Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | double_gel_and_single_gel_rma_expression_values_collapsed_to_symbols.class.cls #double_gel_8d_versus_double_gel_72h.class.cls #double_gel_8d_versus_double_gel_72h_repos |
| Phenotype | class.cls#double_gel_8d_versus_double_gel_72h_repos |
| Upregulated in class | double_gel_8d |
| GeneSet | FRASOR_ER_DN |
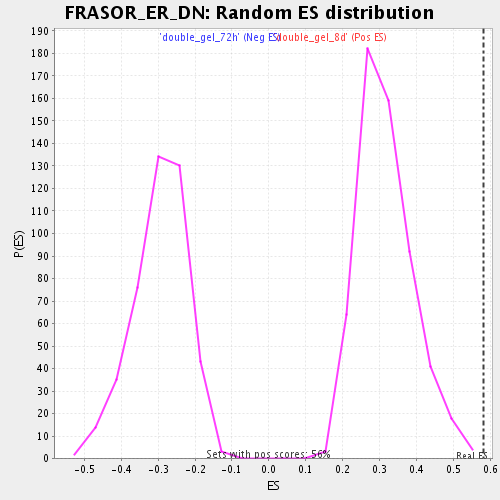
| Enrichment Score (ES) | 0.5806696 |
| Normalized Enrichment Score (NES) | 1.8344219 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.026933268 |
| FWER p-Value | 0.905 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | CD24 | CD24 Entrez, Source | CD24 molecule | 17 | 0.726 | 0.1430 | Yes |
| 2 | DBN1 | DBN1 Entrez, Source | drebrin 1 | 151 | 0.362 | 0.2049 | Yes |
| 3 | ABCG1 | ABCG1 Entrez, Source | ATP-binding cassette, sub-family G (WHITE), member 1 | 212 | 0.312 | 0.2624 | Yes |
| 4 | ECH1 | ECH1 Entrez, Source | enoyl Coenzyme A hydratase 1, peroxisomal | 241 | 0.289 | 0.3177 | Yes |
| 5 | LAMB2 | LAMB2 Entrez, Source | laminin, beta 2 (laminin S) | 351 | 0.229 | 0.3551 | Yes |
| 6 | TGFB2 | TGFB2 Entrez, Source | transforming growth factor, beta 2 | 367 | 0.223 | 0.3983 | Yes |
| 7 | IER3 | IER3 Entrez, Source | immediate early response 3 | 598 | 0.165 | 0.4137 | Yes |
| 8 | CBX6 | CBX6 Entrez, Source | chromobox homolog 6 | 704 | 0.150 | 0.4357 | Yes |
| 9 | ENC1 | ENC1 Entrez, Source | ectodermal-neural cortex (with BTB-like domain) | 1029 | 0.119 | 0.4349 | Yes |
| 10 | ANXA3 | ANXA3 Entrez, Source | annexin A3 | 1062 | 0.117 | 0.4556 | Yes |
| 11 | ERBB2 | ERBB2 Entrez, Source | v-erb-b2 erythroblastic leukemia viral oncogene homolog 2, neuro/glioblastoma derived oncogene homolog (avian) | 1072 | 0.116 | 0.4780 | Yes |
| 12 | ABCC5 | ABCC5 Entrez, Source | ATP-binding cassette, sub-family C (CFTR/MRP), member 5 | 1263 | 0.106 | 0.4849 | Yes |
| 13 | DUSP1 | DUSP1 Entrez, Source | dual specificity phosphatase 1 | 1297 | 0.104 | 0.5031 | Yes |
| 14 | MYO1B | MYO1B Entrez, Source | myosin IB | 1311 | 0.104 | 0.5227 | Yes |
| 15 | L1CAM | L1CAM Entrez, Source | L1 cell adhesion molecule | 1321 | 0.103 | 0.5426 | Yes |
| 16 | SELENBP1 | SELENBP1 Entrez, Source | selenium binding protein 1 | 1492 | 0.094 | 0.5485 | Yes |
| 17 | BIK | BIK Entrez, Source | BCL2-interacting killer (apoptosis-inducing) | 1540 | 0.092 | 0.5632 | Yes |
| 18 | NELL2 | NELL2 Entrez, Source | NEL-like 2 (chicken) | 1551 | 0.091 | 0.5807 | Yes |
| 19 | BMP4 | BMP4 Entrez, Source | bone morphogenetic protein 4 | 1840 | 0.078 | 0.5746 | No |
| 20 | DUSP4 | DUSP4 Entrez, Source | dual specificity phosphatase 4 | 2162 | 0.068 | 0.5640 | No |
| 21 | FMO5 | FMO5 Entrez, Source | flavin containing monooxygenase 5 | 2218 | 0.067 | 0.5731 | No |
| 22 | STXBP1 | STXBP1 Entrez, Source | syntaxin binding protein 1 | 2596 | 0.058 | 0.5562 | No |
| 23 | SLIT2 | SLIT2 Entrez, Source | slit homolog 2 (Drosophila) | 2885 | 0.051 | 0.5447 | No |
| 24 | CLU | CLU Entrez, Source | clusterin | 3299 | 0.043 | 0.5221 | No |
| 25 | STC1 | STC1 Entrez, Source | stanniocalcin 1 | 3324 | 0.042 | 0.5286 | No |
| 26 | PIB5PA | PIB5PA Entrez, Source | phosphatidylinositol (4,5) bisphosphate 5-phosphatase, A | 3373 | 0.041 | 0.5332 | No |
| 27 | LITAF | LITAF Entrez, Source | lipopolysaccharide-induced TNF factor | 3654 | 0.036 | 0.5193 | No |
| 28 | ARNT2 | ARNT2 Entrez, Source | aryl-hydrocarbon receptor nuclear translocator 2 | 3702 | 0.035 | 0.5227 | No |
| 29 | CTSL | CTSL Entrez, Source | cathepsin L | 3892 | 0.032 | 0.5149 | No |
| 30 | BCAS1 | BCAS1 Entrez, Source | breast carcinoma amplified sequence 1 | 3953 | 0.031 | 0.5165 | No |
| 31 | GNS | GNS Entrez, Source | glucosamine (N-acetyl)-6-sulfatase (Sanfilippo disease IIID) | 4298 | 0.025 | 0.4957 | No |
| 32 | BLNK | BLNK Entrez, Source | B-cell linker | 4539 | 0.022 | 0.4820 | No |
| 33 | IL1R1 | IL1R1 Entrez, Source | interleukin 1 receptor, type I | 4666 | 0.020 | 0.4765 | No |
| 34 | FOLH1 | FOLH1 Entrez, Source | folate hydrolase (prostate-specific membrane antigen) 1 | 5352 | 0.010 | 0.4270 | No |
| 35 | CTSH | CTSH Entrez, Source | cathepsin H | 5387 | 0.010 | 0.4264 | No |
| 36 | TRG@ | TRG@ Entrez, Source | T cell receptor gamma locus | 5636 | 0.007 | 0.4091 | No |
| 37 | IFI30 | IFI30 Entrez, Source | interferon, gamma-inducible protein 30 | 6191 | -0.000 | 0.3675 | No |
| 38 | NDRG1 | NDRG1 Entrez, Source | N-myc downstream regulated gene 1 | 6962 | -0.010 | 0.3116 | No |
| 39 | KYNU | KYNU Entrez, Source | kynureninase (L-kynurenine hydrolase) | 7289 | -0.014 | 0.2899 | No |
| 40 | EPOR | EPOR Entrez, Source | erythropoietin receptor | 7361 | -0.015 | 0.2876 | No |
| 41 | PRPS1 | PRPS1 Entrez, Source | phosphoribosyl pyrophosphate synthetase 1 | 8663 | -0.033 | 0.1963 | No |
| 42 | PVALB | PVALB Entrez, Source | parvalbumin | 9188 | -0.040 | 0.1649 | No |
| 43 | MB | MB Entrez, Source | myoglobin | 10581 | -0.064 | 0.0729 | No |
| 44 | TGFB3 | TGFB3 Entrez, Source | transforming growth factor, beta 3 | 10957 | -0.072 | 0.0590 | No |
| 45 | EFNA1 | EFNA1 Entrez, Source | ephrin-A1 | 11985 | -0.102 | 0.0021 | No |
| 46 | GNE | GNE Entrez, Source | glucosamine (UDP-N-acetyl)-2-epimerase/N-acetylmannosamine kinase | 12371 | -0.119 | -0.0031 | No |
| 47 | PPP1R3C | PPP1R3C Entrez, Source | protein phosphatase 1, regulatory (inhibitor) subunit 3C | 13293 | -0.383 | 0.0037 | No |