.149 | -0.0279 | No |
| 103 | NTF3 | NTF3 Entrez, Source | neurotrophin 3 | 12465 | -0.152 | -0.0197 | No |
| 104 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 12470 | -0.153 | -0.0074 | No |
| 105 | SLC30A1 | SLC30A1 Entrez, Source | solute carrier family 30 (zinc transporter), member 1 | 13129 | -0.260 | -0.0357 | No |
| 106 | MAT2A | MAT2A Entrez, Source | methionine adenosyltransferase II, alpha | 13326 | -0.630 | 0.0012 | No |