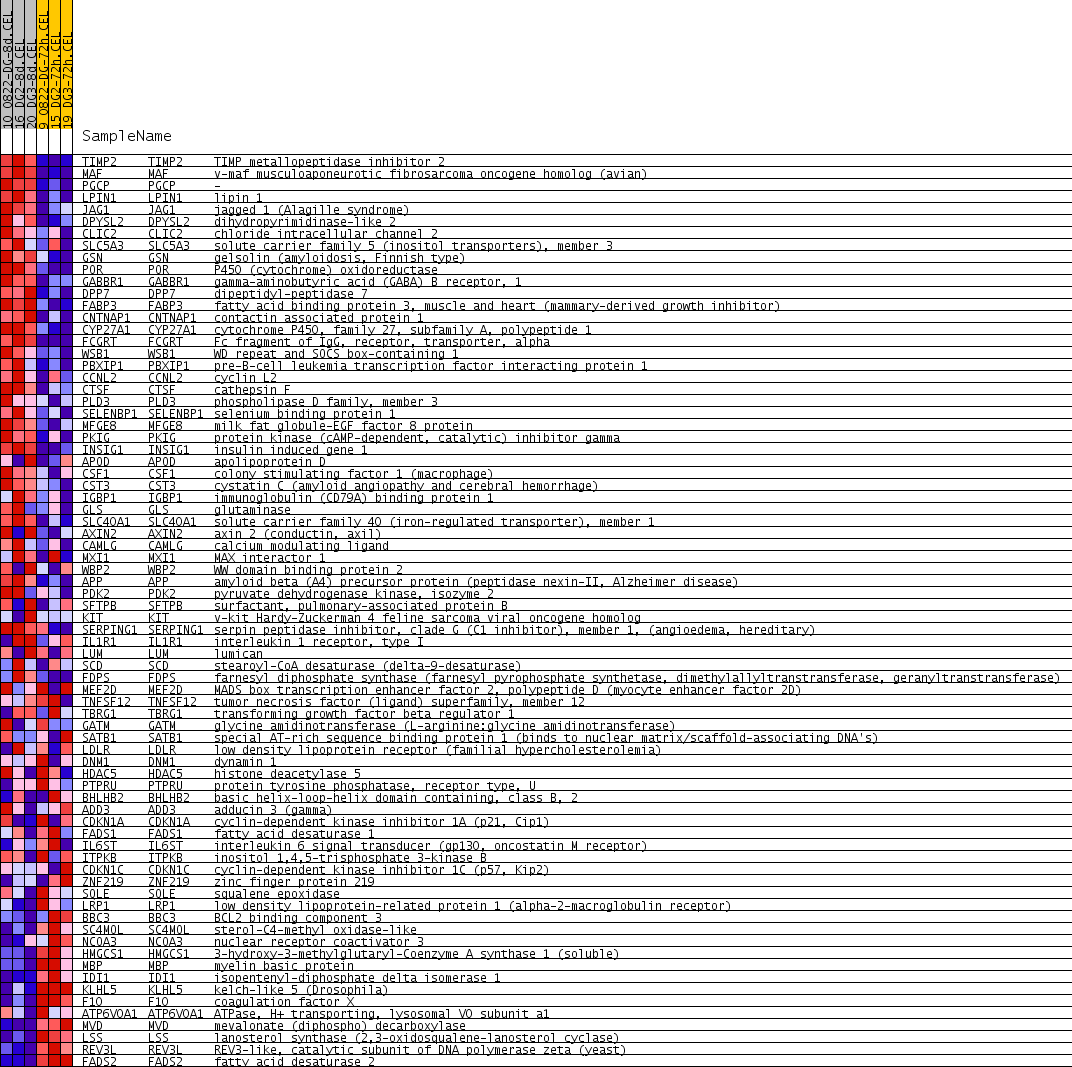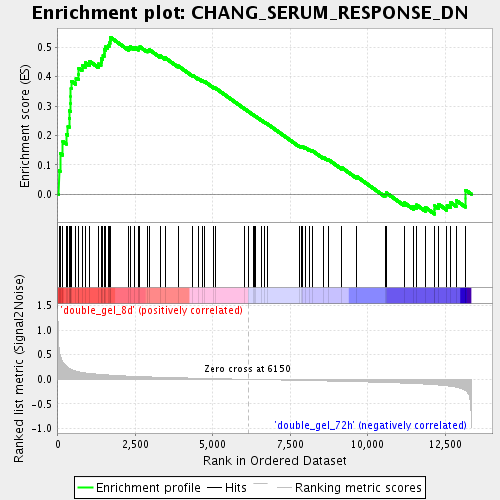
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | CHANG_SERUM_RESPONSE_DN |
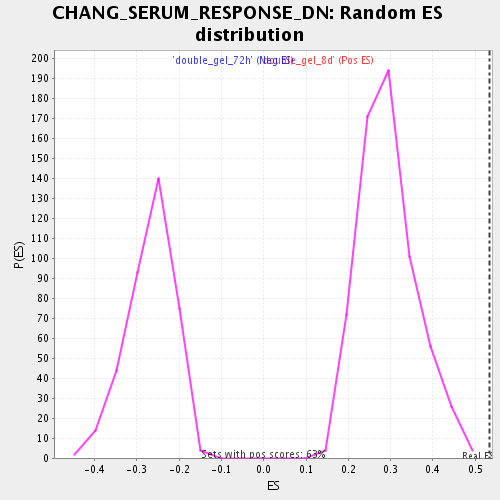
| Enrichment Score (ES) | 0.5340361 |
| Normalized Enrichment Score (NES) | 1.8183098 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.028904414 |
| FWER p-Value | 0.945 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TIMP2 | TIMP2 Entrez, Source | TIMP metallopeptidase inhibitor 2 | 35 | 0.642 | 0.0798 | Yes |
| 2 | MAF | MAF Entrez, Source | v-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) | 73 | 0.484 | 0.1393 | Yes |
| 3 | PGCP | PGCP Entrez, Source | - | 159 | 0.356 | 0.1786 | Yes |
| 4 | LPIN1 | LPIN1 Entrez, Source | lipin 1 | 269 | 0.266 | 0.2046 | Yes |
| 5 | JAG1 | JAG1 Entrez, Source | jagged 1 (Alagille syndrome) | 323 | 0.239 | 0.2313 | Yes |
| 6 | DPYSL2 | DPYSL2 Entrez, Source | dihydropyrimidinase-like 2 | 366 | 0.223 | 0.2568 | Yes |
| 7 | CLIC2 | CLIC2 Entrez, Source | chloride intracellular channel 2 | 378 | 0.216 | 0.2837 | Yes |
| 8 | SLC5A3 | SLC5A3 Entrez, Source | solute carrier family 5 (inositol transporters), member 3 | 411 | 0.207 | 0.3078 | Yes |
| 9 | GSN | GSN Entrez, Source | gelsolin (amyloidosis, Finnish type) | 417 | 0.205 | 0.3338 | Yes |
| 10 | POR | POR Entrez, Source | P450 (cytochrome) oxidoreductase | 421 | 0.204 | 0.3598 | Yes |
| 11 | GABBR1 | GABBR1 Entrez, Source | gamma-aminobutyric acid (GABA) B receptor, 1 | 432 | 0.200 | 0.3848 | Yes |
| 12 | DPP7 | DPP7 Entrez, Source | dipeptidyl-peptidase 7 | 582 | 0.167 | 0.3950 | Yes |
| 13 | FABP3 | FABP3 Entrez, Source | fatty acid binding protein 3, muscle and heart (mammary-derived growth inhibitor) | 666 | 0.155 | 0.4087 | Yes |
| 14 | CNTNAP1 | CNTNAP1 Entrez, Source | contactin associated protein 1 | 673 | 0.154 | 0.4281 | Yes |
| 15 | CYP27A1 | CYP27A1 Entrez, Source | cytochrome P450, family 27, subfamily A, polypeptide 1 | 781 | 0.141 | 0.4382 | Yes |
| 16 | FCGRT | FCGRT Entrez, Source | Fc fragment of IgG, receptor, transporter, alpha | 882 | 0.131 | 0.4475 | Yes |
| 17 | WSB1 | WSB1 Entrez, Source | WD repeat and SOCS box-containing 1 | 1023 | 0.119 | 0.4522 | Yes |
| 18 | PBXIP1 | PBXIP1 Entrez, Source | pre-B-cell leukemia transcription factor interacting protein 1 | 1318 | 0.103 | 0.4433 | Yes |
| 19 | CCNL2 | CCNL2 Entrez, Source | cyclin L2 | 1392 | 0.099 | 0.4505 | Yes |
| 20 | CTSF | CTSF Entrez, Source | cathepsin F | 1408 | 0.098 | 0.4620 | Yes |
| 21 | PLD3 | PLD3 Entrez, Source | phospholipase D family, member 3 | 1454 | 0.096 | 0.4709 | Yes |
| 22 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 1492 | 0.094 | 0.4803 | Yes |
| 23 | MFGE8 | MFGE8 Entrez, Source | milk fat globule-EGF factor 8 protein | 1496 | 0.094 | 0.4921 | Yes |
| 24 | PKIG | PKIG Entrez, Source | protein kinase (cAMP-dependent, catalytic) inhibitor gamma | 1533 | 0.092 | 0.5013 | Yes |
| 25 | INSIG1 | INSIG1 Entrez, Source | insulin induced gene 1 | 1616 | 0.089 | 0.5065 | Yes |
| 26 | APOD | APOD Entrez, Source | apolipoprotein D | 1653 | 0.087 | 0.5149 | Yes |
| 27 | CSF1 | CSF1 Entrez, Source | colony stimulating factor 1 (macrophage) | 1683 | 0.086 | 0.5237 | Yes |
| 28 | CST3 | CST3 Entrez, Source | cystatin C (amyloid angiopathy and cerebral hemorrhage) | 1692 | 0.085 | 0.5340 | Yes |
| 29 | IGBP1 | IGBP1 Entrez, Source | immunoglobulin (CD79A) binding protein 1 | 2280 | 0.065 | 0.4981 | No |
| 30 | GLS | GLS Entrez, Source | glutaminase | 2338 | 0.064 | 0.5020 | No |
| 31 | SLC40A1 | SLC40A1 Entrez, Source | solute carrier family 40 (iron-regulated transporter), member 1 | 2463 | 0.061 | 0.5005 | No |
| 32 | AXIN2 | AXIN2 Entrez, Source | axin 2 (conductin, axil) | 2598 | 0.058 | 0.4977 | No |
| 33 | CAMLG | CAMLG Entrez, Source | calcium modulating ligand | 2647 | 0.056 | 0.5014 | No |
| 34 | MXI1 | MXI1 Entrez, Source | MAX interactor 1 | 2881 | 0.051 | 0.4904 | No |
| 35 | WBP2 | WBP2 Entrez, Source | WW domain binding protein 2 | 2942 | 0.050 | 0.4923 | No |
| 36 | APP | APP Entrez, Source | amyloid beta (A4) precursor protein (peptidase nexin-II, Alzheimer disease) | 3298 | 0.043 | 0.4710 | No |
| 37 | PDK2 | PDK2 Entrez, Source | pyruvate dehydrogenase kinase, isozyme 2 | 3456 | 0.040 | 0.4642 | No |
| 38 | SFTPB | SFTPB Entrez, Source | surfactant, pulmonary-associated protein B | 3879 | 0.032 | 0.4366 | No |
| 39 | KIT | KIT Entrez, Source | v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog | 4345 | 0.025 | 0.4047 | No |
| 40 | SERPING1 | SERPING1 Entrez, Source | serpin peptidase inhibitor, clade G (C1 inhibitor), member 1, (angioedema, hereditary) | 4547 | 0.022 | 0.3924 | No |
| 41 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 4666 | 0.020 | 0.3860 | No |
| 42 | LUM | LUM Entrez, Source | lumican | 4728 | 0.019 | 0.3839 | No |
| 43 | SCD | SCD Entrez, Source | stearoyl-CoA desaturase (delta-9-desaturase) | 5035 | 0.015 | 0.3628 | No |
| 44 | FDPS | FDPS Entrez, Source | farnesyl diphosphate synthase (farnesyl pyrophosphate synthetase, dimethylallyltranstransferase, geranyltranstransferase) | 5077 | 0.014 | 0.3615 | No |
| 45 | MEF2D | MEF2D Entrez, Source | MADS box transcription enhancer factor 2, polypeptide D (myocyte enhancer factor 2D) | 6025 | 0.002 | 0.2904 | No |
| 46 | TNFSF12 | TNFSF12 Entrez, Source | tumor necrosis factor (ligand) superfamily, member 12 | 6140 | 0.000 | 0.2818 | No |
| 47 | TBRG1 | TBRG1 Entrez, Source | transforming growth factor beta regulator 1 | 6314 | -0.002 | 0.2690 | No |
| 48 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 6337 | -0.002 | 0.2676 | No |
| 49 | SATB1 | SATB1 Entrez, Source | special AT-rich sequence binding protein 1 (binds to nuclear matrix/scaffold-associating DNA's) | 6392 | -0.003 | 0.2640 | No |
| 50 | LDLR | LDLR Entrez, Source | low density lipoprotein receptor (familial hypercholesterolemia) | 6572 | -0.005 | 0.2512 | No |
| 51 | DNM1 | DNM1 Entrez, Source | dynamin 1 | 6675 | -0.007 | 0.2443 | No |
| 52 | HDAC5 | HDAC5 Entrez, Source | histone deacetylase 5 | 6756 | -0.008 | 0.2393 | No |
| 53 | PTPRU | PTPRU Entrez, Source | protein tyrosine phosphatase, receptor type, U | 7795 | -0.021 | 0.1637 | No |
| 54 | BHLHB2 | BHLHB2 Entrez, Source | basic helix-loop-helix domain containing, class B, 2 | 7852 | -0.021 | 0.1622 | No |
| 55 | ADD3 | ADD3 Entrez, Source | adducin 3 (gamma) | 7901 | -0.022 | 0.1614 | No |
| 56 | CDKN1A | CDKN1A Entrez, Source | cyclin-dependent kinase inhibitor 1A (p21, Cip1) | 7976 | -0.023 | 0.1588 | No |
| 57 | FADS1 | FADS1 Entrez, Source | fatty acid desaturase 1 | 8129 | -0.025 | 0.1506 | No |
| 58 | IL6ST | IL6ST Entrez, Source | interleukin 6 signal transducer (gp130, oncostatin M receptor) | 8212 | -0.027 | 0.1479 | No |
| 59 | ITPKB | ITPKB Entrez, Source | inositol 1,4,5-trisphosphate 3-kinase B | 8579 | -0.032 | 0.1244 | No |
| 60 | CDKN1C | CDKN1C Entrez, Source | cyclin-dependent kinase inhibitor 1C (p57, Kip2) | 8731 | -0.034 | 0.1174 | No |
| 61 | ZNF219 | ZNF219 Entrez, Source | zinc finger protein 219 | 9165 | -0.040 | 0.0899 | No |
| 62 | SQLE | SQLE Entrez, Source | squalene epoxidase | 9644 | -0.048 | 0.0600 | No |
| 63 | LRP1 | LRP1 Entrez, Source | low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor) | 10559 | -0.063 | -0.0008 | No |
| 64 | BBC3 | BBC3 Entrez, Source | BCL2 binding component 3 | 10590 | -0.064 | 0.0052 | No |
| 65 | SC4MOL | SC4MOL Entrez, Source | sterol-C4-methyl oxidase-like | 11182 | -0.078 | -0.0294 | No |
| 66 | NCOA3 | NCOA3 Entrez, Source | nuclear receptor coactivator 3 | 11477 | -0.086 | -0.0405 | No |
| 67 | HMGCS1 | HMGCS1 Entrez, Source | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 (soluble) | 11576 | -0.088 | -0.0365 | No |
| 68 | MBP | MBP Entrez, Source | myelin basic protein | 11862 | -0.098 | -0.0454 | No |
| 69 | IDI1 | IDI1 Entrez, Source | isopentenyl-diphosphate delta isomerase 1 | 12159 | -0.110 | -0.0536 | No |
| 70 | KLHL5 | KLHL5 Entrez, Source | kelch-like 5 (Drosophila) | 12161 | -0.110 | -0.0396 | No |
| 71 | F10 | F10 Entrez, Source | coagulation factor X | 12278 | -0.115 | -0.0336 | No |
| 72 | ATP6V0A1 | ATP6V0A1 Entrez, Source | ATPase, H+ transporting, lysosomal V0 subunit a1 | 12556 | -0.132 | -0.0375 | No |
| 73 | MVD | MVD Entrez, Source | mevalonate (diphospho) decarboxylase | 12678 | -0.140 | -0.0286 | No |
| 74 | LSS | LSS Entrez, Source | lanosterol synthase (2,3-oxidosqualene-lanosterol cyclase) | 12854 | -0.159 | -0.0214 | No |
| 75 | REV3L | REV3L Entrez, Source | REV3-like, catalytic subunit of DNA polymerase zeta (yeast) | 13154 | -0.226 | -0.0150 | No |
| 76 | FADS2 | FADS2 Entrez, Source | fatty acid desaturase 2 | 13155 | -0.226 | 0.0141 | No |