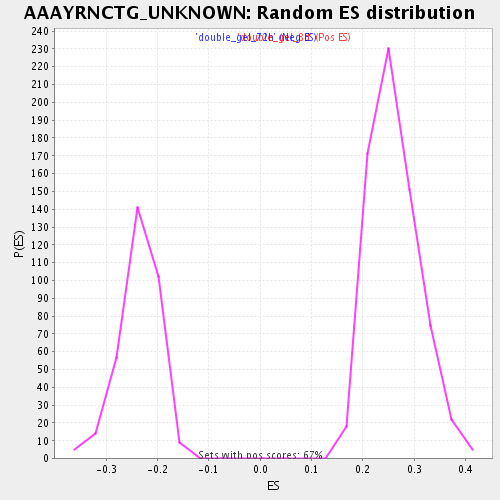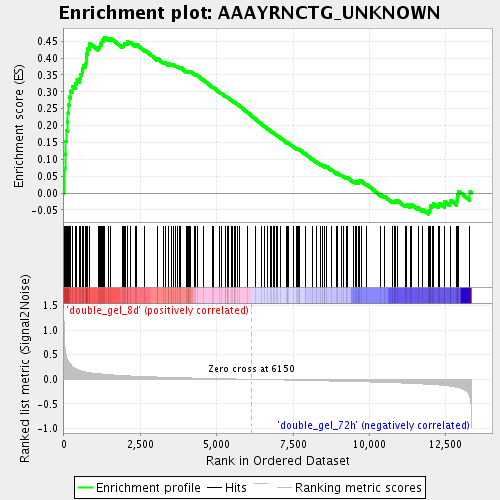
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | AAAYRNCTG_UNKNOWN |
| Enrichment Score (ES) | 0.46160728 |
| Normalized Enrichment Score (NES) | 1.7671963 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.037418775 |
| FWER p-Value | 0.996 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | COL1A2 | COL1A2 Entrez, Source | collagen, type I, alpha 2 | 3 | 1.046 | 0.0734 | Yes |
| 2 | TMEM27 | TMEM27 Entrez, Source | transmembrane protein 27 | 36 | 0.635 | 0.1157 | Yes |
| 3 | RAB30 | RAB30 Entrez, Source | RAB30, member RAS oncogene family | 55 | 0.564 | 0.1540 | Yes |
| 4 | RAPGEF4 | RAPGEF4 Entrez, Source | Rap guanine nucleotide exchange factor (GEF) 4 | 78 | 0.472 | 0.1856 | Yes |
| 5 | PRELP | PRELP Entrez, Source | proline/arginine-rich end leucine-rich repeat protein | 119 | 0.401 | 0.2108 | Yes |
| 6 | FST | FST Entrez, Source | follistatin | 132 | 0.388 | 0.2372 | Yes |
| 7 | CITED2 | CITED2 Entrez, Source | Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 2 | 145 | 0.367 | 0.2621 | Yes |
| 8 | CRAT | CRAT Entrez, Source | carnitine acetyltransferase | 170 | 0.348 | 0.2848 | Yes |
| 9 | PPP2R3A | PPP2R3A Entrez, Source | protein phosphatase 2 (formerly 2A), regulatory subunit B'', alpha | 222 | 0.305 | 0.3023 | Yes |
| 10 | DNAJB4 | DNAJB4 Entrez, Source | DnaJ (Hsp40) homolog, subfamily B, member 4 | 278 | 0.263 | 0.3167 | Yes |
| 11 | LTBP1 | LTBP1 Entrez, Source | latent transforming growth factor beta binding protein 1 | 372 | 0.219 | 0.3250 | Yes |
| 12 | TMSB4X | TMSB4X Entrez, Source | thymosin, beta 4, X-linked | 427 | 0.202 | 0.3351 | Yes |
| 13 | DDAH2 | DDAH2 Entrez, Source | dimethylarginine dimethylaminohydrolase 2 | 522 | 0.178 | 0.3405 | Yes |
| 14 | CRH | CRH Entrez, Source | corticotropin releasing hormone | 541 | 0.173 | 0.3514 | Yes |
| 15 | TEF | TEF Entrez, Source | thyrotrophic embryonic factor | 592 | 0.166 | 0.3593 | Yes |
| 16 | PXN | PXN Entrez, Source | paxillin | 617 | 0.163 | 0.3689 | Yes |
| 17 | FGF10 | FGF10 Entrez, Source | fibroblast growth factor 10 | 632 | 0.160 | 0.3791 | Yes |
| 18 | OTX2 | OTX2 Entrez, Source | orthodenticle homolog 2 (Drosophila) | 708 | 0.150 | 0.3840 | Yes |
| 19 | LMO7 | LMO7 Entrez, Source | LIM domain 7 | 728 | 0.147 | 0.3929 | Yes |
| 20 | PROK2 | PROK2 Entrez, Source | prokineticin 2 | 732 | 0.146 | 0.4030 | Yes |
| 21 | MYH10 | MYH10 Entrez, Source | myosin, heavy chain 10, non-muscle | 744 | 0.145 | 0.4123 | Yes |
| 22 | NCDN | NCDN Entrez, Source | neurochondrin | 774 | 0.142 | 0.4202 | Yes |
| 23 | KLF5 | KLF5 Entrez, Source | Kruppel-like factor 5 (intestinal) | 780 | 0.141 | 0.4297 | Yes |
| 24 | IHPK2 | IHPK2 Entrez, Source | inositol hexaphosphate kinase 2 | 835 | 0.136 | 0.4352 | Yes |
| 25 | FCER1A | FCER1A Entrez, Source | Fc fragment of IgE, high affinity I, receptor for; alpha polypeptide | 844 | 0.135 | 0.4441 | Yes |
| 26 | B4GALT6 | B4GALT6 Entrez, Source | UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 6 | 1117 | 0.114 | 0.4315 | Yes |
| 27 | CYBRD1 | CYBRD1 Entrez, Source | cytochrome b reductase 1 | 1178 | 0.111 | 0.4347 | Yes |
| 28 | WBP1 | WBP1 Entrez, Source | WW domain binding protein 1 | 1181 | 0.110 | 0.4423 | Yes |
| 29 | NR2F1 | NR2F1 Entrez, Source | nuclear receptor subfamily 2, group F, member 1 | 1234 | 0.108 | 0.4459 | Yes |
| 30 | ERBB4 | ERBB4 Entrez, Source | v-erb-a erythroblastic leukemia viral oncogene homolog 4 (avian) | 1249 | 0.107 | 0.4524 | Yes |
| 31 | MGAT1 | MGAT1 Entrez, Source | mannosyl (alpha-1,3-)-glycoprotein beta-1,2-N-acetylglucosaminyltransferase | 1288 | 0.105 | 0.4569 | Yes |
| 32 | GNAQ | GNAQ Entrez, Source | guanine nucleotide binding protein (G protein), q polypeptide | 1323 | 0.103 | 0.4616 | Yes |
| 33 | AQP2 | AQP2 Entrez, Source | aquaporin 2 (collecting duct) | 1456 | 0.096 | 0.4584 | No |
| 34 | MYF6 | MYF6 Entrez, Source | myogenic factor 6 (herculin) | 1536 | 0.092 | 0.4588 | No |
| 35 | SCN8A | SCN8A Entrez, Source | sodium channel, voltage gated, type VIII, alpha | 1920 | 0.076 | 0.4351 | No |
| 36 | FGF12 | FGF12 Entrez, Source | fibroblast growth factor 12 | 1947 | 0.075 | 0.4384 | No |
| 37 | NRG1 | NRG1 Entrez, Source | neuregulin 1 | 1967 | 0.074 | 0.4422 | No |
| 38 | STC2 | STC2 Entrez, Source | stanniocalcin 2 | 2009 | 0.073 | 0.4442 | No |
| 39 | TRDN | TRDN Entrez, Source | triadin | 2077 | 0.071 | 0.4441 | No |
| 40 | PRKCQ | PRKCQ Entrez, Source | protein kinase C, theta | 2087 | 0.070 | 0.4483 | No |
| 41 | DUSP4 | DUSP4 Entrez, Source | dual specificity phosphatase 4 | 2162 | 0.068 | 0.4475 | No |
| 42 | FKRP | FKRP Entrez, Source | fukutin related protein | 2335 | 0.064 | 0.4390 | No |
| 43 | DGKB | DGKB Entrez, Source | diacylglycerol kinase, beta 90kDa | 2364 | 0.063 | 0.4413 | No |
| 44 | CAPN1 | CAPN1 Entrez, Source | calpain 1, (mu/I) large subunit | 2648 | 0.056 | 0.4238 | No |
| 45 | VIP | VIP Entrez, Source | vasoactive intestinal peptide | 3062 | 0.047 | 0.3958 | No |
| 46 | MYH1 | MYH1 Entrez, Source | myosin, heavy chain 1, skeletal muscle, adult | 3074 | 0.047 | 0.3983 | No |
| 47 | PPAP2B | PPAP2B Entrez, Source | phosphatidic acid phosphatase type 2B | 3267 | 0.043 | 0.3867 | No |
| 48 | RTN1 | RTN1 Entrez, Source | reticulon 1 | 3317 | 0.042 | 0.3860 | No |
| 49 | GDNF | GDNF Entrez, Source | glial cell derived neurotrophic factor | 3413 | 0.040 | 0.3816 | No |
| 50 | PTHR1 | PTHR1 Entrez, Source | parathyroid hormone receptor 1 | 3437 | 0.040 | 0.3827 | No |
| 51 | PHEX | PHEX Entrez, Source | phosphate regulating endopeptidase homolog, X-linked (hypophosphatemia, vitamin D resistant rickets) | 3518 | 0.038 | 0.3793 | No |
| 52 | DHH | DHH Entrez, Source | desert hedgehog homolog (Drosophila) | 3533 | 0.038 | 0.3809 | No |
| 53 | ST7L | ST7L Entrez, Source | suppression of tumorigenicity 7 like | 3578 | 0.037 | 0.3802 | No |
| 54 | GJB4 | GJB4 Entrez, Source | gap junction protein, beta 4 (connexin 30.3) | 3665 | 0.036 | 0.3762 | No |
| 55 | PPP1R9B | PPP1R9B Entrez, Source | protein phosphatase 1, regulatory subunit 9B, spinophilin | 3726 | 0.035 | 0.3741 | No |
| 56 | SCN5A | SCN5A Entrez, Source | sodium channel, voltage-gated, type V, alpha (long QT syndrome 3) | 3779 | 0.034 | 0.3725 | No |
| 57 | WNT2B | WNT2B Entrez, Source | wingless-type MMTV integration site family, member 2B | 3814 | 0.033 | 0.3723 | No |
| 58 | RNF146 | RNF146 Entrez, Source | ring finger protein 146 | 4005 | 0.030 | 0.3600 | No |
| 59 | NFATC4 | NFATC4 Entrez, Source | nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 | 4051 | 0.029 | 0.3587 | No |
| 60 | SDCBP | SDCBP Entrez, Source | syndecan binding protein (syntenin) | 4064 | 0.029 | 0.3598 | No |
| 61 | CLDN5 | CLDN5 Entrez, Source | claudin 5 (transmembrane protein deleted in velocardiofacial syndrome) | 4087 | 0.029 | 0.3602 | No |
| 62 | CRYGD | CRYGD Entrez, Source | crystallin, gamma D | 4116 | 0.028 | 0.3601 | No |
| 63 | ZIC1 | ZIC1 Entrez, Source | Zic family member 1 (odd-paired homolog, Drosophila) | 4145 | 0.028 | 0.3599 | No |
| 64 | BCL2L1 | BCL2L1 Entrez, Source | BCL2-like 1 | 4270 | 0.026 | 0.3523 | No |
| 65 | EIF5 | EIF5 Entrez, Source | eukaryotic translation initiation factor 5 | 4315 | 0.025 | 0.3507 | No |
| 66 | COL12A1 | COL12A1 Entrez, Source | collagen, type XII, alpha 1 | 4363 | 0.024 | 0.3489 | No |
| 67 | GFI1 | GFI1 Entrez, Source | growth factor independent 1 | 4553 | 0.022 | 0.3361 | No |
| 68 | GRIN2B | GRIN2B Entrez, Source | glutamate receptor, ionotropic, N-methyl D-aspartate 2B | 4866 | 0.017 | 0.3136 | No |
| 69 | NMT1 | NMT1 Entrez, Source | N-myristoyltransferase 1 | 4905 | 0.017 | 0.3119 | No |
| 70 | SCN3B | SCN3B Entrez, Source | sodium channel, voltage-gated, type III, beta | 5097 | 0.014 | 0.2984 | No |
| 71 | HIP1R | HIP1R Entrez, Source | huntingtin interacting protein 1 related | 5145 | 0.013 | 0.2958 | No |
| 72 | PHF1 | PHF1 Entrez, Source | PHD finger protein 1 | 5282 | 0.011 | 0.2863 | No |
| 73 | ID1 | ID1 Entrez, Source | inhibitor of DNA binding 1, dominant negative helix-loop-helix protein | 5296 | 0.011 | 0.2861 | No |
| 74 | ATPIF1 | ATPIF1 Entrez, Source | ATPase inhibitory factor 1 | 5346 | 0.010 | 0.2831 | No |
| 75 | TTC17 | TTC17 Entrez, Source | tetratricopeptide repeat domain 17 | 5348 | 0.010 | 0.2838 | No |
| 76 | ESRRG | ESRRG Entrez, Source | estrogen-related receptor gamma | 5378 | 0.010 | 0.2823 | No |
| 77 | EPHA7 | EPHA7 Entrez, Source | EPH receptor A7 | 5489 | 0.009 | 0.2745 | No |
| 78 | NFYB | NFYB Entrez, Source | nuclear transcription factor Y, beta | 5521 | 0.008 | 0.2727 | No |
| 79 | CACNB2 | CACNB2 Entrez, Source | calcium channel, voltage-dependent, beta 2 subunit | 5571 | 0.007 | 0.2695 | No |
| 80 | KCNIP2 | KCNIP2 Entrez, Source | Kv channel interacting protein 2 | 5580 | 0.007 | 0.2695 | No |
| 81 | MAP6 | MAP6 Entrez, Source | microtubule-associated protein 6 | 5603 | 0.007 | 0.2683 | No |
| 82 | ZFP91 | ZFP91 Entrez, Source | zinc finger protein 91 homolog (mouse) | 5687 | 0.006 | 0.2624 | No |
| 83 | PFN2 | PFN2 Entrez, Source | profilin 2 | 5738 | 0.005 | 0.2590 | No |
| 84 | KCNN3 | KCNN3 Entrez, Source | potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 | 6008 | 0.002 | 0.2387 | No |
| 85 | POFUT1 | POFUT1 Entrez, Source | protein O-fucosyltransferase 1 | 6279 | -0.002 | 0.2184 | No |
| 86 | DNAJA2 | DNAJA2 Entrez, Source | DnaJ (Hsp40) homolog, subfamily A, member 2 | 6450 | -0.004 | 0.2057 | No |
| 87 | OMG | OMG Entrez, Source | oligodendrocyte myelin glycoprotein | 6563 | -0.005 | 0.1976 | No |
| 88 | HOXA5 | HOXA5 Entrez, Source | homeobox A5 | 6663 | -0.007 | 0.1906 | No |
| 89 | WT1 | WT1 Entrez, Source | Wilms tumor 1 | 6748 | -0.008 | 0.1847 | No |
| 90 | CNTN1 | CNTN1 Entrez, Source | contactin 1 | 6807 | -0.008 | 0.1809 | No |
| 91 | HSD3B7 | HSD3B7 Entrez, Source | hydroxy-delta-5-steroid dehydrogenase, 3 beta- and steroid delta-isomerase 7 | 6860 | -0.009 | 0.1776 | No |
| 92 | GPR85 | GPR85 Entrez, Source | G protein-coupled receptor 85 | 6883 | -0.009 | 0.1766 | No |
| 93 | SMARCA2 | SMARCA2 Entrez, Source | SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 | 6971 | -0.010 | 0.1707 | No |
| 94 | GABRA3 | GABRA3 Entrez, Source | gamma-aminobutyric acid (GABA) A receptor, alpha 3 | 6994 | -0.011 | 0.1698 | No |
| 95 | CRYGS | CRYGS Entrez, Source | crystallin, gamma S | 7082 | -0.012 | 0.1640 | No |
| 96 | CLTC | CLTC Entrez, Source | clathrin, heavy chain (Hc) | 7295 | -0.014 | 0.1489 | No |
| 97 | NTRK2 | NTRK2 Entrez, Source | neurotrophic tyrosine kinase, receptor, type 2 | 7314 | -0.014 | 0.1486 | No |
| 98 | PPP3CA | PPP3CA Entrez, Source | protein phosphatase 3 (formerly 2B), catalytic subunit, alpha isoform (calcineurin A alpha) | 7318 | -0.014 | 0.1494 | No |
| 99 | PDGFRA | PDGFRA Entrez, Source | platelet-derived growth factor receptor, alpha polypeptide | 7343 | -0.015 | 0.1486 | No |
| 100 | NDUFS4 | NDUFS4 Entrez, Source | NADH dehydrogenase (ubiquinone) Fe-S protein 4, 18kDa (NADH-coenzyme Q reductase) | 7499 | -0.017 | 0.1380 | No |
| 101 | BUB3 | BUB3 Entrez, Source | BUB3 budding uninhibited by benzimidazoles 3 homolog (yeast) | 7602 | -0.018 | 0.1315 | No |
| 102 | STRN3 | STRN3 Entrez, Source | striatin, calmodulin binding protein 3 | 7629 | -0.019 | 0.1309 | No |
| 103 | LRRC4 | LRRC4 Entrez, Source | leucine rich repeat containing 4 | 7670 | -0.019 | 0.1292 | No |
| 104 | NXPH4 | NXPH4 Entrez, Source | neurexophilin 4 | 7712 | -0.020 | 0.1275 | No |
| 105 | ABT1 | ABT1 Entrez, Source | activator of basal transcription 1 | 7714 | -0.020 | 0.1287 | No |
| 106 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 7901 | -0.022 | 0.1162 | No |
| 107 | FOXA1 | FOXA1 Entrez, Source | forkhead box A1 | 8131 | -0.026 | 0.1006 | No |
| 108 | KITLG | KITLG Entrez, Source | KIT ligand | 8254 | -0.027 | 0.0933 | No |
| 109 | SLC4A1 | SLC4A1 Entrez, Source | solute carrier family 4, anion exchanger, member 1 (erythrocyte membrane protein band 3, Diego blood group) | 8397 | -0.029 | 0.0846 | No |
| 110 | VDR | VDR Entrez, Source | vitamin D (1,25- dihydroxyvitamin D3) receptor | 8460 | -0.030 | 0.0820 | No |
| 111 | ZNF403 | ZNF403 Entrez, Source | zinc finger protein 403 | 8532 | -0.031 | 0.0788 | No |
| 112 | BAMBI | BAMBI Entrez, Source | BMP and activin membrane-bound inhibitor homolog (Xenopus laevis) | 8540 | -0.031 | 0.0805 | No |
| 113 | NAV2 | NAV2 Entrez, Source | neuron navigator 2 | 8609 | -0.032 | 0.0776 | No |
| 114 | CASQ2 | CASQ2 Entrez, Source | calsequestrin 2 (cardiac muscle) | 8759 | -0.034 | 0.0687 | No |
| 115 | FGFR1OP2 | FGFR1OP2 Entrez, Source | FGFR1 oncogene partner 2 | 8927 | -0.036 | 0.0586 | No |
| 116 | CRKL | CRKL Entrez, Source | v-crk sarcoma virus CT10 oncogene homolog (avian)-like | 8950 | -0.037 | 0.0596 | No |
| 117 | CMKLR1 | CMKLR1 Entrez, Source | chemokine-like receptor 1 | 9069 | -0.038 | 0.0533 | No |
| 118 | HN1 | HN1 Entrez, Source | hematological and neurological expressed 1 | 9156 | -0.040 | 0.0496 | No |
| 119 | MID1 | MID1 Entrez, Source | midline 1 (Opitz/BBB syndrome) | 9243 | -0.041 | 0.0459 | No |
| 120 | DMRT1 | DMRT1 Entrez, Source | doublesex and mab-3 related transcription factor 1 | 9294 | -0.042 | 0.0451 | No |
| 121 | ATP1A2 | ATP1A2 Entrez, Source | ATPase, Na+/K+ transporting, alpha 2 (+) polypeptide | 9475 | -0.045 | 0.0346 | No |
| 122 | TRIM28 | TRIM28 Entrez, Source | tripartite motif-containing 28 | 9549 | -0.046 | 0.0323 | No |
| 123 | PDLIM2 | PDLIM2 Entrez, Source | PDZ and LIM domain 2 (mystique) | 9564 | -0.046 | 0.0344 | No |
| 124 | CDH2 | CDH2 Entrez, Source | cadherin 2, type 1, N-cadherin (neuronal) | 9642 | -0.048 | 0.0320 | No |
| 125 | PHC2 | PHC2 Entrez, Source | polyhomeotic homolog 2 (Drosophila) | 9654 | -0.048 | 0.0345 | No |
| 126 | ACVR1 | ACVR1 Entrez, Source | activin A receptor, type I | 9665 | -0.048 | 0.0371 | No |
| 127 | CACNA2D3 | CACNA2D3 Entrez, Source | calcium channel, voltage-dependent, alpha 2/delta 3 subunit | 9729 | -0.049 | 0.0358 | No |
| 128 | MRPL13 | MRPL13 Entrez, Source | mitochondrial ribosomal protein L13 | 9917 | -0.052 | 0.0253 | No |
| 129 | IGF1 | IGF1 Entrez, Source | insulin-like growth factor 1 (somatomedin C) | 10373 | -0.060 | -0.0050 | No |
| 130 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 10486 | -0.062 | -0.0092 | No |
| 131 | MTA2 | MTA2 Entrez, Source | metastasis associated 1 family, member 2 | 10748 | -0.067 | -0.0242 | No |
| 132 | FGFR1 | FGFR1 Entrez, Source | fibroblast growth factor receptor 1 (fms-related tyrosine kinase 2, Pfeiffer syndrome) | 10807 | -0.069 | -0.0238 | No |
| 133 | MGAT4A | MGAT4A Entrez, Source | mannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme A | 10840 | -0.069 | -0.0213 | No |
| 134 | PCYT1B | PCYT1B Entrez, Source | phosphate cytidylyltransferase 1, choline, beta | 10932 | -0.072 | -0.0232 | No |
| 135 | PMCH | PMCH Entrez, Source | pro-melanin-concentrating hormone | 11172 | -0.078 | -0.0359 | No |
| 136 | XRCC1 | XRCC1 Entrez, Source | X-ray repair complementing defective repair in Chinese hamster cells 1 | 11227 | -0.079 | -0.0344 | No |
| 137 | PLEKHM1 | PLEKHM1 Entrez, Source | pleckstrin homology domain containing, family M (with RUN domain) member 1 | 11344 | -0.083 | -0.0374 | No |
| 138 | FGD4 | FGD4 Entrez, Source | FYVE, RhoGEF and PH domain containing 4 | 11381 | -0.084 | -0.0342 | No |
| 139 | CSNK1A1 | CSNK1A1 Entrez, Source | casein kinase 1, alpha 1 | 11589 | -0.089 | -0.0437 | No |
| 140 | PARK2 | PARK2 Entrez, Source | Parkinson disease (autosomal recessive, juvenile) 2, parkin | 11750 | -0.094 | -0.0492 | No |
| 141 | NMI | NMI Entrez, Source | N-myc (and STAT) interactor | 11925 | -0.100 | -0.0554 | No |
| 142 | CDC23 | CDC23 Entrez, Source | CDC23 (cell division cycle 23, yeast, homolog) | 11974 | -0.102 | -0.0519 | No |
| 143 | SLC6A1 | SLC6A1 Entrez, Source | solute carrier family 6 (neurotransmitter transporter, GABA), member 1 | 11982 | -0.102 | -0.0452 | No |
| 144 | EFNA1 | EFNA1 Entrez, Source | ephrin-A1 | 11985 | -0.102 | -0.0382 | No |
| 145 | EDG5 | EDG5 Entrez, Source | endothelial differentiation, sphingolipid G-protein-coupled receptor, 5 | 12054 | -0.105 | -0.0359 | No |
| 146 | CHRM1 | CHRM1 Entrez, Source | cholinergic receptor, muscarinic 1 | 12093 | -0.107 | -0.0313 | No |
| 147 | SIPA1 | SIPA1 Entrez, Source | signal-induced proliferation-associated gene 1 | 12263 | -0.115 | -0.0360 | No |
| 148 | PLEC1 | PLEC1 Entrez, Source | plectin 1, intermediate filament binding protein 500kDa | 12297 | -0.116 | -0.0304 | No |
| 149 | DHRS4 | DHRS4 Entrez, Source | dehydrogenase/reductase (SDR family) member 4 | 12452 | -0.124 | -0.0334 | No |
| 150 | USP1 | USP1 Entrez, Source | ubiquitin specific peptidase 1 | 12463 | -0.125 | -0.0253 | No |
| 151 | NUP54 | NUP54 Entrez, Source | nucleoporin 54kDa | 12635 | -0.137 | -0.0287 | No |
| 152 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 12659 | -0.138 | -0.0207 | No |
| 153 | DYRK1A | DYRK1A Entrez, Source | dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A | 12862 | -0.160 | -0.0248 | No |
| 154 | VKORC1L1 | VKORC1L1 Entrez, Source | vitamin K epoxide reductase complex, subunit 1-like 1 | 12869 | -0.161 | -0.0139 | No |
| 155 | INHBC | INHBC Entrez, Source | inhibin, beta C | 12890 | -0.163 | -0.0039 | No |
| 156 | DMD | DMD Entrez, Source | dystrophin (muscular dystrophy, Duchenne and Becker types) | 12922 | -0.168 | 0.0055 | No |
| 157 | CA3 | CA3 Entrez, Source | carbonic anhydrase III, muscle specific | 13289 | -0.373 | 0.0040 | No |