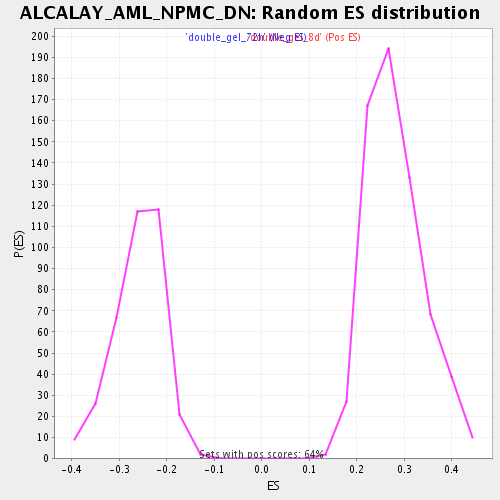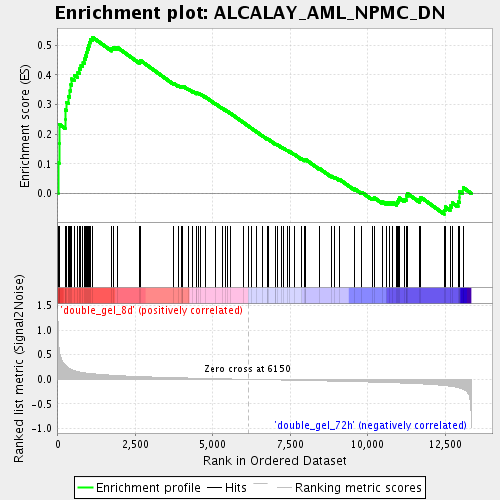
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | ALCALAY_AML_NPMC_DN |
| Enrichment Score (ES) | 0.52719146 |
| Normalized Enrichment Score (NES) | 1.8772627 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.019528592 |
| FWER p-Value | 0.747 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | IGFBP7 | IGFBP7 Entrez, Source | insulin-like growth factor binding protein 7 | 5 | 0.972 | 0.1046 | Yes |
| 2 | HLA-DMA | HLA-DMA Entrez, Source | major histocompatibility complex, class II, DM alpha | 40 | 0.620 | 0.1690 | Yes |
| 3 | IL1R2 | IL1R2 Entrez, Source | interleukin 1 receptor, type II | 46 | 0.599 | 0.2333 | Yes |
| 4 | SPARC | SPARC Entrez, Source | secreted protein, acidic, cysteine-rich (osteonectin) | 234 | 0.297 | 0.2512 | Yes |
| 5 | CD48 | CD48 Entrez, Source | CD48 molecule | 239 | 0.291 | 0.2823 | Yes |
| 6 | IFITM2 | IFITM2 Entrez, Source | interferon induced transmembrane protein 2 (1-8D) | 281 | 0.260 | 0.3073 | Yes |
| 7 | CD96 | CD96 Entrez, Source | CD96 molecule | 341 | 0.233 | 0.3281 | Yes |
| 8 | HNRPD | HNRPD Entrez, Source | heterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) | 389 | 0.213 | 0.3475 | Yes |
| 9 | GSN | GSN Entrez, Source | gelsolin (amyloidosis, Finnish type) | 417 | 0.205 | 0.3676 | Yes |
| 10 | PROM1 | PROM1 Entrez, Source | prominin 1 | 448 | 0.196 | 0.3865 | Yes |
| 11 | ACAA2 | ACAA2 Entrez, Source | acetyl-Coenzyme A acyltransferase 2 (mitochondrial 3-oxoacyl-Coenzyme A thiolase) | 528 | 0.177 | 0.3997 | Yes |
| 12 | ITGA9 | ITGA9 Entrez, Source | integrin, alpha 9 | 643 | 0.158 | 0.4082 | Yes |
| 13 | ITM2C | ITM2C Entrez, Source | integral membrane protein 2C | 696 | 0.150 | 0.4205 | Yes |
| 14 | PLAGL1 | PLAGL1 Entrez, Source | pleiomorphic adenoma gene-like 1 | 742 | 0.145 | 0.4328 | Yes |
| 15 | MLLT3 | MLLT3 Entrez, Source | myeloid/lymphoid or mixed-lineage leukemia (trithorax homolog, Drosophila); translocated to, 3 | 808 | 0.139 | 0.4429 | Yes |
| 16 | TNFRSF14 | TNFRSF14 Entrez, Source | tumor necrosis factor receptor superfamily, member 14 (herpesvirus entry mediator) | 852 | 0.134 | 0.4541 | Yes |
| 17 | CD69 | CD69 Entrez, Source | CD69 molecule | 898 | 0.129 | 0.4647 | Yes |
| 18 | IL2RG | IL2RG Entrez, Source | interleukin 2 receptor, gamma (severe combined immunodeficiency) | 927 | 0.127 | 0.4763 | Yes |
| 19 | PBX2 | PBX2 Entrez, Source | pre-B-cell leukemia transcription factor 2 | 944 | 0.126 | 0.4886 | Yes |
| 20 | ACAT1 | ACAT1 Entrez, Source | acetyl-Coenzyme A acetyltransferase 1 (acetoacetyl Coenzyme A thiolase) | 990 | 0.123 | 0.4985 | Yes |
| 21 | RGS10 | RGS10 Entrez, Source | regulator of G-protein signalling 10 | 1009 | 0.121 | 0.5102 | Yes |
| 22 | TRAK2 | TRAK2 Entrez, Source | trafficking protein, kinesin binding 2 | 1042 | 0.118 | 0.5205 | Yes |
| 23 | B4GALT6 | B4GALT6 Entrez, Source | UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 6 | 1117 | 0.114 | 0.5272 | Yes |
| 24 | PTTG1 | PTTG1 Entrez, Source | pituitary tumor-transforming 1 | 1734 | 0.083 | 0.4897 | No |
| 25 | HADH | HADH Entrez, Source | hydroxyacyl-Coenzyme A dehydrogenase | 1791 | 0.081 | 0.4942 | No |
| 26 | WASF1 | WASF1 Entrez, Source | WAS protein family, member 1 | 1914 | 0.076 | 0.4932 | No |
| 27 | PIP5K1B | PIP5K1B Entrez, Source | phosphatidylinositol-4-phosphate 5-kinase, type I, beta | 2643 | 0.056 | 0.4443 | No |
| 28 | ANK1 | ANK1 Entrez, Source | ankyrin 1, erythrocytic | 2657 | 0.056 | 0.4494 | No |
| 29 | FKBP1B | FKBP1B Entrez, Source | FK506 binding protein 1B, 12.6 kDa | 3733 | 0.035 | 0.3720 | No |
| 30 | BIRC5 | BIRC5 Entrez, Source | baculoviral IAP repeat-containing 5 (survivin) | 3898 | 0.032 | 0.3631 | No |
| 31 | PXMP2 | PXMP2 Entrez, Source | peroxisomal membrane protein 2, 22kDa | 3983 | 0.031 | 0.3600 | No |
| 32 | GNB5 | GNB5 Entrez, Source | guanine nucleotide binding protein (G protein), beta 5 | 4018 | 0.030 | 0.3607 | No |
| 33 | TRH | TRH Entrez, Source | thyrotropin-releasing hormone | 4036 | 0.030 | 0.3626 | No |
| 34 | TUBG1 | TUBG1 Entrez, Source | tubulin, gamma 1 | 4212 | 0.027 | 0.3523 | No |
| 35 | HPCAL1 | HPCAL1 Entrez, Source | hippocalcin-like 1 | 4353 | 0.025 | 0.3444 | No |
| 36 | CCNF | CCNF Entrez, Source | cyclin F | 4470 | 0.023 | 0.3381 | No |
| 37 | CCNA2 | CCNA2 Entrez, Source | cyclin A2 | 4471 | 0.023 | 0.3405 | No |
| 38 | GNG7 | GNG7 Entrez, Source | guanine nucleotide binding protein (G protein), gamma 7 | 4525 | 0.022 | 0.3389 | No |
| 39 | RPS6KA2 | RPS6KA2 Entrez, Source | ribosomal protein S6 kinase, 90kDa, polypeptide 2 | 4610 | 0.021 | 0.3348 | No |
| 40 | ASNA1 | ASNA1 Entrez, Source | arsA arsenite transporter, ATP-binding, homolog 1 (bacterial) | 4755 | 0.019 | 0.3260 | No |
| 41 | HMGN3 | HMGN3 Entrez, Source | high mobility group nucleosomal binding domain 3 | 5092 | 0.014 | 0.3022 | No |
| 42 | INSL3 | INSL3 Entrez, Source | insulin-like 3 (Leydig cell) | 5304 | 0.011 | 0.2874 | No |
| 43 | SERPINI1 | SERPINI1 Entrez, Source | serpin peptidase inhibitor, clade I (neuroserpin), member 1 | 5393 | 0.010 | 0.2819 | No |
| 44 | INPP4B | INPP4B Entrez, Source | inositol polyphosphate-4-phosphatase, type II, 105kDa | 5469 | 0.009 | 0.2771 | No |
| 45 | KCNIP2 | KCNIP2 Entrez, Source | Kv channel interacting protein 2 | 5580 | 0.007 | 0.2696 | No |
| 46 | STOM | STOM Entrez, Source | stomatin | 5982 | 0.002 | 0.2396 | No |
| 47 | KIF2C | KIF2C Entrez, Source | kinesin family member 2C | 6164 | -0.000 | 0.2260 | No |
| 48 | CDKN2C | CDKN2C Entrez, Source | cyclin-dependent kinase inhibitor 2C (p18, inhibits CDK4) | 6239 | -0.001 | 0.2205 | No |
| 49 | EVL | EVL Entrez, Source | Enah/Vasp-like | 6399 | -0.003 | 0.2088 | No |
| 50 | BAALC | BAALC Entrez, Source | brain and acute leukemia, cytoplasmic | 6612 | -0.006 | 0.1935 | No |
| 51 | FOXM1 | FOXM1 Entrez, Source | forkhead box M1 | 6751 | -0.008 | 0.1839 | No |
| 52 | EGFL7 | EGFL7 Entrez, Source | EGF-like-domain, multiple 7 | 6762 | -0.008 | 0.1840 | No |
| 53 | SEPT9 | SEPT9 Entrez, Source | septin 9 | 6782 | -0.008 | 0.1834 | No |
| 54 | OGG1 | OGG1 Entrez, Source | 8-oxoguanine DNA glycosylase | 7030 | -0.011 | 0.1659 | No |
| 55 | NINJ2 | NINJ2 Entrez, Source | ninjurin 2 | 7033 | -0.011 | 0.1670 | No |
| 56 | CD38 | CD38 Entrez, Source | CD38 molecule | 7086 | -0.012 | 0.1643 | No |
| 57 | CRHBP | CRHBP Entrez, Source | corticotropin releasing hormone binding protein | 7209 | -0.013 | 0.1565 | No |
| 58 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 7289 | -0.014 | 0.1521 | No |
| 59 | CORO2A | CORO2A Entrez, Source | coronin, actin binding protein, 2A | 7394 | -0.015 | 0.1459 | No |
| 60 | HMMR | HMMR Entrez, Source | hyaluronan-mediated motility receptor (RHAMM) | 7489 | -0.017 | 0.1406 | No |
| 61 | FAAH | FAAH Entrez, Source | fatty acid amide hydrolase | 7626 | -0.018 | 0.1323 | No |
| 62 | CIITA | CIITA Entrez, Source | class II, major histocompatibility complex, transactivator | 7855 | -0.021 | 0.1174 | No |
| 63 | RAD23B | RAD23B Entrez, Source | RAD23 homolog B (S. cerevisiae) | 7961 | -0.023 | 0.1120 | No |
| 64 | CHEK1 | CHEK1 Entrez, Source | CHK1 checkpoint homolog (S. pombe) | 7980 | -0.023 | 0.1131 | No |
| 65 | CCNB1 | CCNB1 Entrez, Source | cyclin B1 | 8004 | -0.024 | 0.1139 | No |
| 66 | S100B | S100B Entrez, Source | S100 calcium binding protein B | 8430 | -0.030 | 0.0851 | No |
| 67 | KIF4A | KIF4A Entrez, Source | kinesin family member 4A | 8827 | -0.035 | 0.0590 | No |
| 68 | SERPINF1 | SERPINF1 Entrez, Source | serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 1 | 8926 | -0.036 | 0.0555 | No |
| 69 | SLC1A4 | SLC1A4 Entrez, Source | solute carrier family 1 (glutamate/neutral amino acid transporter), member 4 | 9073 | -0.038 | 0.0486 | No |
| 70 | TMEM106C | TMEM106C Entrez, Source | transmembrane protein 106C | 9570 | -0.046 | 0.0162 | No |
| 71 | CD200 | CD200 Entrez, Source | CD200 molecule | 9804 | -0.050 | 0.0041 | No |
| 72 | KIFC1 | KIFC1 Entrez, Source | kinesin family member C1 | 10151 | -0.056 | -0.0160 | No |
| 73 | NUP210 | NUP210 Entrez, Source | nucleoporin 210kDa | 10202 | -0.057 | -0.0137 | No |
| 74 | HGF | HGF Entrez, Source | hepatocyte growth factor (hepapoietin A; scatter factor) | 10486 | -0.062 | -0.0283 | No |
| 75 | CCNB2 | CCNB2 Entrez, Source | cyclin B2 | 10615 | -0.065 | -0.0310 | No |
| 76 | VEGF | VEGF Entrez, Source | vascular endothelial growth factor | 10686 | -0.066 | -0.0292 | No |
| 77 | CDKN3 | CDKN3 Entrez, Source | cyclin-dependent kinase inhibitor 3 (CDK2-associated dual specificity phosphatase) | 10809 | -0.069 | -0.0310 | No |
| 78 | POLD1 | POLD1 Entrez, Source | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | 10927 | -0.072 | -0.0321 | No |
| 79 | MECR | MECR Entrez, Source | mitochondrial trans-2-enoyl-CoA reductase | 10955 | -0.072 | -0.0263 | No |
| 80 | TYMS | TYMS Entrez, Source | thymidylate synthetase | 10993 | -0.073 | -0.0212 | No |
| 81 | RHD | RHD Entrez, Source | Rh blood group, D antigen | 11014 | -0.074 | -0.0148 | No |
| 82 | CDC20 | CDC20 Entrez, Source | CDC20 cell division cycle 20 homolog (S. cerevisiae) | 11187 | -0.078 | -0.0194 | No |
| 83 | ARL2 | ARL2 Entrez, Source | ADP-ribosylation factor-like 2 | 11242 | -0.080 | -0.0148 | No |
| 84 | SYNGR1 | SYNGR1 Entrez, Source | synaptogyrin 1 | 11243 | -0.080 | -0.0062 | No |
| 85 | MTHFD1 | MTHFD1 Entrez, Source | methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1, methenyltetrahydrofolate cyclohydrolase, formyltetrahydrofolate synthetase | 11282 | -0.081 | -0.0004 | No |
| 86 | NPR3 | NPR3 Entrez, Source | natriuretic peptide receptor C/guanylate cyclase C (atrionatriuretic peptide receptor C) | 11674 | -0.091 | -0.0201 | No |
| 87 | DUT | DUT Entrez, Source | dUTP pyrophosphatase | 11707 | -0.092 | -0.0125 | No |
| 88 | HAAO | HAAO Entrez, Source | 3-hydroxyanthranilate 3,4-dioxygenase | 12465 | -0.125 | -0.0562 | No |
| 89 | RBM3 | RBM3 Entrez, Source | RNA binding motif (RNP1, RRM) protein 3 | 12503 | -0.128 | -0.0451 | No |
| 90 | POU4F1 | POU4F1 Entrez, Source | POU domain, class 4, transcription factor 1 | 12659 | -0.138 | -0.0419 | No |
| 91 | ABCG2 | ABCG2 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 2 | 12719 | -0.143 | -0.0309 | No |
| 92 | SRD5A1 | SRD5A1 Entrez, Source | steroid-5-alpha-reductase, alpha polypeptide 1 (3-oxo-5 alpha-steroid delta 4-dehydrogenase alpha 1) | 12915 | -0.167 | -0.0276 | No |
| 93 | GAMT | GAMT Entrez, Source | guanidinoacetate N-methyltransferase | 12960 | -0.175 | -0.0121 | No |
| 94 | DHRS3 | DHRS3 Entrez, Source | dehydrogenase/reductase (SDR family) member 3 | 12962 | -0.175 | 0.0067 | No |
| 95 | HOMER2 | HOMER2 Entrez, Source | homer homolog 2 (Drosophila) | 13074 | -0.203 | 0.0202 | No |