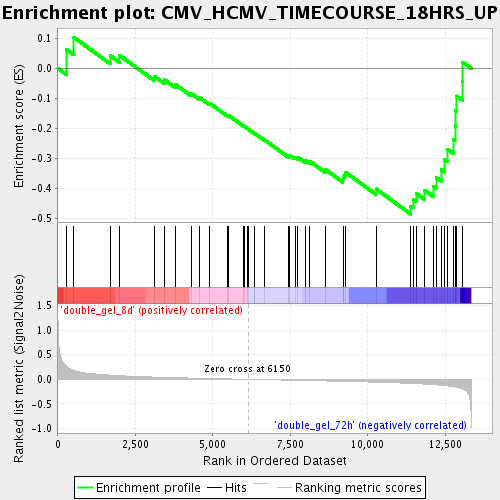
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_72h |
| GeneSet | CMV_HCMV_TIMECOURSE_18HRS_UP |
| Enrichment Score (ES) | -0.48562154 |
| Normalized Enrichment Score (NES) | -1.6290042 |
| Nominal p-value | 0.012406948 |
| FDR q-value | 0.18822421 |
| FWER p-Value | 1.0 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | TSC22D3 | TSC22D3 Entrez, Source | TSC22 domain family, member 3 | 276 | 0.263 | 0.0634 | No |
| 2 | NEU1 | NEU1 Entrez, Source | sialidase 1 (lysosomal sialidase) | 510 | 0.181 | 0.1037 | No |
| 3 | PTGER2 | PTGER2 Entrez, Source | prostaglandin E receptor 2 (subtype EP2), 53kDa | 1691 | 0.085 | 0.0422 | No |
| 4 | KCNS3 | KCNS3 Entrez, Source | potassium voltage-gated channel, delayed-rectifier, subfamily S, member 3 | 1982 | 0.074 | 0.0440 | No |
| 5 | DYNC1I1 | DYNC1I1 Entrez, Source | dynein, cytoplasmic 1, intermediate chain 1 | 3108 | 0.046 | -0.0258 | No |
| 6 | WDR47 | WDR47 Entrez, Source | WD repeat domain 47 | 3443 | 0.040 | -0.0382 | No |
| 7 | DMN | DMN Entrez, Source | desmuslin | 3794 | 0.034 | -0.0538 | No |
| 8 | BRAF | BRAF Entrez, Source | v-raf murine sarcoma viral oncogene homolog B1 | 4295 | 0.026 | -0.0832 | No |
| 9 | RFK | RFK Entrez, Source | riboflavin kinase | 4568 | 0.022 | -0.0968 | No |
| 10 | OLFM1 | OLFM1 Entrez, Source | olfactomedin 1 | 4900 | 0.017 | -0.1163 | No |
| 11 | DR1 | DR1 Entrez, Source | down-regulator of transcription 1, TBP-binding (negative cofactor 2) | 5479 | 0.009 | -0.1570 | No |
| 12 | CRLF3 | CRLF3 Entrez, Source | cytokine receptor-like factor 3 | 5514 | 0.008 | -0.1570 | No |
| 13 | CCNC | CCNC Entrez, Source | cyclin C | 5995 | 0.002 | -0.1924 | No |
| 14 | MEF2D | MEF2D Entrez, Source | MADS box transcription enhancer factor 2, polypeptide D (myocyte enhancer factor 2D) | 6025 | 0.002 | -0.1940 | No |
| 15 | RNF2 | RNF2 Entrez, Source | ring finger protein 2 | 6125 | 0.000 | -0.2014 | No |
| 16 | RCHY1 | RCHY1 Entrez, Source | ring finger and CHY zinc finger domain containing 1 | 6142 | 0.000 | -0.2025 | No |
| 17 | STAM2 | STAM2 Entrez, Source | signal transducing adaptor molecule (SH3 domain and ITAM motif) 2 | 6329 | -0.002 | -0.2158 | No |
| 18 | GATM | GATM Entrez, Source | glycine amidinotransferase (L-arginine:glycine amidinotransferase) | 6337 | -0.002 | -0.2156 | No |
| 19 | PDLIM3 | PDLIM3 Entrez, Source | PDZ and LIM domain 3 | 6654 | -0.006 | -0.2373 | No |
| 20 | EP300 | EP300 Entrez, Source | E1A binding protein p300 | 7440 | -0.016 | -0.2912 | No |
| 21 | SON | SON Entrez, Source | SON DNA binding protein | 7481 | -0.016 | -0.2890 | No |
| 22 | ANP32E | ANP32E Entrez, Source | acidic (leucine-rich) nuclear phosphoprotein 32 family, member E | 7651 | -0.019 | -0.2957 | No |
| 23 | BRMS1 | BRMS1 Entrez, Source | breast cancer metastasis suppressor 1 | 7742 | -0.020 | -0.2961 | No |
| 24 | NUP153 | NUP153 Entrez, Source | nucleoporin 153kDa | 7992 | -0.023 | -0.3073 | No |
| 25 | STCH | STCH Entrez, Source | stress 70 protein chaperone, microsome-associated, 60kDa | 8120 | -0.025 | -0.3088 | No |
| 26 | PPM1A | PPM1A Entrez, Source | protein phosphatase 1A (formerly 2C), magnesium-dependent, alpha isoform | 8630 | -0.033 | -0.3366 | No |
| 27 | POLE3 | POLE3 Entrez, Source | polymerase (DNA directed), epsilon 3 (p17 subunit) | 9201 | -0.040 | -0.3666 | No |
| 28 | DLG1 | DLG1 Entrez, Source | discs, large homolog 1 (Drosophila) | 9232 | -0.041 | -0.3558 | No |
| 29 | ISL1 | ISL1 Entrez, Source | ISL1 transcription factor, LIM/homeodomain, (islet-1) | 9285 | -0.042 | -0.3465 | No |
| 30 | PDE4D | PDE4D Entrez, Source | phosphodiesterase 4D, cAMP-specific (phosphodiesterase E3 dunce homolog, Drosophila) | 10269 | -0.058 | -0.4018 | No |
| 31 | TESK2 | TESK2 Entrez, Source | testis-specific kinase 2 | 11384 | -0.084 | -0.4589 | Yes |
| 32 | ATP6V1C1 | ATP6V1C1 Entrez, Source | ATPase, H+ transporting, lysosomal 42kDa, V1 subunit C1 | 11466 | -0.085 | -0.4376 | Yes |
| 33 | RPS6KB1 | RPS6KB1 Entrez, Source | ribosomal protein S6 kinase, 70kDa, polypeptide 1 | 11564 | -0.088 | -0.4167 | Yes |
| 34 | ARHGEF5 | ARHGEF5 Entrez, Source | Rho guanine nucleotide exchange factor (GEF) 5 | 11836 | -0.097 | -0.4061 | Yes |
| 35 | TMF1 | TMF1 Entrez, Source | TATA element modulatory factor 1 | 12133 | -0.109 | -0.3936 | Yes |
| 36 | CASP2 | CASP2 Entrez, Source | caspase 2, apoptosis-related cysteine peptidase (neural precursor cell expressed, developmentally down-regulated 2) | 12210 | -0.112 | -0.3636 | Yes |
| 37 | CRY1 | CRY1 Entrez, Source | cryptochrome 1 (photolyase-like) | 12383 | -0.120 | -0.3380 | Yes |
| 38 | SSR1 | SSR1 Entrez, Source | signal sequence receptor, alpha (translocon-associated protein alpha) | 12483 | -0.127 | -0.3049 | Yes |
| 39 | GCH1 | GCH1 Entrez, Source | GTP cyclohydrolase 1 (dopa-responsive dystonia) | 12570 | -0.133 | -0.2689 | Yes |
| 40 | ZBTB1 | ZBTB1 Entrez, Source | zinc finger and BTB domain containing 1 | 12766 | -0.148 | -0.2361 | Yes |
| 41 | PPAT | PPAT Entrez, Source | phosphoribosyl pyrophosphate amidotransferase | 12816 | -0.154 | -0.1904 | Yes |
| 42 | SLC12A2 | SLC12A2 Entrez, Source | solute carrier family 12 (sodium/potassium/chloride transporters), member 2 | 12841 | -0.158 | -0.1416 | Yes |
| 43 | GCLC | GCLC Entrez, Source | glutamate-cysteine ligase, catalytic subunit | 12866 | -0.161 | -0.0920 | Yes |
| 44 | TRIM23 | TRIM23 Entrez, Source | tripartite motif-containing 23 | 13053 | -0.198 | -0.0426 | Yes |
| 45 | FNBP4 | FNBP4 Entrez, Source | formin binding protein 4 | 13066 | -0.201 | 0.0208 | Yes |

